

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
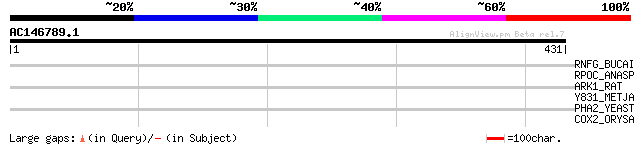
Query= AC146789.1 + phase: 0
(431 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RNFG_BUCAI (P57217) Electron transport complex protein rnfG homolog 33 1.4
RPOC_ANASP (P22704) DNA-directed RNA polymerase gamma chain (EC ... 31 7.0
ARK1_RAT (P26817) Beta-adrenergic receptor kinase 1 (EC 2.7.1.12... 31 7.0
Y831_METJA (Q58241) Hypothetical protein MJ0831 30 9.2
PHA2_YEAST (P32452) Prephenate dehydratase (EC 4.2.1.51) (PDT) 30 9.2
COX2_ORYSA (P04373) Cytochrome c oxidase polypeptide II (EC 1.9.... 30 9.2
>RNFG_BUCAI (P57217) Electron transport complex protein rnfG homolog
Length = 164
Score = 33.1 bits (74), Expect = 1.4
Identities = 30/111 (27%), Positives = 52/111 (46%), Gaps = 13/111 (11%)
Query: 119 RSNKRIELIYLTTMADGYAGMRNYSWGAMTLAYLYGELADACRPGHR---ALGGSVTLLT 175
++ K + I TT DGY+G N + AY GE+ +A H+ +G + L
Sbjct: 34 KNKKPVAAIVETTAPDGYSGAIN----MLVAAYFNGEIINARVLSHKETPGIGDKIDLSI 89
Query: 176 TWFLKHFPGFF--SVDLNTDYLEKYPVAARWKLEKGHGEGITYRSLLDRIQ 224
+ ++ F G + S++ L KY K+E+ G IT +S+ + I+
Sbjct: 90 SNWITRFTGMYVASIEDKDFKLRKY----GGKIEQFTGATITPQSVTNSIK 136
>RPOC_ANASP (P22704) DNA-directed RNA polymerase gamma chain (EC
2.7.7.6) (RNAP gamma subunit) (Transcriptase gamma
chain) (RNA polymerase gamma subunit)
Length = 625
Score = 30.8 bits (68), Expect = 7.0
Identities = 21/75 (28%), Positives = 33/75 (44%), Gaps = 13/75 (17%)
Query: 177 WFLKHFPGFFSVDLNTD--------YLEKYPVAARWKLEKGHGEGITYRSLLDRIQLDDV 228
W+LK P + S+ L+ Y Y V L G+ E +TY+ LL Q ++
Sbjct: 116 WYLKGIPSYISILLDMPLRDVEQIVYFNSYVV-----LSPGNAETLTYKQLLSEDQWLEI 170
Query: 229 CWRPYEEHREIQDFE 243
+ Y E ++Q E
Sbjct: 171 EDQIYSEDSQLQGVE 185
>ARK1_RAT (P26817) Beta-adrenergic receptor kinase 1 (EC 2.7.1.126)
(Beta-ARK-1) (G-protein coupled receptor kinase 2)
Length = 689
Score = 30.8 bits (68), Expect = 7.0
Identities = 31/103 (30%), Positives = 50/103 (48%), Gaps = 11/103 (10%)
Query: 3 MGEMTVTLDDVACLMHLP-IEGRMLAHGKK---MPKHEGAALLMTYL-GVAQHEAEKICN 57
M ++ L DV+ LM + + A K +P+ +++ YL + EKI +
Sbjct: 1 MADLEAVLADVSYLMAMEKSKATPAARASKKILLPEPSIRSVMQKYLEDRGEVTFEKIFS 60
Query: 58 QEYGGYISYPRLRDFYTSYLGRANVLVGTEDPEEVEELERVRT 100
Q+ G Y RDFY ++L A LV E EE+E+ E++ T
Sbjct: 61 QKLG----YLLFRDFYLNHLEEAKPLV--EFYEEIEKYEKLET 97
>Y831_METJA (Q58241) Hypothetical protein MJ0831
Length = 432
Score = 30.4 bits (67), Expect = 9.2
Identities = 10/28 (35%), Positives = 18/28 (63%)
Query: 256 VRRVYRHLPERVLRQYGYVQTIPRHPTD 283
+ +V + + E++L YGY++ I H TD
Sbjct: 8 IEKVTKAIKEKILNHYGYIRVITHHDTD 35
>PHA2_YEAST (P32452) Prephenate dehydratase (EC 4.2.1.51) (PDT)
Length = 368
Score = 30.4 bits (67), Expect = 9.2
Identities = 40/158 (25%), Positives = 64/158 (40%), Gaps = 21/158 (13%)
Query: 252 IMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIVQMFIDFRTHTLKADARGV 311
+ G + Y H + L+Q+ + +DV LP SI Q F T D V
Sbjct: 43 LFLGPKGTYSH--QAALQQF-------QSTSDVEYLPAASIPQCFNQLENDT-SIDYSVV 92
Query: 312 QAGEDTR-RVADGYVLWYTRVSHPQILPPIPGDLPRPANEEQIIAEQWQRYEARSSPDTY 370
T +V Y L R+ + P P D R + ++IAEQ+ P T+
Sbjct: 93 PLENSTNGQVVFSYDLLRDRMIKKALSLPAPADTNRITPDIEVIAEQY-------VPITH 145
Query: 371 DMVSG-AVAYADAQLG--QEEVMSMTPQQWYEAMTHMR 405
++S + A LG +E ++ PQ W + ++R
Sbjct: 146 CLISPIQLPNGIASLGNFEEVIIHSHPQVWGQVECYLR 183
>COX2_ORYSA (P04373) Cytochrome c oxidase polypeptide II (EC
1.9.3.1)
Length = 260
Score = 30.4 bits (67), Expect = 9.2
Identities = 26/110 (23%), Positives = 43/110 (38%), Gaps = 17/110 (15%)
Query: 318 RRVADGYVLWYTRVSHPQILP------------PIPGDLPRPANEEQIIAEQWQR---YE 362
+R+ G + R P ++P + G L PA + I QW R Y
Sbjct: 75 QRIVHGTTIEIIRTIFPSVIPLFIAIPSFALLYSMDGVLVDPAITIKAIGHQWYRSYEYS 134
Query: 363 ARSSPDTYDMV--SGAVAYADAQLGQEEVMSMTPQQWYEAMTHMREQIAP 410
+S D + S + D +LGQ ++ + + A TH+R + P
Sbjct: 135 DYNSSDEQSLTFDSYTIPEDDPELGQSRLLEVDNRVVVPAKTHLRMIVTP 184
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,380,686
Number of Sequences: 164201
Number of extensions: 2465910
Number of successful extensions: 4952
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4951
Number of HSP's gapped (non-prelim): 7
length of query: 431
length of database: 59,974,054
effective HSP length: 113
effective length of query: 318
effective length of database: 41,419,341
effective search space: 13171350438
effective search space used: 13171350438
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146789.1


