

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146755.8 + phase: 0 /pseudo
(1711 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
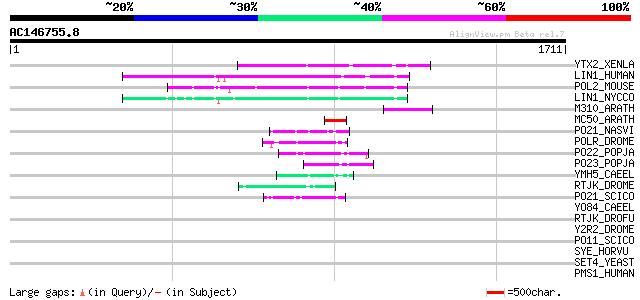
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 177 2e-43
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 175 1e-42
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 173 4e-42
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 166 7e-40
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 118 1e-25
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 84 4e-15
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 75 2e-12
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 73 8e-12
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 65 2e-09
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 55 1e-06
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 49 1e-04
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 48 3e-04
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 47 3e-04
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 45 0.002
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 42 0.015
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 39 0.094
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 37 0.36
SYE_HORVU (Q43768) Glutamyl-tRNA synthetase (EC 6.1.1.17) (Gluta... 36 0.80
SET4_YEAST (P42948) SET domain protein 4 34 3.0
PMS1_HUMAN (P54277) PMS1 protein homolog 1 (DNA mismatch repair ... 34 4.0
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein (ORF
2)
Length = 1308
Score = 177 bits (449), Expect = 2e-43
Identities = 165/615 (26%), Positives = 269/615 (42%), Gaps = 43/615 (6%)
Query: 703 DLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTK--LSSTRAD 760
D ++FF+A + +I L +GT + +++ AR+++ +LF+ +S +
Sbjct: 369 DRGSRFFYALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRARSFYQNLFSPDPISPDACE 428
Query: 761 VISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEI 820
+ +S+ L TL+E A M +K PG DG F+Q FW+ G +
Sbjct: 429 ELWDGLPVVSERRKERLETPITLDELSQALRLMPHNKSPGLDGLTIEFFQFFWDTLGPDF 488
Query: 821 HKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKN 880
H+ E + G P ++L+PK +K+WRP++L + YK+VAK ++ RLK+
Sbjct: 489 HRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKS 548
Query: 881 VLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWE 940
VL + I QS VPGR+I DN + +L+H+ + + L LD KA+DR+D +
Sbjct: 549 VLAEVIHPDQSYTVPGRTIFDNVFLIRDLLHFAR---RTGLSLAFLSLDQEKAFDRVDHQ 605
Query: 941 YLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILC 1000
YL + F +++ ++ + + V +N PL GRG+RQG PLS L+ L
Sbjct: 606 YLIGTLQAYSFGPQFVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLA 665
Query: 1001 AEGLSALIRDAERRGVI--TGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTT 1058
E L+R V+ R+ SA A +L A D RA E +
Sbjct: 666 IEPFLCLLRKRLTGLVLKEPDMRVVLSAYADDVILVAQDLVDLERAQECQ--------EV 717
Query: 1059 YEAASGQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVLGTGKYLGL-PSMIGRSKKAT 1117
Y AAS IN KS + + + + + KYLG+ S
Sbjct: 718 YAAASSARINWSKSSGLLEGSLKVDFLPPAFRDISWESKI--IKYLGVYLSAEEYPVSQN 775
Query: 1118 FKFVKDRIWNRINSWS--SRCLSQAGREILIKSVLQSIPSYVMSIFLLPSTFIGEIEKML 1175
F +++ + R+ W ++ LS GR ++I ++ S Y + FI +I++ L
Sbjct: 776 FIELEECVLTRLGKWKGFAKVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRL 835
Query: 1176 NAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRA----FNLAMIGKQAWKLISNP 1231
F W G HW+S S+P GG G +R+ F L I + L ++P
Sbjct: 836 LDFLW-------IGKHWVSAGVSSLPLKEGGQGVVCIRSQVHTFRLQQIQRY---LYADP 885
Query: 1232 DSLITKLLKAKY--FPHSDYFSASIGHNPSYVWRSLWSVRDYLKHGLK-WSI------GT 1282
L + Y + Y P R+L ++ Y + LK WS+ G
Sbjct: 886 SPQWCTLASSFYRQVRNMGYDRQLFIIEPEGFLRNLSTLPAYYQDTLKTWSMVSVLRQGA 945
Query: 1283 GETISVWNQPWLKDP 1297
E + N+P L +P
Sbjct: 946 TEGEDILNEPLLYNP 960
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 175 bits (443), Expect = 1e-42
Identities = 196/922 (21%), Positives = 380/922 (40%), Gaps = 67/922 (7%)
Query: 349 GPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDS 408
G ++ + N GL +P L +++ P + ET LK I + +
Sbjct: 2 GSNSHITILTLNVNGLNAPIKRHRLANWIKSQDPSVCCIQETHLTCRDTHRLK-IKGWRN 60
Query: 409 CFTVDRIGRGGGLAFLWKNSANCTITNFSQN---HIDVEVDDLLIGKWRLTGFYGMPENG 465
+ + + G+A L + + T ++ H + + + + Y P G
Sbjct: 61 IYQANGKQKKAGVAILVSDKTDFKPTKIKRDKEGHYIMVKGSIQQEELTILNIYA-PNTG 119
Query: 466 RRKESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFD 525
+ L +L R ++GDFN LS+ ++ R ++ I+ A+ A L D
Sbjct: 120 APRFIKQVLSDLQRDLDSHTIIMGDFNTPLSTLDRSTR-QKINKDIQELNSALHQADLID 178
Query: 526 LHMTGYA-FTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTC 584
++ T + T + K D L + + E ++ SDH + L
Sbjct: 179 IYRTLHPKSTEYTFFSAPHHTYSKTDHILGSKTLLSKCKRT--EIITNCLSDHSAIKLEL 236
Query: 585 KPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYPEEGITQKLSYCAEDLTDWSR--- 641
+ + +K N L + +++ K + E + +Y ++L D ++
Sbjct: 237 RIKKLTQNHSTTWKLNNLLLNDYWVHNEMKAEIKKFFETNENKDTTY--QNLWDTAKAVC 294
Query: 642 -------NNNNFRRDISKVQKKIEKLR-------THVTAANVSYFNSLKNKLDKLLVQDD 687
N + +++ SK+ I +L+ T+ A+ ++ +L ++ Q
Sbjct: 295 RGKFIALNAHKRKQERSKIDTLISQLKELEKQEQTNSKASRRQEIIKIRAELKEIETQKT 354
Query: 688 LFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYF 747
L ++++++++ + + +R+ N+I+ ++N G + E+Q R Y+
Sbjct: 355 LQKINESRSWFFEKINKIDRPLARLIKKKREKNQIDTIKNDRGDITTDPTEIQTTIREYY 414
Query: 748 SHLFTKLSSTRADVISKVAT----SISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDG 803
HL+ ++ + T ++ ++ L T E + S+ K PGP+G
Sbjct: 415 KHLYANKLENLEEMDKFLDTYTLPRLNQEEVESLNRPITSSEIEAIINSLPNKKSPGPEG 474
Query: 804 FNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKD-WRPIAL 862
F FYQ + + K + G+ P +I LIPK TT K+ +RPI+L
Sbjct: 475 FTAEFYQRYKEELVPFLLKLFQSIEKEGILPNSFYEASIILIPKPGRDTTKKENFRPISL 534
Query: 863 CNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQG 922
N+ K++ K+LAN+++ + K I Q F+P N +I +I ++ +TK +
Sbjct: 535 MNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGWFNIRKSINIIQHIN-RTKDT-N 592
Query: 923 DVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIP 982
+ + +D KA+D++ ++ + ++ +++ I + +++NG ++
Sbjct: 593 HMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPPL 652
Query: 983 GRGIRQGDPLSPYLFILCAEGLSALIR-DAERRGVITGTRICTSAPAISHLLFADDCFLF 1041
G RQG PLSP L + E L+ IR + E +G+ G + LFADD ++
Sbjct: 653 KTGTRQGCPLSPLLPNIVLEVLARAIRQEKEIKGIQLGKE------EVKLSLFADDMIVY 706
Query: 1042 FRASEQEASVMKNILTTYEAASGQAINLQKSEMYC---SRNTPTECQNRIATTLGVKQVL 1098
A + +++ + SG IN+QKS+ + +R T ++ + + T+ K++
Sbjct: 707 LENPIVSAQNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPFTIASKRI- 765
Query: 1099 GTGKYLGLPSMIGRSKKATFKFVKDRIWNRI----NSWSSRCLSQAGREILIKSVLQSIP 1154
KYLG+ + R K FK + N I N W + S GR ++K + +P
Sbjct: 766 ---KYLGI--QLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAI--LP 818
Query: 1155 SYVMSI----FLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFK 1210
+ LP TF E+EK F W A+ ++ LS GG+
Sbjct: 819 KVIYRFNAIPIKLPMTFFTELEKTTLKFIWNQKRAH------IAKSTLSQKNKAGGITLP 872
Query: 1211 SLRAFNLAMIGKQAWKLISNPD 1232
+ + A + K AW N D
Sbjct: 873 DFKLYYKATVTKTAWYWYQNRD 894
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 173 bits (438), Expect = 4e-42
Identities = 182/773 (23%), Positives = 326/773 (41%), Gaps = 60/773 (7%)
Query: 487 VIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGLFDLHMTGYAFT-WFKSLGTARAV 545
++GDFN LSS+++ + + ++ + + L D++ T Y T +
Sbjct: 168 IVGDFNTPLSSKDRSWKQKLNRDTVK-LTEVMKQMDLTDIYRTFYPKTKGYTFFSAPHGT 226
Query: 546 EEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYPLWLTCKPVTAVNRNPHRFKFENAWLA 605
K+D + + N + N +E + SDH+ L L + +K N L
Sbjct: 227 FSKIDHIIGHKTGLNRYKN--IEIVPCILSDHHGLRLIFNNNINNGKPTFTWKLNNTLLN 284
Query: 606 EPDFKQQVQQRWKLYPEEGITQKLSYCAEDLTDWSRNNNNFRRD----ISKVQKKIE--- 658
+ K+ +++ K + E + +Y +L D + F R +S +KK E
Sbjct: 285 DTLVKEGIKKEIKDFLEFNENEATTY--PNLWDTMKA---FLRGKLIALSASKKKRETAH 339
Query: 659 --KLRTHVTAANVSYFNS-----------LKNKLDKLLVQDDLFWKQRAKTFWYKEGDLN 705
L TH+ A NS L+ +++++ + + + +++++++ +
Sbjct: 340 TSSLTTHLKALEKKEANSPKRSRRQEIIKLRGEINQVETRRTIQRINQTRSWFFEKINKI 399
Query: 706 TKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLF-TKLSSTRADVISK 764
K R I + N G + +E+QN R+++ L+ TKL + D + K
Sbjct: 400 DKPLARLTKGHRDKILINKIRNEKGDITTDPEEIQNTIRSFYKRLYSTKLENL--DEMDK 457
Query: 765 V-----ATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSE 819
++ D L + + +E + S+ K PGPDGF+ FYQ F
Sbjct: 458 FLDRYQVPKLNQDQVDHLNSPISPKEIEAVINSLPTKKSPGPDGFSAEFYQTFKEDLIPI 517
Query: 820 IHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTT-MKDWRPIALCNVVYKMVAKVLANRL 878
+HK + G P I LIPK T ++++RPI+L N+ K++ K+LANR+
Sbjct: 518 LHKLFHKIEVEGTLPNSFYEATITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRI 577
Query: 879 KNVLDKCISTSQSAFVPGRSILDNALVAIELIHYM-KAKTKGSQGDVALKLDISKAYDRL 937
+ + I Q F+PG N +I +IHY+ K K K + + LD KA+D++
Sbjct: 578 QEHIKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINKLKDK---NHMIISLDAEKAFDKI 634
Query: 938 DWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLF 997
++ ++ + ++ I + VNG ++ + G RQG PLSPYLF
Sbjct: 635 QHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLF 694
Query: 998 ILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILT 1057
+ E L+ IR + I G +I IS L ADD ++ + + N++
Sbjct: 695 NIVLEVLARAIRQQKE---IKGIQIGKEEVKIS--LLADDMIVYISDPKNSTRELLNLIN 749
Query: 1058 TYEAASGQAINLQKSEMYC-SRNTPTECQNRIATTLGVKQVLGTGKYLG--LPSMIGRSK 1114
++ G IN KS + ++N E + R T + V KYLG L +
Sbjct: 750 SFGEVVGYKINSNKSMAFLYTKNKQAEKEIRETTPFSI--VTNNIKYLGVTLTKEVKDLY 807
Query: 1115 KATFKFVKDRIWNRINSWSSRCLSQAGREILIKSVL--QSIPSYVMSIFLLPSTFIGEIE 1172
FK +K I + W S GR ++K + ++I + +P+ F E+E
Sbjct: 808 DKNFKSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELE 867
Query: 1173 KMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGFKSLRAFNLAMIGKQAW 1225
+ F W + ++ L +T GG+ L+ + A++ K AW
Sbjct: 868 GAICKFVWNNKKPR------IAKSLLKDKRTSGGITMPDLKLYYRAIVIKTAW 914
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 166 bits (419), Expect = 7e-40
Identities = 190/916 (20%), Positives = 373/916 (39%), Gaps = 69/916 (7%)
Query: 349 GPPRKMSFIVWNCRGLGSPSTVPTLKYLVRTYKPEGIFLSETMAALNKIEELKYILSFDS 408
G + +S N GL P L ++ KP+ + E+ L LK + + S
Sbjct: 2 GLSKGLSIFSINVNGLNCPLKRHRLADWIQKLKPDICCIQESHLTLKDKYRLK-VKGWSS 60
Query: 409 CFTVDRIGRGGGLAFLWKNSANCTITNFSQN---HIDVEVDDLLIGKWRLTGFYGMPENG 465
F + + G+A L+ ++ T ++ H + + + Y N
Sbjct: 61 IFQANGKQKKAGIAILFADAIGFKPTKIRKDKDGHFIFVKGNTQYDEISIINIYAPNHNA 120
Query: 466 RR--KESWAFLKNLARTSSLPWCVIGDFNDILSSEEKKGRSERAPWLIRGFRQAVLDAGL 523
+ +E+ + NL ++S+ V+GDFN L+ ++ + + + ++ + L
Sbjct: 121 PQFIRETLTDMSNLISSTSI---VVGDFNTPLAVLDRSSKKKLSKEIL-DLNSTIQHLDL 176
Query: 524 FDLHMTGYA----FTWFKSLGTARAVEEKLDRALVNSDWCNMFPNAGVECLSTTSSDHYP 579
D++ T + +T+F S A K+D L + + F +E + SDH+
Sbjct: 177 TDIYRTFHPNKTEYTFFSS---AHGTYSKIDHILGHKSNLSKFKK--IEIIPCIFSDHHG 231
Query: 580 LWLTCKPVTAVNRNPHRFKFENAWLAEPDFKQQVQQRWKLYPEEGITQKLSYCAEDLTDW 639
+ + ++ + +K N L + ++++ + E+ Q +Y ++L D
Sbjct: 232 IKVELNNNRNLHTHTKTWKLNNLMLKDTWVIDEIKKEITKFLEQNNNQDTNY--QNLWDT 289
Query: 640 SR--------------------NNNNFRRDISKVQKKIEKLRTHVTAANVSYFNSLKNKL 679
++ NN + +++K+ ++ + N++
Sbjct: 290 AKAVLRGKFIALQAFLKKTEREEVNNLMGHLKQLEKEEHSNPKPSRRKEITKIRAELNEI 349
Query: 680 DKLLVQDDLFWKQRAKTFWYKEGDLNTKFFHAAATSRRKVNRIEHLENSNGTVCRKEDEL 739
+ + + ++K++++++ + K +R + I + N N + E+
Sbjct: 350 ENKRIIQQI---NKSKSWFFEKINKIDKPLANLTRKKRVKSLISSIRNGNDEITTDPSEI 406
Query: 740 QNIARNYF----SHLFTKLSSTRADVISKVATSISDDDNCFLTAAFTLEEFKTAAFSMQA 795
Q I Y+ SH + L + + +S + L + E + ++
Sbjct: 407 QKILNEYYKKLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPK 466
Query: 796 DKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPK-GDAQTTM 854
K PGPDGF FYQ F + + G+ P NI LIPK G T
Sbjct: 467 KKSPGPDGFTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPGKDPTRK 526
Query: 855 KDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELI-HYM 913
+++RPI+L N+ K++ K+L NR++ + K I Q F+PG N +I +I H
Sbjct: 527 ENYRPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHIN 586
Query: 914 KAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVN 973
K K K + L +D KA+D + ++ + ++ +++ I +++N
Sbjct: 587 KLKNK---DHMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILN 643
Query: 974 GVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLL 1033
GV++ G RQG PLSP LF + E L+ IR+ + I G I + +S L
Sbjct: 644 GVKLKSFPLRSGTRQGCPLSPLLFNIVMEVLAIAIREEK---AIKGIHIGSEEIKLS--L 698
Query: 1034 FADDCFLFFRASEQEASVMKNILTTYEAASGQAINLQKSEMYCSRNTPTECQNRIATTLG 1093
FADD ++ + + + ++ Y SG IN KS + N + + + ++
Sbjct: 699 FADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNN-NQAEKTVKDSIP 757
Query: 1094 VKQVLGTGKYLG--LPSMIGRSKKATFKFVKDRIWNRINSWSSRCLSQAGREILIK-SVL 1150
V KYLG L + K ++ ++ I +N W + S GR ++K S+L
Sbjct: 758 FTVVPKKMKYLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSIL 817
Query: 1151 -QSIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTFGGMGF 1209
++I ++ P ++ ++EK++ F W ++ LS GG+
Sbjct: 818 PKAIYNFNAIPIKAPLSYFKDLEKIILHFIWNQKKPQ------IAKTLLSNKNKAGGITL 871
Query: 1210 KSLRAFNLAMIGKQAW 1225
LR + +++ K AW
Sbjct: 872 PDLRLYYKSIVIKTAW 887
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 118 bits (296), Expect = 1e-25
Identities = 56/153 (36%), Positives = 88/153 (56%), Gaps = 2/153 (1%)
Query: 1152 SIPSYVMSIFLLPSTFIGEIEKMLNAFWWGHNSANSRGMHWLSWERLSVPKTF-GGMGFK 1210
++P Y MS F L ++ + FWW + N R + W++W++L K GG+GF+
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWS-SCENKRKISWVAWQKLCKSKEDDGGLGFR 60
Query: 1211 SLRAFNLAMIGKQAWKLISNPDSLITKLLKAKYFPHSDYFSASIGHNPSYVWRSLWSVRD 1270
L FN A++ KQ++++I P +L+++LL+++YFPHS S+G PSY WRS+ R+
Sbjct: 61 DLGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGRE 120
Query: 1271 YLKHGLKWSIGTGETISVWNQPWLKDPLCLQPV 1303
L GL +IG G VW W+ D L P+
Sbjct: 121 LLSRGLLRTIGDGIHTKVWLDRWIMDETPLPPL 153
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 83.6 bits (205), Expect = 4e-15
Identities = 38/67 (56%), Positives = 49/67 (72%)
Query: 971 LVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAIS 1030
++NG G + P RG+RQGDPLSPYLFILC E LS L R A+ +G + G R+ ++P I+
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRIN 72
Query: 1031 HLLFADD 1037
HLLFADD
Sbjct: 73 HLLFADD 79
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 75.1 bits (183), Expect = 2e-12
Identities = 67/250 (26%), Positives = 104/250 (40%), Gaps = 23/250 (9%)
Query: 800 GPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRP 859
GPDG WN I + +G P + LIPK +RP
Sbjct: 336 GPDGMTT----TAWNSIDECIKSLFNMIMYHGQCPRRYLDSRTVLIPKEPGTMDPACFRP 391
Query: 860 IALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKG 919
+++ +V + ++LANR+ + T Q AF+ + +N + +I + K KG
Sbjct: 392 LSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENTSLLSAMIKEARMKIKG 449
Query: 920 SQGDVALKLDISKAYDRLDWEYLRDIMIQMRFS---EKWIRWIMLCVETVDYTVLVNGVQ 976
+A+ LD+ KA+D ++ + D + + + +I W+ +T V G
Sbjct: 450 LY--IAI-LDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKGRW 506
Query: 977 VGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSAPAISHLLFAD 1036
+ P RG+RQGDPLSP LF + DA R + T A I L+FAD
Sbjct: 507 IRP---ARGVRQGDPLSPLLF--------NCVMDAVLRRLPENTGFLMGAEKIGALVFAD 555
Query: 1037 DCFLFFRASE 1046
D L E
Sbjct: 556 DLVLLAETRE 565
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 72.8 bits (177), Expect = 8e-12
Identities = 68/271 (25%), Positives = 115/271 (42%), Gaps = 34/271 (12%)
Query: 779 AAFTLEEFKTAAFSMQADKCPGPDGFNP--------GFYQNFWNMCGSEIHKAGCEWLEN 830
+A T ++ + + S+ + PGPDG P G N+ L
Sbjct: 342 SAITEQDLRASRVSLSSS--PGPDGITPKSAREVPSGIMLRIMNLI-----------LWC 388
Query: 831 GVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQ 890
G P + IPK +D+RPI++ +V+ + + +LA RL + ++ Q
Sbjct: 389 GNLPHSIRLARTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSINW--DPRQ 446
Query: 891 SAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMR 950
F+P DNA + ++L+ ++ K + LD+SKA+D L + D +
Sbjct: 447 RGFLPTDGCADNATI-VDLV--LRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYG 503
Query: 951 FSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRD 1010
+ ++ ++ E ++ +G +P RG++QGDPLSP LF L + L +
Sbjct: 504 APKGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTL-- 561
Query: 1011 AERRGVITGTRICTSAPAISHLLFADDCFLF 1041
G G I +A FADD LF
Sbjct: 562 PSEIGAKVGNAITNAA------AFADDLVLF 586
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 65.1 bits (157), Expect = 2e-09
Identities = 70/289 (24%), Positives = 119/289 (40%), Gaps = 26/289 (8%)
Query: 828 LENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCIS 887
L G P + LIPK +WRPI + + + +++ ++LA RL+ ++ +
Sbjct: 47 LLRGHVPTPWTAMRTTLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVE--LH 104
Query: 888 TSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMI 947
+Q + L N+L+ L Y+ ++ + + + LD+ KA+D + + +
Sbjct: 105 PAQKGYARIDGTLVNSLL---LDTYISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQ 161
Query: 948 QMRFSEKWIRWIMLCVETVDYTVLVN-GVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSA 1006
++ E +I + T+ V G Q + RG++QGDPLSP+LF + L
Sbjct: 162 RLGIDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLC 221
Query: 1007 LIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTT---YEAAS 1063
++ G G I L FADD L E ++ L T +
Sbjct: 222 SLQSTPGIGGTIGEE------KIPVLAFADDLLLL----EDNDVLLPTTLATVANFFRLR 271
Query: 1064 GQAINLQKSEMYCSRNTPTECQNRIATTLGVKQVL-------GTGKYLG 1105
G ++N +KS + C R L V VL T +YLG
Sbjct: 272 GMSLNAKKSVSISVAASGGVCIPRTKPFLRVDNVLLPVTDRMQTFRYLG 320
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 55.5 bits (132), Expect = 1e-06
Identities = 58/226 (25%), Positives = 94/226 (40%), Gaps = 19/226 (8%)
Query: 907 IELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETV 966
I L HY+K + + + LDI KA+D + + M + +IM +
Sbjct: 7 IMLEHYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMSTITDA 66
Query: 967 DYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSALIRDAERRGVITGTRICTSA 1026
++V G + G++QGDPLSP LF + + L + D E+ G T A
Sbjct: 67 YTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLND-EQPGA-----SMTPA 120
Query: 1027 PAISHLLFADDCFLFFRASEQEASVMKNILTT--YEAASGQAINLQK-----SEMYCSRN 1079
I+ L FADD L +++ V ++ TT Y G +N +K + R+
Sbjct: 121 CKIASLAFADDLLLL---EDRDIDVPNSLATTCAYFRTRGMTLNPEKCASISAATVSGRS 177
Query: 1080 TPTECQNRIATTLGVKQV--LGTGKYLGLP-SMIGRSKKATFKFVK 1122
P + +K + + T KYLGL S G +K + +
Sbjct: 178 VPRSKPSFTIDGRYIKPLGGINTFKYLGLTFSSTGVAKPTVYNLTR 223
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 48.9 bits (115), Expect = 1e-04
Identities = 51/236 (21%), Positives = 93/236 (38%), Gaps = 25/236 (10%)
Query: 824 GCEWLENGVFPPHLNSTNIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLD 883
G + + P I IPK ++ ++RPI+L + +++ +++ +R+++
Sbjct: 652 GMQSFSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYS 711
Query: 884 KCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDWEYLR 943
+S Q F+ RS + + +I L H + K L D +KA+D++ L
Sbjct: 712 HLLSPHQHGFLNFRSCPSSLVRSISLYHSILKNEKSLD---ILFFDFAKAFDKVSHPILL 768
Query: 944 DIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIP-GRGIRQGDPLSPYLFILCAE 1002
+ + W + ++V +N P G+ QG P LFIL
Sbjct: 769 KKLALFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFILF-- 826
Query: 1003 GLSALIRDAERRGVITGTRICTSAPAISHLLFADDCFLFFRASEQEASVMKNILTT 1058
++ L+ D E P I FADD +F S ++N + T
Sbjct: 827 -INDLLIDLE--------------PNIHVSCFADDIKIF----HHNPSTLQNSIDT 863
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 47.8 bits (112), Expect = 3e-04
Identities = 73/310 (23%), Positives = 119/310 (37%), Gaps = 28/310 (9%)
Query: 706 TKFFHAAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADVISKV 765
TK F A T + + +N +G CR E + N FT + + +V
Sbjct: 368 TKRFKAQPTPKSAI------KNPSGGWCRTSLEKTEVFANNLEQRFTPYNYAPESLCRQV 421
Query: 766 ATSISDDDNCFLT-AAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAG 824
+ L +A TLEE K + K PG D + + + +
Sbjct: 422 EEYLESPFQMSLPLSAVTLEEVKNLIAKLPLKKAPGEDLLDNRTIRLLPDQALQFLALIF 481
Query: 825 CEWLENGVFPPHLNSTNIALIPK-GDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLD 883
L+ G FP S +I +I K G T + +RP +L + K++ +++ NRL D
Sbjct: 482 NSVLDVGYFPKAWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTCKD 541
Query: 884 --KCISTSQSAF-----VPGR--SILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAY 934
K I Q F P + +++ AL A+E Y V LDI +A+
Sbjct: 542 VTKAIPKFQFGFRLQHGTPEQLHRVVNFALEAMENKEYA----------VGAFLDIQQAF 591
Query: 935 DRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSP 994
DR+ W + F + + +E + V V+G + G+ QG L P
Sbjct: 592 DRV-WHPGLLYKAKRLFPPQLYLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGP 650
Query: 995 YLFILCAEGL 1004
L+ + A +
Sbjct: 651 TLYSVFASDM 660
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 47.4 bits (111), Expect = 3e-04
Identities = 64/258 (24%), Positives = 112/258 (42%), Gaps = 20/258 (7%)
Query: 782 TLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCE-WLENGVFPPHLNST 840
+L E K+A S + K GPDG P WN + ++ G P + +
Sbjct: 160 SLIEIKSARASNE--KGAGPDGVTP----RSWNALDDRYKRLLYNIFVFYGRVPSPIKGS 213
Query: 841 NIALIPKGDAQTTMKDWRPIALCNVVYKMVAKVLANRLKNVLDKCISTSQSAFVPGRSIL 900
PK + +RP+++C+V+ + K+LA R V Q+A++P +
Sbjct: 214 RTVFTPKIEGGPDPGVFRPLSICSVILREFNKILARRF--VSCYTYDERQTAYLP----I 267
Query: 901 DNALVAIELIHYMKAKTKGSQGDVALK-LDISKAYDRLDWEYLRDIMIQMRFSEKWIRWI 959
D + + ++ + A+ K + ++ + LD+ KA++ + L D + + + +I
Sbjct: 268 DGVCINVSMLTAIIAEAKRLRKELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDYI 327
Query: 960 MLCVETVDYTVLVNG-VQVGPLIPGRGIRQGDPLSPYLFILCAE-GLSALIRDAERRGVI 1017
V + G ++ ++ G+ QGDPLS LF L E L AL + E R I
Sbjct: 328 ADMYNNVITEMQFEGKCELASIL--AGVYQGDPLSGPLFTLAYEKALRAL--NNEGRFDI 383
Query: 1018 TGTRICTSAPAISHLLFA 1035
R+ SA + LL A
Sbjct: 384 ADVRVNASAYSDDGLLLA 401
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 44.7 bits (104), Expect = 0.002
Identities = 28/96 (29%), Positives = 49/96 (50%), Gaps = 1/96 (1%)
Query: 911 HYMKAKTKGSQGDVALKLDISKAYDRLDWEYLRDIMIQMRFSEKWIRWIMLCVETVDYTV 970
++ K +G+Q DV LD SKA+D++ + L D + ++ ++ IRW+ + + + V
Sbjct: 7 NWTKNIDEGAQTDVKY-LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKV 65
Query: 971 LVNGVQVGPLIPGRGIRQGDPLSPYLFILCAEGLSA 1006
V P G+ QG +SP LF + +SA
Sbjct: 66 KVGNTLSEPKKTVCGVPQGSVISPVLFGIFVNEISA 101
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 42.0 bits (97), Expect = 0.015
Identities = 80/380 (21%), Positives = 140/380 (36%), Gaps = 30/380 (7%)
Query: 652 KVQKKIEKLRTHVTAANVSYFNS-LKNKLDKLLVQDDLFWKQRAKTFWYKEGDLNTKFFH 710
++++K+ + A + +S L N+L K+L + +G N +
Sbjct: 307 RLKRKVRREYARTGDARIQQIHSRLANRLHKVLNRRKQSQIDNLLENLDTDGSTNFSLWR 366
Query: 711 AAATSRRKVNRIEHLENSNGTVCRKEDELQNIARNYFSHLFTKLSSTRADVISKVATSIS 770
+ + + N G CR E + N+ F L+ VA S+
Sbjct: 367 ITKRYKTQATPNSAIRNPAGGWCRTSREKTEVFANHLEQRFKALAFAPESHSLMVAESLQ 426
Query: 771 DDDNCFLTA-AFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLE 829
L A TLEE K ++ K PG D + + + + L
Sbjct: 427 TPFQMALPADPVTLEEVKELVSKLKPKKAPGEDLLDNRTIRLLPDQALLYLVLIFNSILR 486
Query: 830 NGVFPPHLNSTNIALIPK-GDAQTTMKDWRPIALCNVVYKMVAKVLANRL--KNVLDKCI 886
G FP + +I +I K G + +RP +L + KM+ +++ NR+ + + I
Sbjct: 487 VGYFPKARPTASIIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRILTSEEVTRAI 546
Query: 887 STSQSAF-----VPGR--SILDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDW 939
Q F P + +++ AL A+E K + GS LDI +A+DR+ W
Sbjct: 547 PKFQFGFRLQHGTPEQLHRVVNFALEALE-----KKEYAGS-----CFLDIQQAFDRV-W 595
Query: 940 EYLRDIMIQMRFSEKWIRWIMLCVETVDYTVLVNGVQVGPLIPGRGIRQGDPLSPYLFIL 999
+ S + + I E ++V +G + G+ QG L P L+
Sbjct: 596 HPGLLYKAKSLLSPQLFQLIKSFWEGRKFSVTADGCRSSVKFIEAGVPQGSVLGPTLY-- 653
Query: 1000 CAEGLSALIRDAERRGVITG 1019
S D + +TG
Sbjct: 654 -----SIFTADMPNQNAVTG 668
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 39.3 bits (90), Expect = 0.094
Identities = 38/181 (20%), Positives = 72/181 (38%), Gaps = 16/181 (8%)
Query: 763 SKVATSISDDDNCFLTAAFTLEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHK 822
S+ T+I+++ + A + E T +++ + PG DG N + W +
Sbjct: 420 SEAPTAIAEE----VPPALEVFEVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLAS 475
Query: 823 AGCEWLENGVFPPHLNSTNIALIPKGDAQTTMK--DWRPIALCNVVYKMVAKVLANRLKN 880
+ G FP + + KG + + +R I L V K++ ++ NR++
Sbjct: 476 LFSRCIRLGYFPAEWKCPRVVSLLKGPDKDKCEPSSYRGICLLPVFGKVLEAIMVNRVRE 535
Query: 881 VLDKCISTSQSAFVPGRSILDNALVAIELIHYMKAKTKGSQGDVALK--LDISKAYDRLD 938
VL + Q F GR + D ++K+ S L +D A+D ++
Sbjct: 536 VLPEGCRW-QFGFRQGRCVED-------AWRHVKSSVGASAAQYVLGTFVDFKGAFDNVE 587
Query: 939 W 939
W
Sbjct: 588 W 588
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 37.4 bits (85), Expect = 0.36
Identities = 34/160 (21%), Positives = 64/160 (39%), Gaps = 8/160 (5%)
Query: 783 LEEFKTAAFSMQADKCPGPDGFNPGFYQNFWNMCGSEIHKAGCEWLENGVFPPHLNSTNI 842
++E + + K PGPDG + W + + L FP ++
Sbjct: 407 MDEVSDSVRRCKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLYKQCLLESYFPQKWKIASL 466
Query: 843 ALIPK--GDAQTTMKDWRPIALCNVVYKMVAKVLANRL-KNVLDKCISTSQSAFVPGRSI 899
++ K ++ +RPI L + + K++ ++ RL + ++D +S Q AF G+S
Sbjct: 467 VILLKLLDRIRSDPGSYRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKST 526
Query: 900 LDNALVAIELIHYMKAKTKGSQGDVALKLDISKAYDRLDW 939
D + + K + L +D A+D L W
Sbjct: 527 EDAWRCVQRHVECSEMKYV-----IGLNIDFQGAFDNLGW 561
>SYE_HORVU (Q43768) Glutamyl-tRNA synthetase (EC 6.1.1.17)
(Glutamate--tRNA ligase) (GluRS)
Length = 560
Score = 36.2 bits (82), Expect = 0.80
Identities = 43/149 (28%), Positives = 61/149 (40%), Gaps = 30/149 (20%)
Query: 410 FTVDRIGRGGGL----AFLWKNSANCTITNFSQNHIDVEVDDLLI----GKWRLTGFYGM 461
FT+DR+ + G + W N H+ DLLI +WR TG
Sbjct: 341 FTIDRVNKSGAVFDATKLKWMNG----------QHLRSLPSDLLIKDFEDQWRSTGILLE 390
Query: 462 PENGRRKESWAFLK---NLARTSSLPWCVIGDF--NDILSSEEKKG-----RSERAPWLI 511
E+G KE+ LK +L + C + + ++ LSS+E K SE A LI
Sbjct: 391 SESGFAKEAAELLKEGIDLITDADAALCKLLSYPLHETLSSDEAKSVVEDKLSEVASGLI 450
Query: 512 RGFRQAVLDAGLFDLHMTGYAFTWFKSLG 540
+ LD L + H G+ W KS G
Sbjct: 451 SAYDSGELDQALAEGH-DGWK-KWVKSFG 477
>SET4_YEAST (P42948) SET domain protein 4
Length = 560
Score = 34.3 bits (77), Expect = 3.0
Identities = 21/82 (25%), Positives = 45/82 (54%), Gaps = 3/82 (3%)
Query: 606 EPDFKQQVQQ-RWKLYPEEGITQKLSYCAEDLTDWSRNNNNFRRDISKVQKKIEKLRTHV 664
+ D K QV Q + ++P + ++L + +T + N N ++ ++ ++++ +K + H
Sbjct: 208 DSDTKVQVNQVKPMIFPRKMGDERLFQFSSIVTTSASNTNQHQQSVNNIEEQPKKRQLHY 267
Query: 665 TAANVSYFNSLKNKL--DKLLV 684
TA NS++ KL +KL+V
Sbjct: 268 TAPTTENSNSIRKKLRQEKLVV 289
>PMS1_HUMAN (P54277) PMS1 protein homolog 1 (DNA mismatch repair
protein PMS1)
Length = 932
Score = 33.9 bits (76), Expect = 4.0
Identities = 40/151 (26%), Positives = 65/151 (42%), Gaps = 12/151 (7%)
Query: 554 VNSDWCNMFPNA-GVECLSTTSSDHYPLWLTCKPVTA----VNRNPHRFKFENAWLAEPD 608
+N D CN N + T+ D + KP++A V + +F EN + D
Sbjct: 541 LNEDSCNKKSNVIDNKSGKVTAYDLLSNRVIKKPMSASALFVQDHRPQFLIENPKTSLED 600
Query: 609 FKQQVQQRWKLYPEEGITQKLSYCAEDLTDWSRNNNNFRRDI---SKVQKKIEKLRTHVT 665
Q+++ WK EE +KL Y + D R N+ +R I S++ K + + T
Sbjct: 601 ATLQIEELWKTLSEE---EKLKYEEKATKDLERYNSQMKRAIEQESQMSLKDGRKKIKPT 657
Query: 666 AA-NVSYFNSLKNKLDKLLVQDDLFWKQRAK 695
+A N++ + LK L D+L Q K
Sbjct: 658 SAWNLAQKHKLKTSLSNQPKLDELLQSQIEK 688
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.333 0.142 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 197,722,129
Number of Sequences: 164201
Number of extensions: 8248035
Number of successful extensions: 27786
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 27727
Number of HSP's gapped (non-prelim): 46
length of query: 1711
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1587
effective length of database: 39,613,130
effective search space: 62866037310
effective search space used: 62866037310
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 73 (32.7 bits)
Medicago: description of AC146755.8


