

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
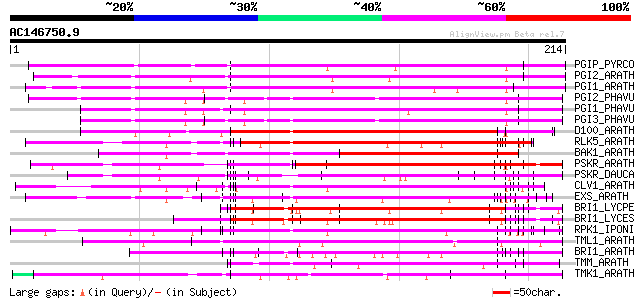
Query= AC146750.9 + phase: 0
(214 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor (Polyg... 133 3e-31
PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor (Pol... 130 2e-30
PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor (Pol... 126 3e-29
PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor (Pol... 100 2e-21
PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor (Pol... 100 2e-21
PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor (Pol... 100 3e-21
D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100 ... 97 4e-20
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 93 5e-19
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 92 1e-18
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 87 3e-17
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 84 3e-16
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 84 3e-16
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 83 4e-16
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 83 5e-16
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 83 5e-16
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 78 1e-14
TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor 76 7e-14
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 74 2e-13
TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM) 68 1e-11
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 62 1e-09
>PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor
(Polygalacturonase-inhibiting protein)
Length = 330
Score = 133 bits (335), Expect = 3e-31
Identities = 78/191 (40%), Positives = 109/191 (56%), Gaps = 3/191 (1%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGC 67
T L L+ ++ S N + CN DDK LL+I+ FG P L W ++T+CCDW V C
Sbjct: 7 TFLSLTLLFSSVLNPALSDLCNPDDKKVLLQIKKAFGDPYV-LASWKSDTDCCDWYCVTC 65
Query: 68 GRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
R+ +TI G +SG +PA G+LPYL L + P +TGPI + +KL+ L++
Sbjct: 66 DST-TNRINSLTIFAGQ-VSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKS 123
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L L +LSG +P FL +LK L +DLS N L+G IP+SL L +L + N+L G I
Sbjct: 124 LRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNKLTGHI 183
Query: 188 PAGLKKFNKNV 198
P +F NV
Sbjct: 184 PISFGQFIGNV 194
Score = 75.1 bits (183), Expect = 1e-13
Identities = 54/178 (30%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLG 144
L+G +P+ LP L L L + K+TG IP SF + + + +L L N LSG IP+
Sbjct: 155 LTGAIPSSLSELPNLGALRL-DRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFA 213
Query: 145 KLK----------------------------------------------RLKEVDLSNNK 158
++ L +D+++NK
Sbjct: 214 QMDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRNLLEFNLSKVEFPTSLTSLDINHNK 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
+ G+IP L + NVS+N+LCG IP G L+ F++ + HN+CLCGAPL CK
Sbjct: 274 IYGSIPVEFTQL-NFQFLNVSYNRLCGQIPVGGKLQSFDEYSYFHNRCLCGAPLPSCK 330
>PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 330
Score = 130 bits (327), Expect = 2e-30
Identities = 77/190 (40%), Positives = 108/190 (56%), Gaps = 5/190 (2%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCG- 68
L LS++ + S A D C+ DDK LLKI+ P L WD T+CC W + CG
Sbjct: 9 LLLSTLLLTTSLAKD--LCHKDDKTTLLKIKKSLNNPY-HLASWDPKTDCCSWYCLECGD 65
Query: 69 RPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
RVT + I G +SG +P E G+LPYL+ L ++ +TG I + +KL+ L L
Sbjct: 66 ATVNHRVTSLIIQDG-EISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLTFL 124
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L +L+GP+P FL +LK L+ +DLS N LSG+IP+SL +L+ L +S N+L G IP
Sbjct: 125 RLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIP 184
Query: 189 AGLKKFNKNV 198
F+ V
Sbjct: 185 ESFGTFSGKV 194
Score = 82.8 bits (203), Expect = 5e-16
Identities = 58/178 (32%), Positives = 80/178 (44%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ +L L L L+ K+TGPIP SF ++ +L L N LSG IP LG
Sbjct: 155 LSGSIPSSLSSLRKLEYLELSRN-KLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLG 213
Query: 145 K----------------------------------------------LKRLKEVDLSNNK 158
K L +D+++N
Sbjct: 214 NPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLNNLDMNHNG 273
Query: 159 LSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
++G+IPA NVS+N+LCG IP G +++F+ F HNKCLCGAPL CK
Sbjct: 274 ITGSIPAEWSKAY-FQLLNVSYNRLCGRIPKGEYIQRFDSYSFFHNKCLCGAPLPSCK 330
>PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 330
Score = 126 bits (317), Expect = 3e-29
Identities = 78/189 (41%), Positives = 108/189 (56%), Gaps = 6/189 (3%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG 66
L LLFL + F + S D CN +DK LLKI+ P L WD T+CC W +
Sbjct: 7 LCLLFLFT--FLTTCLSKDL-CNQNDKNTLLKIKKSLNNPY-HLASWDPQTDCCSWYCLE 62
Query: 67 CGRPYPG-RVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRL 125
CG RVT +TI G +SG +PAE G+LPYL L ++ +TG I + +KL+ L
Sbjct: 63 CGDATVNHRVTALTIFSGQ-ISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLKNL 121
Query: 126 QNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
+ L L +L+GPIP F+ +LK L+ ++LS N LSG+IP+SL L + +S N+L G
Sbjct: 122 RMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKLTG 181
Query: 186 AIPAGLKKF 194
+IP F
Sbjct: 182 SIPESFGSF 190
Score = 86.7 bits (213), Expect = 4e-17
Identities = 64/178 (35%), Positives = 80/178 (43%), Gaps = 51/178 (28%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ-RLQNLDLGSNSLSGPIPSFLG 144
LSG++P+ LP + L L+ K+TG IP SF + +L L N LSGPIP LG
Sbjct: 155 LSGSIPSSLSTLPKILALELSRN-KLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLG 213
Query: 145 KLKRLKEVDLSNNKLSGT-----------------------------IPASLGNLQ---- 171
+ +DLS NKL G IP +LG L
Sbjct: 214 NID-FNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDISKVDIPKTLGILDLNHN 272
Query: 172 -------------SLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPCK 214
L FNVS+N+LCG IP G L+ F+ + HNKCLCGAPL CK
Sbjct: 273 GITGNIPVQWTEAPLQFFNVSYNKLCGHIPTGGKLQTFDSYSYFHNKCLCGAPLEICK 330
>PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 342
Score = 100 bits (250), Expect = 2e-21
Identities = 68/194 (35%), Positives = 102/194 (52%), Gaps = 8/194 (4%)
Query: 8 TLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG- 66
+L + I S S A + CN DK ALL+I+ G P L W T+CC+ +++G
Sbjct: 13 SLSIILVILVSLSTAHSEL-CNPQDKQALLQIKKDLGNPT-TLSSWLPTTDCCNRTWLGV 70
Query: 67 -CGRPYPG-RVTVVTISRGWGLSGT--LPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL 122
C RV + +S G L +P+ NLPYL+ L + + + GPIP + +KL
Sbjct: 71 LCDTDTQTYRVNNLDLS-GLNLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKL 129
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
+L L + ++SG IP FL ++K L +D S N LSGT+P S+ +L +L N+
Sbjct: 130 TQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGNR 189
Query: 183 LCGAIPAGLKKFNK 196
+ GAIP F+K
Sbjct: 190 ISGAIPDSYGSFSK 203
Score = 87.0 bits (214), Expect = 3e-17
Identities = 55/140 (39%), Positives = 79/140 (56%), Gaps = 6/140 (4%)
Query: 76 TVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSL 135
T +TISR L+G +P F NL L+ + L+ + G F + Q + L NSL
Sbjct: 206 TSMTISRN-RLTGKIPPTFANLN-LAFVDLSRN-MLEGDASVLFGSDKNTQKIHLAKNSL 262
Query: 136 SGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKK 193
+ + +G K L +DL NN++ GT+P L L+ L NVSFN LCG IP G L++
Sbjct: 263 AFDLGK-VGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEIPQGGNLQR 321
Query: 194 FNKNVFEHNKCLCGAPLAPC 213
F+ + + +NKCLCG+PL C
Sbjct: 322 FDVSAYANNKCLCGSPLPAC 341
>PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 342
Score = 100 bits (250), Expect = 2e-21
Identities = 63/174 (36%), Positives = 94/174 (53%), Gaps = 7/174 (4%)
Query: 28 CNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG--CGRPYPG-RVTVVTISRGW 84
CN DK ALL+I+ G P L W T+CC+ +++G C RV + +S G
Sbjct: 32 CNPQDKQALLQIKKDLGNPT-TLSSWLPTTDCCNRTWLGVLCDTDTQTYRVNNLDLS-GH 89
Query: 85 GLSGT--LPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
L +P+ NLPYL+ L + + + GPIP + +KL +L L + ++SG IP F
Sbjct: 90 NLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDF 149
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L ++K L +D S N LSGT+P S+ +L +L N++ GAIP F+K
Sbjct: 150 LSQIKTLVTLDFSYNALSGTLPPSISSLPNLGGITFDGNRISGAIPDSYGSFSK 203
Score = 88.2 bits (217), Expect = 1e-17
Identities = 53/153 (34%), Positives = 74/153 (47%), Gaps = 26/153 (16%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SG +P +G+ L ++TG IP +F+ L L +DL N L G G
Sbjct: 190 ISGAIPDSYGSFSKLFTAMTISRNRLTGKIPPTFANLN-LAFVDLSRNMLEGDASVLFGS 248
Query: 146 LKRLKEV-----------------------DLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
K K++ DL NN++ GT+P L L+ L NVSFN
Sbjct: 249 DKNTKKIHLAKNSLAFDLGKVGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLQSLNVSFNN 308
Query: 183 LCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPC 213
LCG IP G LK+F+ + + +NKCLCG+PL C
Sbjct: 309 LCGEIPQGGNLKRFDVSSYANNKCLCGSPLPSC 341
>PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2) (PGIP-3)
Length = 342
Score = 100 bits (248), Expect = 3e-21
Identities = 63/174 (36%), Positives = 94/174 (53%), Gaps = 7/174 (4%)
Query: 28 CNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVG--CGRPYPG-RVTVVTISRGW 84
CN DK ALL+I+ G P L W T+CC+ +++G C RV + +S G
Sbjct: 32 CNPQDKQALLQIKKDLGNPT-TLSSWLPTTDCCNRTWLGVLCDTDTQTYRVNNLDLS-GH 89
Query: 85 GLSGT--LPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
L +P+ NLPYL+ L + + + GPIP + +KL +L L + ++SG IP F
Sbjct: 90 NLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDF 149
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L ++K L +D S N LSGT+P S+ +L +L N++ GAIP F+K
Sbjct: 150 LSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGNRISGAIPDSYGSFSK 203
Score = 87.0 bits (214), Expect = 3e-17
Identities = 55/140 (39%), Positives = 78/140 (55%), Gaps = 6/140 (4%)
Query: 76 TVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSL 135
T +TISR L+G +P F NL L+ + L+ + G F + Q + L NSL
Sbjct: 206 TSMTISRN-RLTGKIPPTFANLN-LAFVDLSRN-MLQGDASVLFGSDKNTQKIHLAKNSL 262
Query: 136 SGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKK 193
+ +G K L +DL NN++ GT+P L L+ L NVSFN LCG IP G L++
Sbjct: 263 DFDLEK-VGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEIPQGGNLQR 321
Query: 194 FNKNVFEHNKCLCGAPLAPC 213
F+ + + +NKCLCG+PL C
Sbjct: 322 FDVSAYANNKCLCGSPLPAC 341
>D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100
precursor
Length = 372
Score = 96.7 bits (239), Expect = 4e-20
Identities = 63/199 (31%), Positives = 100/199 (49%), Gaps = 18/199 (9%)
Query: 28 CNADDKAALLKIRDHFGGPK-GRLDDWDNNTECC-DWSFVGCGRPYPGRVTVVTI----- 80
C+ D+ AL + P G + W NT+CC +W + C P GRVT +++
Sbjct: 27 CSPKDQTALNAFKSSLSEPNLGIFNTWSENTDCCKEWYGISCD-PDSGRVTDISLRGESE 85
Query: 81 -------SRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN 133
R +SG++ +L L+ L LA+ +TG IP + L L+ LDL N
Sbjct: 86 DAIFQKAGRSGYMSGSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGN 145
Query: 134 SLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA---G 190
++G IP+ +GKL +L ++L+ N++SG IPASL +L L ++ N + G IPA
Sbjct: 146 KITGEIPAEIGKLSKLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADFGS 205
Query: 191 LKKFNKNVFEHNKCLCGAP 209
LK ++ + N+ P
Sbjct: 206 LKMLSRVLLGRNELTGSIP 224
Score = 90.9 bits (224), Expect = 2e-18
Identities = 50/127 (39%), Positives = 68/127 (53%), Gaps = 3/127 (2%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+ G +P GN+ LS+L+L + +TGPIP S L +L N+L G IP G
Sbjct: 243 IEGPIPEWMGNMKVLSLLNL-DCNSLTGPIPGSLLSNSGLDVANLSRNALEGTIPDVFGS 301
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNK 203
L +DLS+N LSG IP SL + + + ++S N+LCG IP G F N+
Sbjct: 302 KTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRIPTGFPFDHLEATSFSDNQ 361
Query: 204 CLCGAPL 210
CLCG PL
Sbjct: 362 CLCGGPL 368
Score = 77.8 bits (190), Expect = 2e-14
Identities = 41/103 (39%), Positives = 67/103 (64%), Gaps = 1/103 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
++G +PAE G L L++L+LAE +++G IP S + L L++L+L N ++G IP+ G
Sbjct: 147 ITGEIPAEIGKLSKLAVLNLAEN-QMSGEIPASLTSLIELKHLELTENGITGVIPADFGS 205
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
LK L V L N+L+G+IP S+ ++ L+ ++S N + G IP
Sbjct: 206 LKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHIEGPIP 248
Score = 77.4 bits (189), Expect = 2e-14
Identities = 43/106 (40%), Positives = 60/106 (56%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
+SG +PA +L L L L E +TG IP F L+ L + LG N L+G IP +
Sbjct: 171 MSGEIPASLTSLIELKHLELTENG-ITGVIPADFGSLKMLSRVLLGRNELTGSIPESISG 229
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
++RL ++DLS N + G IP +GN++ LS N+ N L G IP L
Sbjct: 230 MERLADLDLSKNHIEGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSL 275
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 92.8 bits (229), Expect = 5e-19
Identities = 50/115 (43%), Positives = 74/115 (63%), Gaps = 4/115 (3%)
Query: 90 LPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRL 149
+P++ GNL L +L LA + GPIP S S+L L NLDL N L+G IPS++ +LK +
Sbjct: 204 IPSQLGNLTELQVLWLAGC-NLVGPIPPSLSRLTSLVNLDLTFNQLTGSIPSWITQLKTV 262
Query: 150 KEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN---KNVFEH 201
++++L NN SG +P S+GN+ +L +F+ S N+L G IP L N N+FE+
Sbjct: 263 EQIELFNNSFSGELPESMGNMTTLKRFDASMNKLTGKIPDNLNLLNLESLNLFEN 317
Score = 80.9 bits (198), Expect = 2e-15
Identities = 49/112 (43%), Positives = 65/112 (57%), Gaps = 1/112 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG++P E G+L + +S AE +G IP S KL++L LDL N LSG IP L
Sbjct: 464 SGSIPNEIGSLNGIIEISGAEND-FSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGW 522
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV 198
K L E++L+NN LSG IP +G L L+ ++S NQ G IP L+ NV
Sbjct: 523 KNLNELNLANNHLSGEIPKEVGILPVLNYLDLSSNQFSGEIPLELQNLKLNV 574
Score = 77.4 bits (189), Expect = 2e-14
Identities = 48/117 (41%), Positives = 67/117 (57%), Gaps = 2/117 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P F LP LS+L L++ TG IP + + L NL + N SG IP+ +G
Sbjct: 415 LSGQIPHGFWGLPRLSLLELSDN-SFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNEIGS 473
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHN 202
L + E+ + N SG IP SL L+ LS+ ++S NQL G IP L+ + KN+ E N
Sbjct: 474 LNGIIEISGAENDFSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGW-KNLNELN 529
Score = 74.7 bits (182), Expect = 1e-13
Identities = 41/105 (39%), Positives = 62/105 (59%), Gaps = 2/105 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG LP GN+ L A M K+TG IP++ + L L++L+L N L GP+P + +
Sbjct: 273 SGELPESMGNMTTLKRFD-ASMNKLTGKIPDNLNLLN-LESLNLFENMLEGPLPESITRS 330
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
K L E+ L NN+L+G +P+ LG L ++S+N+ G IPA +
Sbjct: 331 KTLSELKLFNNRLTGVLPSQLGANSPLQYVDLSYNRFSGEIPANV 375
Score = 70.9 bits (172), Expect = 2e-12
Identities = 61/188 (32%), Positives = 79/188 (41%), Gaps = 13/188 (6%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTEC--CDWSF 64
+ LL LSS Y + + D K L P L W +N + C W
Sbjct: 6 ILLLCLSSTYLPSLSLNQDATILRQAKLGL-------SDPAQSLSSWSDNNDVTPCKWLG 58
Query: 65 VGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
V C V V +S + L G P+ +LP L LSL + F
Sbjct: 59 VSCDAT--SNVVSVDLS-SFMLVGPFPSILCHLPSLHSLSLYNNSINGSLSADDFDTCHN 115
Query: 125 LQNLDLGSNSLSGPIPSFLG-KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQL 183
L +LDL N L G IP L L LK +++S N LS TIP+S G + L N++ N L
Sbjct: 116 LISLDLSENLLVGSIPKSLPFNLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAGNFL 175
Query: 184 CGAIPAGL 191
G IPA L
Sbjct: 176 SGTIPASL 183
Score = 63.9 bits (154), Expect = 3e-10
Identities = 44/130 (33%), Positives = 66/130 (49%), Gaps = 27/130 (20%)
Query: 86 LSGTLPAEFG-NLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
L G++P NLP L L ++ ++ IP+SF + ++L++L+L N LSG IP+ LG
Sbjct: 126 LVGSIPKSLPFNLPNLKFLEISGN-NLSDTIPSSFGEFRKLESLNLAGNFLSGTIPASLG 184
Query: 145 KLKRLKEVDLSNN-------------------------KLSGTIPASLGNLQSLSQFNVS 179
+ LKE+ L+ N L G IP SL L SL +++
Sbjct: 185 NVTTLKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLT 244
Query: 180 FNQLCGAIPA 189
FNQL G+IP+
Sbjct: 245 FNQLTGSIPS 254
Score = 63.2 bits (152), Expect = 4e-10
Identities = 42/103 (40%), Positives = 55/103 (52%), Gaps = 1/103 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G LP++ G L + L+ + +G IP + +L+ L L NS SG I + LGK
Sbjct: 343 LTGVLPSQLGANSPLQYVDLS-YNRFSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGK 401
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
K L V LSNNKLSG IP L LS +S N G+IP
Sbjct: 402 CKSLTRVRLSNNKLSGQIPHGFWGLPRLSLLELSDNSFTGSIP 444
Score = 62.4 bits (150), Expect = 8e-10
Identities = 36/110 (32%), Positives = 57/110 (51%), Gaps = 1/110 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG + G L+ + L+ K++G IP+ F L RL L+L NS +G IP +
Sbjct: 392 SGEISNNLGKCKSLTRVRLSNN-KLSGQIPHGFWGLPRLSLLELSDNSFTGSIPKTIIGA 450
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
K L + +S N+ SG+IP +G+L + + + + N G IP L K +
Sbjct: 451 KNLSNLRISKNRFSGSIPNEIGSLNGIIEISGAENDFSGEIPESLVKLKQ 500
Score = 59.3 bits (142), Expect = 6e-09
Identities = 45/129 (34%), Positives = 64/129 (48%), Gaps = 26/129 (20%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P NL L L+L E + GP+P S ++ + L L L +N L+G +PS LG
Sbjct: 296 LTGKIPDNL-NLLNLESLNLFEN-MLEGPLPESITRSKTLSELKLFNNRLTGVLPSQLGA 353
Query: 146 LKRLKEVDLSNNKLSGTIPA------------------------SLGNLQSLSQFNVSFN 181
L+ VDLS N+ SG IPA +LG +SL++ +S N
Sbjct: 354 NSPLQYVDLSYNRFSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRLSNN 413
Query: 182 QLCGAIPAG 190
+L G IP G
Sbjct: 414 KLSGQIPHG 422
Score = 37.4 bits (85), Expect = 0.026
Identities = 31/84 (36%), Positives = 35/84 (40%), Gaps = 26/84 (30%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E G LP L+ LDL SN SG IP L
Sbjct: 535 LSGEIPKEVGILPVLNY-------------------------LDLSSNQFSGEIPLELQN 569
Query: 146 LKRLKEVDLSNNKLSGTIPASLGN 169
LK L ++LS N LSG IP N
Sbjct: 570 LK-LNVLNLSYNHLSGKIPPLYAN 592
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 91.7 bits (226), Expect = 1e-18
Identities = 62/155 (40%), Positives = 79/155 (50%), Gaps = 5/155 (3%)
Query: 35 ALLKIRDHFGGPKGRLDDWDNNTEC-CDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAE 93
AL +++ P L WD C W V C VT V + LSG L +
Sbjct: 31 ALSALKNSLADPNKVLQSWDATLVTPCTWFHVTCNSD--NSVTRVDLGNA-NLSGQLVMQ 87
Query: 94 FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVD 153
G LP L L L +TG IP L L +LDL N+LSGPIPS LG+LK+L+ +
Sbjct: 88 LGQLPNLQYLELYSN-NITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRLKKLRFLR 146
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
L+NN LSG IP SL + +L ++S N L G IP
Sbjct: 147 LNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIP 181
Score = 55.8 bits (133), Expect = 7e-08
Identities = 28/69 (40%), Positives = 43/69 (61%)
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
+DLG+ +LSG + LG+L L+ ++L +N ++GTIP LGNL L ++ N L G I
Sbjct: 73 VDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPI 132
Query: 188 PAGLKKFNK 196
P+ L + K
Sbjct: 133 PSTLGRLKK 141
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 87.0 bits (214), Expect = 3e-17
Identities = 49/105 (46%), Positives = 66/105 (62%), Gaps = 3/105 (2%)
Query: 111 VTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNL 170
++GPI F L++L DL N+LSG IPS L + L+ +DLSNN+LSG+IP SL L
Sbjct: 535 LSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSIPVSLQQL 594
Query: 171 QSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCLCGAPLAPC 213
LS+F+V++N L G IP+G + F + FE N LCG PC
Sbjct: 595 SFLSKFSVAYNNLSGVIPSGGQFQTFPNSSFESNH-LCGEHRFPC 638
Score = 75.1 bits (183), Expect = 1e-13
Identities = 55/186 (29%), Positives = 89/186 (47%), Gaps = 35/186 (18%)
Query: 9 LLFLSSI--YFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN---TECCDWS 63
++FL+ + +F S + C+ D AL RD + + D W N+ T+CC+W+
Sbjct: 10 VIFLTELLCFFYSSESQTTSRCHPHDLEAL---RDFIAHLEPKPDGWINSSSSTDCCNWT 66
Query: 64 FVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ 123
+ C GRV + E GN K++G + S KL
Sbjct: 67 GITCNSNNTGRV--------------IRLELGN------------KKLSGKLSESLGKLD 100
Query: 124 RLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQL 183
++ L+L N + IP + LK L+ +DLS+N LSG IP S+ NL +L F++S N+
Sbjct: 101 EIRVLNLSRNFIKDSIPLSIFNLKNLQTLDLSSNDLSGGIPTSI-NLPALQSFDLSSNKF 159
Query: 184 CGAIPA 189
G++P+
Sbjct: 160 NGSLPS 165
Score = 67.0 bits (162), Expect = 3e-11
Identities = 36/84 (42%), Positives = 50/84 (58%)
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGN 169
++TG +P S LQ LDL N L+G IPS++G K L +DLSNN +G IP SL
Sbjct: 426 RLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSFTGEIPKSLTK 485
Query: 170 LQSLSQFNVSFNQLCGAIPAGLKK 193
L+SL+ N+S N+ P +K+
Sbjct: 486 LESLTSRNISVNEPSPDFPFFMKR 509
Score = 63.5 bits (153), Expect = 3e-10
Identities = 39/115 (33%), Positives = 63/115 (53%), Gaps = 2/115 (1%)
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
G G +P N P L++L+L ++G + + + + L +LDLG+N +G +P L
Sbjct: 279 GFIGGIPKSLANSPSLNLLNLRNN-SLSGRLMLNCTAMIALNSLDLGTNRFNGRLPENLP 337
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA-GLKKFNKNV 198
KRLK V+L+ N G +P S N +SLS F++S + L A G+ + KN+
Sbjct: 338 DCKRLKNVNLARNTFHGQVPESFKNFESLSYFSLSNSSLANISSALGILQHCKNL 392
Score = 51.6 bits (122), Expect = 1e-06
Identities = 33/102 (32%), Positives = 50/102 (48%), Gaps = 1/102 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P + +L L++L + E +++G + L L LD+ N SG IP +
Sbjct: 208 LTGNIPEDLFHLKRLNLLGIQEN-RLSGSLSREIRNLSSLVRLDVSWNLFSGEIPDVFDE 266
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L +LK N G IP SL N SL+ N+ N L G +
Sbjct: 267 LPQLKFFLGQTNGFIGGIPKSLANSPSLNLLNLRNNSLSGRL 308
Score = 49.3 bits (116), Expect = 7e-06
Identities = 36/105 (34%), Positives = 45/105 (42%), Gaps = 1/105 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G + FG L L L M +TG IP L+RL L + N LSG + + L
Sbjct: 185 AGNFTSGFGKCVLLEHLCLG-MNDLTGNIPEDLFHLKRLNLLGIQENRLSGSLSREIRNL 243
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L +D+S N SG IP L L F N G IP L
Sbjct: 244 SSLVRLDVSWNLFSGEIPDVFDELPQLKFFLGQTNGFIGGIPKSL 288
Score = 46.6 bits (109), Expect = 4e-05
Identities = 24/71 (33%), Positives = 43/71 (59%)
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
++L+ L + + L+G +P +L L+ +DLS N+L+G IP+ +G+ ++L ++S N
Sbjct: 415 EKLKVLVVANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNS 474
Query: 183 LCGAIPAGLKK 193
G IP L K
Sbjct: 475 FTGEIPKSLTK 485
Score = 45.1 bits (105), Expect = 1e-04
Identities = 32/106 (30%), Positives = 52/106 (48%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG+L E NL L L ++ +G IP+ F +L +L+ +N G IP L
Sbjct: 232 LSGSLSREIRNLSSLVRLDVS-WNLFSGEIPDVFDELPQLKFFLGQTNGFIGGIPKSLAN 290
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L ++L NN LSG + + + +L+ ++ N+ G +P L
Sbjct: 291 SPSLNLLNLRNNSLSGRLMLNCTAMIALNSLDLGTNRFNGRLPENL 336
Score = 40.4 bits (93), Expect = 0.003
Identities = 33/112 (29%), Positives = 56/112 (49%), Gaps = 8/112 (7%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLD-----LGSNSLSGPIPSF 142
G +P F N LS SL+ I ++ LQ +NL L + + P S
Sbjct: 354 GQVPESFKNFESLSYFSLSNSSLAN--ISSALGILQHCKNLTTLVLTLNFHGEALPDDSS 411
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKF 194
L ++LK + ++N +L+G++P L + L ++S+N+L GAIP+ + F
Sbjct: 412 L-HFEKLKVLVVANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDF 462
Score = 39.7 bits (91), Expect = 0.005
Identities = 46/167 (27%), Positives = 72/167 (42%), Gaps = 14/167 (8%)
Query: 12 LSSIYFSKSNASDDF--FCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCDWSFVGCGR 69
L+S S + S DF F ++ A L+ FG P ++ NN W G +
Sbjct: 489 LTSRNISVNEPSPDFPFFMKRNESARALQYNQIFGFPP-TIELGHNNLSGPIWEEFGNLK 547
Query: 70 PYPGRVTVVTISRGWG-LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNL 128
++ V + W LSG++P+ + L L L+ +++G IP S +L L
Sbjct: 548 ----KLHVFDLK--WNALSGSIPSSLSGMTSLEALDLSNN-RLSGSIPVSLQQLSFLSKF 600
Query: 129 DLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSG--TIPASLGNLQSL 173
+ N+LSG IPS G+ + +N L G P S G +L
Sbjct: 601 SVAYNNLSGVIPSG-GQFQTFPNSSFESNHLCGEHRFPCSEGTESAL 646
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 83.6 bits (205), Expect = 3e-16
Identities = 54/131 (41%), Positives = 75/131 (57%), Gaps = 8/131 (6%)
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
GL P+ F + LS SL G I F L++L L+L +N+LSG IP+ L
Sbjct: 525 GLQYNQPSSFPPMIDLSYNSL------NGSIWPEFGDLRQLHVLNLKNNNLSGNIPANLS 578
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL--KKFNKNVFEHN 202
+ L+ +DLS+N LSG IP SL L LS F+V++N+L G IP G+ + F + FE N
Sbjct: 579 GMTSLEVLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKLSGPIPTGVQFQTFPNSSFEGN 638
Query: 203 KCLCGAPLAPC 213
+ LCG +PC
Sbjct: 639 QGLCGEHASPC 649
Score = 72.8 bits (177), Expect = 6e-13
Identities = 59/208 (28%), Positives = 84/208 (40%), Gaps = 44/208 (21%)
Query: 23 SDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNN------TECCDWSFVGCGRPY----- 71
S + CN++D AL G + +D W N + CCDW + C
Sbjct: 24 SQNLTCNSNDLKAL---EGFMRGLESSIDGWKWNESSSFSSNCCDWVGISCKSSVSLGLD 80
Query: 72 ----PGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQN 127
GRV + + R LSG L L L +L+L ++G I S L L+
Sbjct: 81 DVNESGRVVELELGRR-KLSGKLSESVAKLDQLKVLNLTH-NSLSGSIAASLLNLSNLEV 138
Query: 128 LDLGSNSLSGPIPSFL------------------------GKLKRLKEVDLSNNKLSGTI 163
LDL SN SG PS + L R++E+DL+ N G+I
Sbjct: 139 LDLSSNDFSGLFPSLINLPSLRVLNVYENSFHGLIPASLCNNLPRIREIDLAMNYFDGSI 198
Query: 164 PASLGNLQSLSQFNVSFNQLCGAIPAGL 191
P +GN S+ ++ N L G+IP L
Sbjct: 199 PVGIGNCSSVEYLGLASNNLSGSIPQEL 226
Score = 67.0 bits (162), Expect = 3e-11
Identities = 41/110 (37%), Positives = 58/110 (52%), Gaps = 2/110 (1%)
Query: 88 GTLPAEF-GNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
G +PA NLP + + LA M G IP ++ L L SN+LSG IP L +L
Sbjct: 171 GLIPASLCNNLPRIREIDLA-MNYFDGSIPVGIGNCSSVEYLGLASNNLSGSIPQELFQL 229
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L + L NN+LSG + + LG L +L + ++S N+ G IP + NK
Sbjct: 230 SNLSVLALQNNRLSGALSSKLGKLSNLGRLDISSNKFSGKIPDVFLELNK 279
Score = 62.8 bits (151), Expect = 6e-10
Identities = 39/104 (37%), Positives = 52/104 (49%), Gaps = 1/104 (0%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G++P GN + L LA ++G IP +L L L L +N LSG + S LGKL
Sbjct: 196 GSIPVGIGNCSSVEYLGLASN-NLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLS 254
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L +D+S+NK SG IP L L F+ N G +P L
Sbjct: 255 NLGRLDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSL 298
Score = 61.2 bits (147), Expect = 2e-09
Identities = 36/102 (35%), Positives = 56/102 (54%), Gaps = 1/102 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG++P E L LS+L+L + +++G + + KL L LD+ SN SG IP +
Sbjct: 218 LSGSIPQELFQLSNLSVLAL-QNNRLSGALSSKLGKLSNLGRLDISSNKFSGKIPDVFLE 276
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
L +L +N +G +P SL N +S+S ++ N L G I
Sbjct: 277 LNKLWYFSAQSNLFNGEMPRSLSNSRSISLLSLRNNTLSGQI 318
Score = 57.8 bits (138), Expect = 2e-08
Identities = 37/81 (45%), Positives = 45/81 (54%), Gaps = 4/81 (4%)
Query: 93 EFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEV 152
+F NL L + S ++ G +P S LQ LDL N LSG IP +LG L L +
Sbjct: 423 QFKNLKVLIIASC----QLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYL 478
Query: 153 DLSNNKLSGTIPASLGNLQSL 173
DLSNN G IP SL +LQSL
Sbjct: 479 DLSNNTFIGEIPHSLTSLQSL 499
Score = 55.8 bits (133), Expect = 7e-08
Identities = 33/106 (31%), Positives = 55/106 (51%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG L ++ G L L L ++ K +G IP+ F +L +L SN +G +P L
Sbjct: 242 LSGALSSKLGKLSNLGRLDISSN-KFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSLSN 300
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+ + + L NN LSG I + + +L+ +++ N G+IP+ L
Sbjct: 301 SRSISLLSLRNNTLSGQIYLNCSAMTNLTSLDLASNSFSGSIPSNL 346
Score = 55.1 bits (131), Expect = 1e-07
Identities = 35/93 (37%), Positives = 49/93 (52%), Gaps = 1/93 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G +P N +S+LSL ++G I + S + L +LDL SNS SG IPS L
Sbjct: 291 NGEMPRSLSNSRSISLLSLRNNT-LSGQIYLNCSAMTNLTSLDLASNSFSGSIPSNLPNC 349
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVS 179
RLK ++ + K IP S N QSL+ + S
Sbjct: 350 LRLKTINFAKIKFIAQIPESFKNFQSLTSLSFS 382
Score = 52.0 bits (123), Expect = 1e-06
Identities = 31/82 (37%), Positives = 44/82 (52%)
Query: 121 KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSF 180
+ + L+ L + S L G +P +L L+ +DLS N+LSGTIP LG+L SL ++S
Sbjct: 423 QFKNLKVLIIASCQLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSN 482
Query: 181 NQLCGAIPAGLKKFNKNVFEHN 202
N G IP L V + N
Sbjct: 483 NTFIGEIPHSLTSLQSLVSKEN 504
Score = 51.2 bits (121), Expect = 2e-06
Identities = 35/108 (32%), Positives = 51/108 (46%), Gaps = 1/108 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P F L L S A+ G +P S S + + L L +N+LSG I +
Sbjct: 267 SGKIPDVFLELNKLWYFS-AQSNLFNGEMPRSLSNSRSISLLSLRNNTLSGQIYLNCSAM 325
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKF 194
L +DL++N SG+IP++L N L N + + IP K F
Sbjct: 326 TNLTSLDLASNSFSGSIPSNLPNCLRLKTINFAKIKFIAQIPESFKNF 373
Score = 50.4 bits (119), Expect = 3e-06
Identities = 44/142 (30%), Positives = 59/142 (40%), Gaps = 37/142 (26%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L GT+P N P L +L L+ +++G IP L L LDL +N+ G IP L
Sbjct: 437 LRGTVPQWLSNSPSLQLLDLS-WNQLSGTIPPWLGSLNSLFYLDLSNNTFIGEIPHSLTS 495
Query: 146 LKRL--KE----------------------------------VDLSNNKLSGTIPASLGN 169
L+ L KE +DLS N L+G+I G+
Sbjct: 496 LQSLVSKENAVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGD 555
Query: 170 LQSLSQFNVSFNQLCGAIPAGL 191
L+ L N+ N L G IPA L
Sbjct: 556 LRQLHVLNLKNNNLSGNIPANL 577
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 83.6 bits (205), Expect = 3e-16
Identities = 44/121 (36%), Positives = 71/121 (58%), Gaps = 3/121 (2%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P E L +LS ++ + +TG IP+S S+ L ++DL N ++G IP + +K
Sbjct: 494 GNIPREIFELKHLSRINTSAN-NITGGIPDSISRCSTLISVDLSRNRINGEIPKGINNVK 552
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNKCL 205
L +++S N+L+G+IP +GN+ SL+ ++SFN L G +P G FN+ F N L
Sbjct: 553 NLGTLNISGNQLTGSIPTGIGNMTSLTTLDLSFNDLSGRVPLGGQFLVFNETSFAGNTYL 612
Query: 206 C 206
C
Sbjct: 613 C 613
Score = 76.6 bits (187), Expect = 4e-14
Identities = 49/115 (42%), Positives = 66/115 (56%), Gaps = 2/115 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G +P EFG L L +L +A +TG IP S S L+ L L L N+L+G IP L L
Sbjct: 230 TGGVPPEFGGLTKLEILDMASCT-LTGEIPTSLSNLKHLHTLFLHINNLTGHIPPELSGL 288
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK-NVFE 200
LK +DLS N+L+G IP S NL +++ N+ N L G IP + + K VFE
Sbjct: 289 VSLKSLDLSINQLTGEIPQSFINLGNITLINLFRNNLYGQIPEAIGELPKLEVFE 343
Score = 73.2 bits (178), Expect = 4e-13
Identities = 46/124 (37%), Positives = 61/124 (49%), Gaps = 4/124 (3%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
G GLSG PA L L + + TG +P F L +L+ LD+ S +L+G IP+
Sbjct: 201 GAGLSGKSPAFLSRLKNLREMYIGYYNSYTGGVPPEFGGLTKLEILDMASCTLTGEIPTS 260
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK----NV 198
L LK L + L N L+G IP L L SL ++S NQL G IP N+
Sbjct: 261 LSNLKHLHTLFLHINNLTGHIPPELSGLVSLKSLDLSINQLTGEIPQSFINLGNITLINL 320
Query: 199 FEHN 202
F +N
Sbjct: 321 FRNN 324
Score = 67.4 bits (163), Expect = 2e-11
Identities = 65/224 (29%), Positives = 98/224 (43%), Gaps = 40/224 (17%)
Query: 3 LLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGR-LDDWDNNTEC-- 59
L LHL L F ++ D LL ++ GPKG L DW +++
Sbjct: 11 LFLHLYLFFSPCFAYT-------------DMEVLLNLKSSMIGPKGHGLHDWIHSSSPDA 57
Query: 60 -CDWSFVGC---GRPYPGRVTVV----TISRGWGL--------------SGTLPAEFGNL 97
C +S V C R V+ TIS G+ +G LP E +L
Sbjct: 58 HCSFSGVSCDDDARVISLNVSFTPLFGTISPEIGMLTHLVNLTLAANNFTGELPLEMKSL 117
Query: 98 PYLSMLSLAEMPKVTGPIPNSFSK-LQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSN 156
L +L+++ +TG P K + L+ LD +N+ +G +P + +LK+LK +
Sbjct: 118 TSLKVLNISNNGNLTGTFPGEILKAMVDLEVLDTYNNNFNGKLPPEMSELKKLKYLSFGG 177
Query: 157 NKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFE 200
N SG IP S G++QSL ++ L G PA L + KN+ E
Sbjct: 178 NFFSGEIPESYGDIQSLEYLGLNGAGLSGKSPAFLSRL-KNLRE 220
Score = 65.1 bits (157), Expect = 1e-10
Identities = 41/119 (34%), Positives = 63/119 (52%), Gaps = 2/119 (1%)
Query: 73 GRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGS 132
G +T++ + R L G +P G LP L + + E T +P + + L LD+
Sbjct: 313 GNITLINLFRN-NLYGQIPEAIGELPKLEVFEVWEN-NFTLQLPANLGRNGNLIKLDVSD 370
Query: 133 NSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
N L+G IP L + ++L+ + LSNN G IP LG +SL++ + N L G +PAGL
Sbjct: 371 NHLTGLIPKDLCRGEKLEMLILSNNFFFGPIPEELGKCKSLTKIRIVKNLLNGTVPAGL 429
Score = 63.5 bits (153), Expect = 3e-10
Identities = 39/111 (35%), Positives = 62/111 (55%), Gaps = 1/111 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P E L L L L+ + ++TG IP SF L + ++L N+L G IP +G+
Sbjct: 277 LTGHIPPELSGLVSLKSLDLS-INQLTGEIPQSFINLGNITLINLFRNNLYGQIPEAIGE 335
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L +L+ ++ N + +PA+LG +L + +VS N L G IP L + K
Sbjct: 336 LPKLEVFEVWENNFTLQLPANLGRNGNLIKLDVSDNHLTGLIPKDLCRGEK 386
Score = 59.3 bits (142), Expect = 6e-09
Identities = 35/108 (32%), Positives = 57/108 (52%), Gaps = 2/108 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+GT+PA NLP ++++ L + +G +P + S L + L +N SG IP +G
Sbjct: 421 LNGTVPAGLFNLPLVTIIELTDN-FFSGELPVTMSG-DVLDQIYLSNNWFSGEIPPAIGN 478
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L+ + L N+ G IP + L+ LS+ N S N + G IP + +
Sbjct: 479 FPNLQTLFLDRNRFRGNIPREIFELKHLSRINTSANNITGGIPDSISR 526
Score = 53.9 bits (128), Expect = 3e-07
Identities = 33/108 (30%), Positives = 59/108 (54%), Gaps = 1/108 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P F NL +++++L + G IP + +L +L+ ++ N+ + +P+ LG+
Sbjct: 301 LTGEIPQSFINLGNITLINLFRN-NLYGQIPEAIGELPKLEVFEVWENNFTLQLPANLGR 359
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L ++D+S+N L+G IP L + L +S N G IP L K
Sbjct: 360 NGNLIKLDVSDNHLTGLIPKDLCRGEKLEMLILSNNFFFGPIPEELGK 407
Score = 47.4 bits (111), Expect = 3e-05
Identities = 32/109 (29%), Positives = 52/109 (47%), Gaps = 2/109 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G +P + L ML L+ GPIP K + L + + N L+G +P+ L
Sbjct: 373 LTGLIPKDLCRGEKLEMLILSNN-FFFGPIPEELGKCKSLTKIRIVKNLLNGTVPAGLFN 431
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKF 194
L + ++L++N SG +P ++ L Q +S N G IP + F
Sbjct: 432 LPLVTIIELTDNFFSGELPVTMSG-DVLDQIYLSNNWFSGEIPPAIGNF 479
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells
protein) (EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 83.2 bits (204), Expect = 4e-16
Identities = 49/121 (40%), Positives = 69/121 (56%), Gaps = 4/121 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +PA L L++L L+ +TG IP +LQ L+L +N L+G IP G
Sbjct: 616 LSGEIPASLSRLTNLTILDLSGNA-LTGSIPKEMGNSLKLQGLNLANNQLNGHIPESFGL 674
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV---FEHN 202
L L +++L+ NKL G +PASLGNL+ L+ ++SFN L G + + L K V E N
Sbjct: 675 LGSLVKLNLTKNKLDGPVPASLGNLKELTHMDLSFNNLSGELSSELSTMEKLVGLYIEQN 734
Query: 203 K 203
K
Sbjct: 735 K 735
Score = 81.6 bits (200), Expect = 1e-15
Identities = 65/192 (33%), Positives = 96/192 (49%), Gaps = 12/192 (6%)
Query: 7 LTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTEC--CDWSF 64
LT LFL + S+A D + + +L+ + P L W+ ++ CDW
Sbjct: 4 LTALFLFLFFSFSSSAIVDL---SSETTSLISFKRSLENPS-LLSSWNVSSSASHCDWVG 59
Query: 65 VGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
V C GRV +++ L G +P E +L L L LA + +G IP L+
Sbjct: 60 VTC---LLGRVNSLSLP-SLSLRGQIPKEISSLKNLRELCLAGN-QFSGKIPPEIWNLKH 114
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLG-NLQSLSQFNVSFNQL 183
LQ LDL NSL+G +P L +L +L +DLS+N SG++P S +L +LS +VS N L
Sbjct: 115 LQTLDLSGNSLTGLLPRLLSELPQLLYLDLSDNHFSGSLPPSFFISLPALSSLDVSNNSL 174
Query: 184 CGAIPAGLKKFN 195
G IP + K +
Sbjct: 175 SGEIPPEIGKLS 186
Score = 77.0 bits (188), Expect = 3e-14
Identities = 46/117 (39%), Positives = 67/117 (56%), Gaps = 2/117 (1%)
Query: 75 VTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNS 134
+T++ +S G L+G++P E GN L L+LA ++ G IP SF L L L+L N
Sbjct: 630 LTILDLS-GNALTGSIPKEMGNSLKLQGLNLANN-QLNGHIPESFGLLGSLVKLNLTKNK 687
Query: 135 LSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L GP+P+ LG LK L +DLS N LSG + + L ++ L + N+ G IP+ L
Sbjct: 688 LDGPVPASLGNLKELTHMDLSFNNLSGELSSELSTMEKLVGLYIEQNKFTGEIPSEL 744
Score = 72.8 bits (177), Expect = 6e-13
Identities = 42/107 (39%), Positives = 57/107 (53%), Gaps = 1/107 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P+E GN+ L + A GP+P SKL+ L LDL N L IP G+L
Sbjct: 199 SGQIPSEIGNISLLKNFA-APSCFFNGPLPKEISKLKHLAKLDLSYNPLKCSIPKSFGEL 257
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L ++L + +L G IP LGN +SL +SFN L G +P L +
Sbjct: 258 HNLSILNLVSAELIGLIPPELGNCKSLKSLMLSFNSLSGPLPLELSE 304
Score = 71.6 bits (174), Expect = 1e-12
Identities = 43/124 (34%), Positives = 69/124 (54%), Gaps = 3/124 (2%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G +PA GNL L+ + L+ ++G + + S +++L L + N +G IPS LG
Sbjct: 688 LDGPVPASLGNLKELTHMDLS-FNNLSGELSSELSTMEKLVGLYIEQNKFTGEIPSELGN 746
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAG--LKKFNKNVFEHNK 203
L +L+ +D+S N LSG IP + L +L N++ N L G +P+ + +K + NK
Sbjct: 747 LTQLEYLDVSENLLSGEIPTKICGLPNLEFLNLAKNNLRGEVPSDGVCQDPSKALLSGNK 806
Query: 204 CLCG 207
LCG
Sbjct: 807 ELCG 810
Score = 70.1 bits (170), Expect = 4e-12
Identities = 42/104 (40%), Positives = 55/104 (52%), Gaps = 1/104 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G LPAE GN L L L++ ++TG IP KL L L+L +N G IP LG
Sbjct: 460 LEGYLPAEIGNAASLKRLVLSDN-QLTGEIPREIGKLTSLSVLNLNANMFQGKIPVELGD 518
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPA 189
L +DL +N L G IP + L L +S+N L G+IP+
Sbjct: 519 CTSLTTLDLGSNNLQGQIPDKITALAQLQCLVLSYNNLSGSIPS 562
Score = 66.6 bits (161), Expect = 4e-11
Identities = 44/130 (33%), Positives = 60/130 (45%), Gaps = 3/130 (2%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
G L+G LP LP L L L++ P+ F L L +LD+ +NSLSG IP
Sbjct: 122 GNSLTGLLPRLLSELPQLLYLDLSDNHFSGSLPPSFFISLPALSSLDVSNNSLSGEIPPE 181
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP---AGLKKFNKNVF 199
+GKL L + + N SG IP+ +GN+ L F G +P + LK K
Sbjct: 182 IGKLSNLSNLYMGLNSFSGQIPSEIGNISLLKNFAAPSCFFNGPLPKEISKLKHLAKLDL 241
Query: 200 EHNKCLCGAP 209
+N C P
Sbjct: 242 SYNPLKCSIP 251
Score = 65.9 bits (159), Expect = 7e-11
Identities = 40/114 (35%), Positives = 62/114 (54%), Gaps = 11/114 (9%)
Query: 86 LSGTLPA---------EFGNLPYLSMLSLAEMP--KVTGPIPNSFSKLQRLQNLDLGSNS 134
LSG++P+ E +L +L + ++ +++GPIP + L + L +N
Sbjct: 556 LSGSIPSKPSAYFHQIEMPDLSFLQHHGIFDLSYNRLSGPIPEELGECLVLVEISLSNNH 615
Query: 135 LSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
LSG IP+ L +L L +DLS N L+G+IP +GN L N++ NQL G IP
Sbjct: 616 LSGEIPASLSRLTNLTILDLSGNALTGSIPKEMGNSLKLQGLNLANNQLNGHIP 669
Score = 65.9 bits (159), Expect = 7e-11
Identities = 47/123 (38%), Positives = 60/123 (48%), Gaps = 10/123 (8%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSGT+ F L L L ++ G IP KL L LDL SN+ +G IP L K
Sbjct: 389 LSGTIEEVFDGCSSLGELLLTNN-QINGSIPEDLWKLP-LMALDLDSNNFTGEIPKSLWK 446
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK--------FNKN 197
L E S N+L G +PA +GN SL + +S NQL G IP + K N N
Sbjct: 447 STNLMEFTASYNRLEGYLPAEIGNAASLKRLVLSDNQLTGEIPREIGKLTSLSVLNLNAN 506
Query: 198 VFE 200
+F+
Sbjct: 507 MFQ 509
Score = 64.7 bits (156), Expect = 2e-10
Identities = 43/108 (39%), Positives = 57/108 (51%), Gaps = 1/108 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG+LP+ G L L LA + +G IP+ L++L L SN LSG IP L
Sbjct: 317 LSGSLPSWMGKWKVLDSLLLANN-RFSGEIPHEIEDCPMLKHLSLASNLLSGSIPRELCG 375
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L+ +DLS N LSGTI SL + ++ NQ+ G+IP L K
Sbjct: 376 SGSLEAIDLSGNLLSGTIEEVFDGCSSLGELLLTNNQINGSIPEDLWK 423
Score = 63.2 bits (152), Expect = 4e-10
Identities = 36/106 (33%), Positives = 55/106 (50%), Gaps = 2/106 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
++G++P + LP +++ + TG IP S K L N L G +P+ +G
Sbjct: 413 INGSIPEDLWKLPLMALD--LDSNNFTGEIPKSLWKSTNLMEFTASYNRLEGYLPAEIGN 470
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
LK + LS+N+L+G IP +G L SLS N++ N G IP L
Sbjct: 471 AASLKRLVLSDNQLTGEIPREIGKLTSLSVLNLNANMFQGKIPVEL 516
Score = 62.4 bits (150), Expect = 8e-10
Identities = 38/106 (35%), Positives = 57/106 (52%), Gaps = 2/106 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G +P E GN L L L+ ++GP+P S++ L N LSG +PS++GK
Sbjct: 270 LIGLIPPELGNCKSLKSLMLS-FNSLSGPLPLELSEIPLL-TFSAERNQLSGSLPSWMGK 327
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
K L + L+NN+ SG IP + + L +++ N L G+IP L
Sbjct: 328 WKVLDSLLLANNRFSGEIPHEIEDCPMLKHLSLASNLLSGSIPREL 373
Score = 60.8 bits (146), Expect = 2e-09
Identities = 44/118 (37%), Positives = 58/118 (48%), Gaps = 13/118 (11%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF----- 142
G +P E G+ L+ L L + G IP+ + L +LQ L L N+LSG IPS
Sbjct: 510 GKIPVELGDCTSLTTLDLGSN-NLQGQIPDKITALAQLQCLVLSYNNLSGSIPSKPSAYF 568
Query: 143 -------LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L L+ DLS N+LSG IP LG L + ++S N L G IPA L +
Sbjct: 569 HQIEMPDLSFLQHHGIFDLSYNRLSGPIPEELGECLVLVEISLSNNHLSGEIPASLSR 626
Score = 60.1 bits (144), Expect = 4e-09
Identities = 38/102 (37%), Positives = 55/102 (53%), Gaps = 2/102 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LP E +P L+ AE +++G +P+ K + L +L L +N SG IP +
Sbjct: 294 LSGPLPLELSEIPLLTFS--AERNQLSGSLPSWMGKWKVLDSLLLANNRFSGEIPHEIED 351
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
LK + L++N LSG+IP L SL ++S N L G I
Sbjct: 352 CPMLKHLSLASNLLSGSIPRELCGSGSLEAIDLSGNLLSGTI 393
Score = 54.7 bits (130), Expect = 2e-07
Identities = 38/107 (35%), Positives = 52/107 (48%), Gaps = 2/107 (1%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P E + P L LSLA ++G IP L+ +DL N LSG I
Sbjct: 342 SGEIPHEIEDCPMLKHLSLASN-LLSGSIPRELCGSGSLEAIDLSGNLLSGTIEEVFDGC 400
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
L E+ L+NN+++G+IP L L L ++ N G IP L K
Sbjct: 401 SSLGELLLTNNQINGSIPEDLWKL-PLMALDLDSNNFTGEIPKSLWK 446
Score = 53.1 bits (126), Expect = 5e-07
Identities = 29/83 (34%), Positives = 42/83 (49%)
Query: 106 AEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPA 165
A ++ G +P L+ L L N L+G IP +GKL L ++L+ N G IP
Sbjct: 455 ASYNRLEGYLPAEIGNAASLKRLVLSDNQLTGEIPREIGKLTSLSVLNLNANMFQGKIPV 514
Query: 166 SLGNLQSLSQFNVSFNQLCGAIP 188
LG+ SL+ ++ N L G IP
Sbjct: 515 ELGDCTSLTTLDLGSNNLQGQIP 537
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 82.8 bits (203), Expect = 5e-16
Identities = 50/113 (44%), Positives = 66/113 (58%), Gaps = 1/113 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E L L L L + +TGPIP S S +L + L +N LSG IP+ LG+
Sbjct: 487 LSGEIPQELMYLQALENLIL-DFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGR 545
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV 198
L L + L NN +SG IPA LGN QSL +++ N L G+IP L K + N+
Sbjct: 546 LSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNI 598
Score = 80.9 bits (198), Expect = 2e-15
Identities = 46/106 (43%), Positives = 65/106 (60%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G++P+ G+L L L L + +++G IP LQ L+NL L N L+GPIP+ L
Sbjct: 463 LTGSIPSSLGSLSKLKDLILW-LNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSN 521
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+L + LSNN+LSG IPASLG L +L+ + N + G IPA L
Sbjct: 522 CTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAEL 567
Score = 75.5 bits (184), Expect = 9e-14
Identities = 51/134 (38%), Positives = 64/134 (47%), Gaps = 25/134 (18%)
Query: 88 GTLPAEFGNLPYLSMLSLAE------MPK-------------------VTGPIPNSFSKL 122
G LP F NLP L L ++ +P GPIP+S S
Sbjct: 391 GGLPDSFSNLPKLETLDMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNC 450
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
+L +LDL N L+G IPS LG L +LK++ L N+LSG IP L LQ+L + FN
Sbjct: 451 SQLVSLDLSFNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFND 510
Query: 183 LCGAIPAGLKKFNK 196
L G IPA L K
Sbjct: 511 LTGPIPASLSNCTK 524
Score = 73.9 bits (180), Expect = 3e-13
Identities = 44/111 (39%), Positives = 63/111 (56%), Gaps = 9/111 (8%)
Query: 95 GNLPYLSMLSLAEMP-------KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK-- 145
G LP ++L L+ + K G +P+SFS L +L+ LD+ SN+L+G IPS + K
Sbjct: 366 GKLPVDTLLKLSNIKTMVLSFNKFVGGLPDSFSNLPKLETLDMSSNNLTGIIPSGICKDP 425
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
+ LK + L NN G IP SL N L ++SFN L G+IP+ L +K
Sbjct: 426 MNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLTGSIPSSLGSLSK 476
Score = 71.6 bits (174), Expect = 1e-12
Identities = 47/107 (43%), Positives = 59/107 (54%), Gaps = 4/107 (3%)
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGN 169
K+ G IP + L L+LG N LSG IP LG LK + +DLS N+ +GTIP SL +
Sbjct: 674 KLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTS 733
Query: 170 LQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPL-APC 213
L L + ++S N L G IP A F F +N LCG PL PC
Sbjct: 734 LTLLGEIDLSNNNLSGMIPESAPFDTFPDYRFANNS-LCGYPLPLPC 779
Score = 70.1 bits (170), Expect = 4e-12
Identities = 44/108 (40%), Positives = 60/108 (54%), Gaps = 1/108 (0%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P N L L L+ +TG IP+S L +L++L L N LSG IP L L+
Sbjct: 441 GPIPDSLSNCSQLVSLDLS-FNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQ 499
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
L+ + L N L+G IPASL N L+ ++S NQL G IPA L + +
Sbjct: 500 ALENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLS 547
Score = 67.8 bits (164), Expect = 2e-11
Identities = 40/100 (40%), Positives = 55/100 (55%), Gaps = 2/100 (2%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G++P E G + YLS+L+L ++G IP L+ + LDL N +G IP+ L
Sbjct: 675 LEGSIPKELGAMYYLSILNLGH-NDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTS 733
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
L L E+DLSNN LSG IP S + + + N LCG
Sbjct: 734 LTLLGEIDLSNNNLSGMIPES-APFDTFPDYRFANNSLCG 772
Score = 53.1 bits (126), Expect = 5e-07
Identities = 38/114 (33%), Positives = 56/114 (48%), Gaps = 8/114 (7%)
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKL-QRLQNLDLGSNSLSGPIPSFL 143
GL LP+E +L YL + G PN + L + + LDL N+ SG +P L
Sbjct: 295 GLVPKLPSE--SLQYLYLRG----NDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESL 348
Query: 144 GKLKRLKEVDLSNNKLSGTIPA-SLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
G+ L+ VD+SNN SG +P +L L ++ +SFN+ G +P K
Sbjct: 349 GECSSLELVDISNNNFSGKLPVDTLLKLSNIKTMVLSFNKFVGGLPDSFSNLPK 402
Score = 52.8 bits (125), Expect = 6e-07
Identities = 27/64 (42%), Positives = 39/64 (60%)
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
LDL N L G IP LG + L ++L +N LSG IP LG L++++ ++S+N+ G I
Sbjct: 668 LDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTI 727
Query: 188 PAGL 191
P L
Sbjct: 728 PNSL 731
Score = 45.4 bits (106), Expect = 1e-04
Identities = 37/120 (30%), Positives = 54/120 (44%), Gaps = 8/120 (6%)
Query: 82 RGWGLSGTLPA-EFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIP 140
+G L+G++P +F NL YL + + + SF LQ+LDL SN G I
Sbjct: 220 KGNKLAGSIPELDFKNLSYLDLSA-----NNFSTVFPSFKDCSNLQHLDLSSNKFYGDIG 274
Query: 141 SFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFE 200
S L +L ++L+NN+ G +P +SL + N G P L K V E
Sbjct: 275 SSLSSCGKLSFLNLTNNQFVGLVPKLPS--ESLQYLYLRGNDFQGVYPNQLADLCKTVVE 332
Score = 37.0 bits (84), Expect = 0.034
Identities = 41/144 (28%), Positives = 71/144 (48%), Gaps = 13/144 (9%)
Query: 50 LDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGW-GLSGTLPAEFGNLPYLSMLSLA-E 107
L +W ++T+ C ++ V C RV+ + +S + + +L + LP ++ SL +
Sbjct: 61 LQNWLSSTDPCSFTGVSCKN---SRVSSIDLSNTFLSVDFSLVTSY-LLPLSNLESLVLK 116
Query: 108 MPKVTGPIPNSFSKLQ---RLQNLDLGSNSLSGPIP--SFLGKLKRLKEVDLSNNKLSGT 162
++G + S +K Q L ++DL N++SGPI S G LK ++LS N L
Sbjct: 117 NANLSGSL-TSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPP 175
Query: 163 IPASL-GNLQSLSQFNVSFNQLCG 185
L G SL ++S+N + G
Sbjct: 176 GKEMLKGATFSLQVLDLSYNNISG 199
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor
(EC 2.7.1.37) (tBRI1) (Altered brassinolide sensitivity
1) (Systemin receptor SR160)
Length = 1207
Score = 82.8 bits (203), Expect = 5e-16
Identities = 50/113 (44%), Positives = 66/113 (58%), Gaps = 1/113 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E L L L L + +TGPIP S S +L + L +N LSG IP+ LG+
Sbjct: 487 LSGEIPQELMYLQALENLIL-DFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGR 545
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV 198
L L + L NN +SG IPA LGN QSL +++ N L G+IP L K + N+
Sbjct: 546 LSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNI 598
Score = 80.9 bits (198), Expect = 2e-15
Identities = 46/106 (43%), Positives = 65/106 (60%), Gaps = 1/106 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G++P+ G+L L L L + +++G IP LQ L+NL L N L+GPIP+ L
Sbjct: 463 LTGSIPSSLGSLSKLKDLILW-LNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSN 521
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+L + LSNN+LSG IPASLG L +L+ + N + G IPA L
Sbjct: 522 CTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAEL 567
Score = 74.3 bits (181), Expect = 2e-13
Identities = 41/84 (48%), Positives = 51/84 (59%)
Query: 113 GPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQS 172
GPIP+S S +L +LDL N L+G IPS LG L +LK++ L N+LSG IP L LQ+
Sbjct: 441 GPIPDSLSNCSQLVSLDLSFNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQA 500
Query: 173 LSQFNVSFNQLCGAIPAGLKKFNK 196
L + FN L G IPA L K
Sbjct: 501 LENLILDFNDLTGPIPASLSNCTK 524
Score = 72.8 bits (177), Expect = 6e-13
Identities = 46/114 (40%), Positives = 64/114 (55%), Gaps = 6/114 (5%)
Query: 87 SGTLPAEFGNLPYLSMLS--LAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
SG LP + L LS + + K G +P+SFS L +L+ LD+ SN+L+G IPS +
Sbjct: 365 SGKLPVD--TLSKLSNIKTMVLSFNKFVGGLPDSFSNLLKLETLDMSSNNLTGVIPSGIC 422
Query: 145 K--LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
K + LK + L NN G IP SL N L ++SFN L G+IP+ L +K
Sbjct: 423 KDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLTGSIPSSLGSLSK 476
Score = 71.6 bits (174), Expect = 1e-12
Identities = 47/107 (43%), Positives = 59/107 (54%), Gaps = 4/107 (3%)
Query: 110 KVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGN 169
K+ G IP + L L+LG N LSG IP LG LK + +DLS N+ +GTIP SL +
Sbjct: 674 KLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTS 733
Query: 170 LQSLSQFNVSFNQLCGAIP--AGLKKFNKNVFEHNKCLCGAPL-APC 213
L L + ++S N L G IP A F F +N LCG PL PC
Sbjct: 734 LTLLGEIDLSNNNLSGMIPESAPFDTFPDYRFANNS-LCGYPLPIPC 779
Score = 70.1 bits (170), Expect = 4e-12
Identities = 44/108 (40%), Positives = 60/108 (54%), Gaps = 1/108 (0%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLK 147
G +P N L L L+ +TG IP+S L +L++L L N LSG IP L L+
Sbjct: 441 GPIPDSLSNCSQLVSLDLS-FNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQ 499
Query: 148 RLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFN 195
L+ + L N L+G IPASL N L+ ++S NQL G IPA L + +
Sbjct: 500 ALENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLS 547
Score = 67.8 bits (164), Expect = 2e-11
Identities = 40/100 (40%), Positives = 55/100 (55%), Gaps = 2/100 (2%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G++P E G + YLS+L+L ++G IP L+ + LDL N +G IP+ L
Sbjct: 675 LEGSIPKELGAMYYLSILNLGH-NDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTS 733
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCG 185
L L E+DLSNN LSG IP S + + + N LCG
Sbjct: 734 LTLLGEIDLSNNNLSGMIPES-APFDTFPDYRFANNSLCG 772
Score = 67.0 bits (162), Expect = 3e-11
Identities = 43/106 (40%), Positives = 57/106 (53%), Gaps = 3/106 (2%)
Query: 88 GTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK--LQRLQNLDLGSNSLSGPIPSFLGK 145
G LP F NL L L ++ +TG IP+ K + L+ L L +N GPIP L
Sbjct: 391 GGLPDSFSNLLKLETLDMSSN-NLTGVIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSN 449
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+L +DLS N L+G+IP+SLG+L L + NQL G IP L
Sbjct: 450 CSQLVSLDLSFNYLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQEL 495
Score = 60.1 bits (144), Expect = 4e-09
Identities = 40/131 (30%), Positives = 58/131 (43%), Gaps = 1/131 (0%)
Query: 64 FVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQ 123
FVG P RG G P + +L + +G +P S +
Sbjct: 293 FVGLVPKLPSESLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLGECS 352
Query: 124 RLQNLDLGSNSLSGPIP-SFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
L+ +D+ N+ SG +P L KL +K + LS NK G +P S NL L ++S N
Sbjct: 353 SLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLPDSFSNLLKLETLDMSSNN 412
Query: 183 LCGAIPAGLKK 193
L G IP+G+ K
Sbjct: 413 LTGVIPSGICK 423
Score = 52.8 bits (125), Expect = 6e-07
Identities = 27/64 (42%), Positives = 39/64 (60%)
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
LDL N L G IP LG + L ++L +N LSG IP LG L++++ ++S+N+ G I
Sbjct: 668 LDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTI 727
Query: 188 PAGL 191
P L
Sbjct: 728 PNSL 731
Score = 47.4 bits (111), Expect = 3e-05
Identities = 49/180 (27%), Positives = 67/180 (37%), Gaps = 57/180 (31%)
Query: 82 RGWGLSGTLPA-EFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIP 140
+G L+G++P +F NL YL + + + SF LQ+LDL SN G I
Sbjct: 220 KGNKLAGSIPELDFKNLSYLDLSA-----NNFSTVFPSFKDCSNLQHLDLSSNKFYGDIG 274
Query: 141 SFL---GKL--------------------------------------------KRLKEVD 153
S L GKL K + E+D
Sbjct: 275 SSLSSCGKLSFLNLTNNQFVGLVPKLPSESLQYLYLRGNDFQGVYPNQLADLCKTVVELD 334
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP----AGLKKFNKNVFEHNKCLCGAP 209
LS N SG +P SLG SL ++S+N G +P + L V NK + G P
Sbjct: 335 LSYNNFSGMVPESLGECSSLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLP 394
Score = 33.1 bits (74), Expect = 0.49
Identities = 40/144 (27%), Positives = 69/144 (47%), Gaps = 13/144 (9%)
Query: 50 LDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGW-GLSGTLPAEFGNLPYLSMLSLA-E 107
L +W ++T C ++ V C RV+ + +S + + +L + LP ++ SL +
Sbjct: 61 LQNWLSSTGPCSFTGVSCKN---SRVSSIDLSNTFLSVDFSLVTSY-LLPLSNLESLVLK 116
Query: 108 MPKVTGPIPNSFSKLQ---RLQNLDLGSNSLSGPIP--SFLGKLKRLKEVDLSNNKLSGT 162
++G + S +K Q L ++DL N++SGPI S G LK ++LS N L
Sbjct: 117 NANLSGSL-TSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLNLSKNFLDPP 175
Query: 163 IPASL-GNLQSLSQFNVSFNQLCG 185
L SL ++S+N + G
Sbjct: 176 GKEMLKAATFSLQVLDLSYNNISG 199
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 78.2 bits (191), Expect = 1e-14
Identities = 65/218 (29%), Positives = 100/218 (45%), Gaps = 37/218 (16%)
Query: 1 MFLLLHLTLLFL---SSIYFSKSNASDDFFCNADDKAALLKIRDHFGG-PKGRLDDWD-N 55
M + ++ LLFL SSIY + F D AALL + H+ P W+ +
Sbjct: 1 MKVAVNTFLLFLCSTSSIYAA--------FALNSDGAALLSLTRHWTSIPSDITQSWNAS 52
Query: 56 NTECCDWSFVGCGR----------------------PYPGRVTVVTISRGWGLSGTLPAE 93
++ C W V C R + + V +S G G G++P++
Sbjct: 53 DSTPCSWLGVECDRRQFVDTLNLSSYGISGEFGPEISHLKHLKKVVLS-GNGFFGSIPSQ 111
Query: 94 FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVD 153
GN L + L+ TG IP++ LQ L+NL L NSL GP P L + L+ V
Sbjct: 112 LGNCSLLEHIDLSSN-SFTGNIPDTLGALQNLRNLSLFFNSLIGPFPESLLSIPHLETVY 170
Query: 154 LSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+ N L+G+IP+++GN+ L+ + NQ G +P+ L
Sbjct: 171 FTGNGLNGSIPSNIGNMSELTTLWLDDNQFSGPVPSSL 208
Score = 77.8 bits (190), Expect = 2e-14
Identities = 54/147 (36%), Positives = 77/147 (51%), Gaps = 28/147 (19%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK-----------------------L 122
L+G++P+ G+L L+ LSL E +G IP S + L
Sbjct: 583 LNGSIPSTLGSLTELTKLSLGEN-SFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPVGAL 641
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
Q L++L+L SN L+G +P LGKLK L+E+D+S+N LSGT+ L +QSL+ N+S N
Sbjct: 642 QALRSLNLSSNKLNGQLPIDLGKLKMLEELDVSHNNLSGTLRV-LSTIQSLTFINISHNL 700
Query: 183 LCGAIPAGLKKF---NKNVFEHNKCLC 206
G +P L KF + F N LC
Sbjct: 701 FSGPVPPSLTKFLNSSPTSFSGNSDLC 727
Score = 75.5 bits (184), Expect = 9e-14
Identities = 43/132 (32%), Positives = 67/132 (50%), Gaps = 2/132 (1%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
G GL+G++P+ GN+ L+ L L + + +GP+P+S + LQ L L N+L G +P
Sbjct: 173 GNGLNGSIPSNIGNMSELTTLWLDDN-QFSGPVPSSLGNITTLQELYLNDNNLVGTLPVT 231
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK-NVFEH 201
L L+ L +D+ NN L G IP + + + ++S NQ G +P GL F
Sbjct: 232 LNNLENLVYLDVRNNSLVGAIPLDFVSCKQIDTISLSNNQFTGGLPPGLGNCTSLREFGA 291
Query: 202 NKCLCGAPLAPC 213
C P+ C
Sbjct: 292 FSCALSGPIPSC 303
Score = 75.1 bits (183), Expect = 1e-13
Identities = 40/114 (35%), Positives = 67/114 (58%), Gaps = 1/114 (0%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
G +G +P GNL ++ + L+ +++G IP L +L++L+L N L G +PS
Sbjct: 508 GNNFTGPIPPSLGNLKNVTAIYLSSN-QLSGSIPPELGSLVKLEHLNLSHNILKGILPSE 566
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
L +L E+D S+N L+G+IP++LG+L L++ ++ N G IP L + NK
Sbjct: 567 LSNCHKLSELDASHNLLNGSIPSTLGSLTELTKLSLGENSFSGGIPTSLFQSNK 620
Score = 74.7 bits (182), Expect = 1e-13
Identities = 44/111 (39%), Positives = 64/111 (57%), Gaps = 2/111 (1%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G++P++ G L L L E + G +P+ F + Q L DL N+ +GPIP LG
Sbjct: 464 LEGSVPSDLGGCSTLERLILEEN-NLRGGLPD-FVEKQNLLFFDLSGNNFTGPIPPSLGN 521
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
LK + + LS+N+LSG+IP LG+L L N+S N L G +P+ L +K
Sbjct: 522 LKNVTAIYLSSNQLSGSIPPELGSLVKLEHLNLSHNILKGILPSELSNCHK 572
Score = 69.7 bits (169), Expect = 5e-12
Identities = 38/103 (36%), Positives = 57/103 (54%), Gaps = 1/103 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G P ++P+L + + G IP++ + L L L N SGP+PS LG
Sbjct: 152 LIGPFPESLLSIPHLETVYFTGNG-LNGSIPSNIGNMSELTTLWLDDNQFSGPVPSSLGN 210
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+ L+E+ L++N L GT+P +L NL++L +V N L GAIP
Sbjct: 211 ITTLQELYLNDNNLVGTLPVTLNNLENLVYLDVRNNSLVGAIP 253
Score = 67.8 bits (164), Expect = 2e-11
Identities = 40/107 (37%), Positives = 54/107 (50%), Gaps = 1/107 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
+G LP GN L A ++GPIP+ F +L +L L L N SG IP LGK
Sbjct: 273 TGGLPPGLGNCTSLREFG-AFSCALSGPIPSCFGQLTKLDTLYLAGNHFSGRIPPELGKC 331
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKK 193
K + ++ L N+L G IP LG L L ++ N L G +P + K
Sbjct: 332 KSMIDLQLQQNQLEGEIPGELGMLSQLQYLHLYTNNLSGEVPLSIWK 378
Score = 67.0 bits (162), Expect = 3e-11
Identities = 41/113 (36%), Positives = 58/113 (51%), Gaps = 1/113 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P+ FG L L L LA +G IP K + + +L L N L G IP LG
Sbjct: 296 LSGPIPSCFGQLTKLDTLYLAGN-HFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGM 354
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV 198
L +L+ + L N LSG +P S+ +QSL + N L G +P + + + V
Sbjct: 355 LSQLQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNNLSGELPVDMTELKQLV 407
Score = 67.0 bits (162), Expect = 3e-11
Identities = 49/142 (34%), Positives = 67/142 (46%), Gaps = 25/142 (17%)
Query: 75 VTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNS 134
VT + +S LSG++P E G+L L L+L+ + G +P+ S +L LD N
Sbjct: 525 VTAIYLSSNQ-LSGSIPPELGSLVKLEHLNLSHNI-LKGILPSELSNCHKLSELDASHNL 582
Query: 135 LSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASL-----------------------GNLQ 171
L+G IPS LG L L ++ L N SG IP SL G LQ
Sbjct: 583 LNGSIPSTLGSLTELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPVGALQ 642
Query: 172 SLSQFNVSFNQLCGAIPAGLKK 193
+L N+S N+L G +P L K
Sbjct: 643 ALRSLNLSSNKLNGQLPIDLGK 664
Score = 65.9 bits (159), Expect = 7e-11
Identities = 43/120 (35%), Positives = 62/120 (50%), Gaps = 4/120 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L G +P E G L L L L ++G +P S K+Q LQ+L L N+LSG +P + +
Sbjct: 344 LEGEIPGELGMLSQLQYLHLYTN-NLSGEVPLSIWKIQSLQSLQLYQNNLSGELPVDMTE 402
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL---KKFNKNVFEHN 202
LK+L + L N +G IP LG SL +++ N G IP L KK + + +N
Sbjct: 403 LKQLVSLALYENHFTGVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQKKLKRLLLGYN 462
Score = 65.1 bits (157), Expect = 1e-10
Identities = 38/110 (34%), Positives = 58/110 (52%), Gaps = 1/110 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P+ GN+ L L L + + G +P + + L+ L LD+ +NSL G IP
Sbjct: 201 SGPVPSSLGNITTLQELYLNDN-NLVGTLPVTLNNLENLVYLDVRNNSLVGAIPLDFVSC 259
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
K++ + LSNN+ +G +P LGN SL +F L G IP+ + K
Sbjct: 260 KQIDTISLSNNQFTGGLPPGLGNCTSLREFGAFSCALSGPIPSCFGQLTK 309
Score = 62.8 bits (151), Expect = 6e-10
Identities = 39/120 (32%), Positives = 59/120 (48%), Gaps = 1/120 (0%)
Query: 82 RGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPS 141
R L G +P +F + + +SL+ + TG +P L+ S +LSGPIPS
Sbjct: 244 RNNSLVGAIPLDFVSCKQIDTISLSNN-QFTGGLPPGLGNCTSLREFGAFSCALSGPIPS 302
Query: 142 FLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEH 201
G+L +L + L+ N SG IP LG +S+ + NQL G IP L ++ + H
Sbjct: 303 CFGQLTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGMLSQLQYLH 362
Score = 62.0 bits (149), Expect = 1e-09
Identities = 43/127 (33%), Positives = 61/127 (47%), Gaps = 1/127 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG LP + L L L+L E TG IP L+ LDL N +G IP L
Sbjct: 392 LSGELPVDMTELKQLVSLALYEN-HFTGVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCS 450
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCL 205
K+LK + L N L G++P+ LG +L + + N L G +P ++K N F+ +
Sbjct: 451 QKKLKRLLLGYNYLEGSVPSDLGGCSTLERLILEENNLRGGLPDFVEKQNLLFFDLSGNN 510
Query: 206 CGAPLAP 212
P+ P
Sbjct: 511 FTGPIPP 517
Score = 60.1 bits (144), Expect = 4e-09
Identities = 35/105 (33%), Positives = 53/105 (50%), Gaps = 1/105 (0%)
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKL 146
SG +P E G + L L + ++ G IP L +LQ L L +N+LSG +P + K+
Sbjct: 321 SGRIPPELGKCKSMIDLQL-QQNQLEGEIPGELGMLSQLQYLHLYTNNLSGEVPLSIWKI 379
Query: 147 KRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
+ L+ + L N LSG +P + L+ L + N G IP L
Sbjct: 380 QSLQSLQLYQNNLSGELPVDMTELKQLVSLALYENHFTGVIPQDL 424
>TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor
Length = 674
Score = 75.9 bits (185), Expect = 7e-14
Identities = 54/148 (36%), Positives = 71/148 (47%), Gaps = 7/148 (4%)
Query: 71 YPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSF---SKLQRLQN 127
Y ++ V +S G L+G LP NL + ++G +P S LQ
Sbjct: 145 YTSSLSDVDLS-GNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLPEPALPNSTCGNLQV 203
Query: 128 LDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAI 187
LDLG N SG P F+ + K +K +DLS+N G +P LG L+ L N+S N G +
Sbjct: 204 LDLGGNKFSGEFPEFITRFKGVKSLDLSSNVFEGLVPEGLGVLE-LESLNLSHNNFSGML 262
Query: 188 P-AGLKKFNKNVFEHNK-CLCGAPLAPC 213
P G KF FE N LCG PL PC
Sbjct: 263 PDFGESKFGAESFEGNSPSLCGLPLKPC 290
Score = 62.8 bits (151), Expect = 6e-10
Identities = 52/165 (31%), Positives = 76/165 (45%), Gaps = 10/165 (6%)
Query: 29 NADDKAALLKIRDHFGGPKGRL--DDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGWGL 86
++D K L KI+ G L W+++ C W V + +S
Sbjct: 30 SSDVKLLLGKIKSSLQGNSESLLLSSWNSSVPVCQWRGVKWVFSNGSPLQCSDLSSPQWT 89
Query: 87 SGTLPAEFGNLPYLSMLSLAEMPK--VTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
+ +L N L +LSL ++P +TG +P + LQ++ L NSLSG IP LG
Sbjct: 90 NTSL----FNDSSLHLLSL-QLPSANLTGSLPREIGEFSMLQSVFLNINSLSGSIPLELG 144
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNL-QSLSQFNVSFNQLCGAIP 188
L +VDLS N L+G +P S+ NL L F + N L G +P
Sbjct: 145 YTSSLSDVDLSGNALAGVLPPSIWNLCDKLVSFKIHGNNLSGVLP 189
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 74.3 bits (181), Expect = 2e-13
Identities = 49/128 (38%), Positives = 63/128 (48%), Gaps = 23/128 (17%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
LSG +P E G++PYL +L+L ++G IP+ L+ L LDL SN L G IP +
Sbjct: 666 LSGYIPKEIGSMPYLFILNLGH-NDISGSIPDEVGDLRGLNILDLSSNKLDGRIPQAMSA 724
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNVFEHNKCL 205
L L E+DLSNN LSG IP + QF + F F +N L
Sbjct: 725 LTMLTEIDLSNNNLSGPIP-------EMGQF---------------ETFPPAKFLNNPGL 762
Query: 206 CGAPLAPC 213
CG PL C
Sbjct: 763 CGYPLPRC 770
Score = 72.0 bits (175), Expect = 1e-12
Identities = 46/131 (35%), Positives = 67/131 (51%), Gaps = 23/131 (17%)
Query: 86 LSGTLPAEFGNLPYLSMLSL------AEMPK-----------------VTGPIPNSFSKL 122
LSGT+P+ G+L L L L E+P+ +TG IP+ S
Sbjct: 452 LSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNC 511
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
L + L +N L+G IP ++G+L+ L + LSNN SG IPA LG+ +SL +++ N
Sbjct: 512 TNLNWISLSNNRLTGEIPKWIGRLENLAILKLSNNSFSGNIPAELGDCRSLIWLDLNTNL 571
Query: 183 LCGAIPAGLKK 193
G IPA + K
Sbjct: 572 FNGTIPAAMFK 582
Score = 63.2 bits (152), Expect = 4e-10
Identities = 35/80 (43%), Positives = 46/80 (56%)
Query: 112 TGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQ 171
TG IP + S L +L L N LSG IPS LG L +L+++ L N L G IP L ++
Sbjct: 429 TGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVK 488
Query: 172 SLSQFNVSFNQLCGAIPAGL 191
+L + FN L G IP+GL
Sbjct: 489 TLETLILDFNDLTGEIPSGL 508
Score = 58.5 bits (140), Expect = 1e-08
Identities = 43/114 (37%), Positives = 61/114 (52%), Gaps = 5/114 (4%)
Query: 87 SGTLPAE-FGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR-LQNLDLGSNSLSGPI-PSFL 143
SG LP + + L +L L+ + +G +P S + L L LDL SN+ SGPI P+
Sbjct: 353 SGELPMDTLLKMRGLKVLDLS-FNEFSGELPESLTNLSASLLTLDLSSNNFSGPILPNLC 411
Query: 144 GKLKR-LKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGLKKFNK 196
K L+E+ L NN +G IP +L N L ++SFN L G IP+ L +K
Sbjct: 412 QNPKNTLQELYLQNNGFTGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSK 465
Score = 56.2 bits (134), Expect = 5e-08
Identities = 37/104 (35%), Positives = 53/104 (50%), Gaps = 3/104 (2%)
Query: 97 LPYLSMLSLAEMPKVTGPIPNSFS-KLQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLS 155
L L LSLAE K TG IP+ S L LDL N G +P F G L+ + LS
Sbjct: 290 LKSLQYLSLAEN-KFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSCSLLESLALS 348
Query: 156 NNKLSGTIPA-SLGNLQSLSQFNVSFNQLCGAIPAGLKKFNKNV 198
+N SG +P +L ++ L ++SFN+ G +P L + ++
Sbjct: 349 SNNFSGELPMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASL 392
Score = 55.8 bits (133), Expect = 7e-08
Identities = 44/108 (40%), Positives = 54/108 (49%), Gaps = 4/108 (3%)
Query: 87 SGTLPAEFGNLPYLSMLSL-AEMPKVTGPI-PNSFSKLQR-LQNLDLGSNSLSGPIPSFL 143
SG LP NL S+L+L +GPI PN + LQ L L +N +G IP L
Sbjct: 378 SGELPESLTNLS-ASLLTLDLSSNNFSGPILPNLCQNPKNTLQELYLQNNGFTGKIPPTL 436
Query: 144 GKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
L + LS N LSGTIP+SLG+L L + N L G IP L
Sbjct: 437 SNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWLNMLEGEIPQEL 484
Score = 52.4 bits (124), Expect = 8e-07
Identities = 37/106 (34%), Positives = 53/106 (49%), Gaps = 11/106 (10%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF 142
GW LS G L +L++ K++G + S+ L+ LD+ SN+ S IP F
Sbjct: 192 GWVLSDGC----GELKHLAISG----NKISGDV--DVSRCVNLEFLDVSSNNFSTGIP-F 240
Query: 143 LGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
LG L+ +D+S NKLSG ++ L N+S NQ G IP
Sbjct: 241 LGDCSALQHLDISGNKLSGDFSRAISTCTELKLLNISSNQFVGPIP 286
Score = 43.5 bits (101), Expect = 4e-04
Identities = 43/148 (29%), Positives = 66/148 (44%), Gaps = 8/148 (5%)
Query: 47 KGRLDDWDNNTECCDWSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLA 106
K L DW +N C + V C + + + G S + + +L L L L+
Sbjct: 49 KNLLPDWSSNKNPCTFDGVTCRDDKVTSIDLSSKPLNVGFSA-VSSSLLSLTGLESLFLS 107
Query: 107 EMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSF--LGKLKRLKEVDLSNNKLS--GT 162
+ G + + F L +LDL NSLSGP+ + LG LK +++S+N L G
Sbjct: 108 NS-HINGSV-SGFKCSASLTSLDLSRNSLSGPVTTLTSLGSCSGLKFLNVSSNTLDFPGK 165
Query: 163 IPASLGNLQSLSQFNVSFNQLCGAIPAG 190
+ L L SL ++S N + GA G
Sbjct: 166 VSGGL-KLNSLEVLDLSANSISGANVVG 192
>TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM)
Length = 496
Score = 68.2 bits (165), Expect = 1e-11
Identities = 43/133 (32%), Positives = 65/133 (48%), Gaps = 23/133 (17%)
Query: 85 GLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSN----------- 133
G G +P E GNL L +L L + + G IP SF++ L++LDL N
Sbjct: 170 GFLGPIPDELGNLTNLKVLDLHKN-HLNGSIPLSFNRFSGLRSLDLSGNRLTGSIPGFVL 228
Query: 134 -----------SLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQ 182
L+GP+P L L ++DLS N+++G IP S+ L L ++S+N+
Sbjct: 229 PALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINRLNQLVLLDLSYNR 288
Query: 183 LCGAIPAGLKKFN 195
L G P+ L+ N
Sbjct: 289 LSGPFPSSLQGLN 301
Score = 63.2 bits (152), Expect = 4e-10
Identities = 42/114 (36%), Positives = 63/114 (54%), Gaps = 4/114 (3%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQRLQNLDLGSNSLSGPIPSFLGK 145
L+G++P LP LS+L L + +TGP+P + + L +DL N ++GPIP + +
Sbjct: 219 LTGSIPGFV--LPALSVLDLNQN-LLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINR 275
Query: 146 LKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN-QLCGAIPAGLKKFNKNV 198
L +L +DLS N+LSG P+SL L SL + N + IP K KN+
Sbjct: 276 LNQLVLLDLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNL 329
Score = 57.8 bits (138), Expect = 2e-08
Identities = 35/104 (33%), Positives = 53/104 (50%), Gaps = 1/104 (0%)
Query: 86 LSGTLPAEFGNLPYLSMLSLAEMPKVTGPIP-NSFSKLQRLQNLDLGSNSLSGPIPSFLG 144
LSG P+ L L L L K + IP N+F L+ L L L + ++ G IP L
Sbjct: 289 LSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQGSIPKSLT 348
Query: 145 KLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLCGAIP 188
+L L+ + L N L+G IP +++ LS+ ++ N L G +P
Sbjct: 349 RLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVP 392
Score = 57.4 bits (137), Expect = 2e-08
Identities = 35/88 (39%), Positives = 46/88 (51%), Gaps = 1/88 (1%)
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFNQLC 184
LQ L L N GPIP LG L LK +DL N L+G+IP S L ++S N+L
Sbjct: 161 LQTLVLRENGFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNRLT 220
Query: 185 GAIPAGLKKFNKNVFEHNKCLCGAPLAP 212
G+IP G +V + N+ L P+ P
Sbjct: 221 GSIP-GFVLPALSVLDLNQNLLTGPVPP 247
Score = 30.0 bits (66), Expect = 4.2
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 11/91 (12%)
Query: 83 GWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIP-------NSFSKLQRLQNLDLGSN-- 133
G L+G +P EF ++ +LS L L + +TGP+P KL+ N L N
Sbjct: 360 GNNLTGEIPLEFRDVKHLSELRLND-NSLTGPVPFERDTVWRMRRKLRLYNNAGLCVNRD 418
Query: 134 -SLSGPIPSFLGKLKRLKEVDLSNNKLSGTI 163
L S G RL + + S SGT+
Sbjct: 419 SDLDDAFGSKSGSTVRLCDAETSRPAPSGTV 449
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 61.6 bits (148), Expect = 1e-09
Identities = 45/166 (27%), Positives = 72/166 (43%), Gaps = 34/166 (20%)
Query: 10 LFLSSIYFSKSNASDDFFCNADDKA-----ALLKIRDHFGGPKGRLDDWDNNTECCDWSF 64
+F SS+ S+ F ++ + +LL I F P + W N C +W
Sbjct: 297 VFKSSVSVDLDKDSNSFCLSSPGECDPRVKSLLLIASSFDYPPRLAESWKGNDPCTNWIG 356
Query: 65 VGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSKLQR 124
+ C G +TV+++ + L+GT+ EFG ++
Sbjct: 357 IACSN---GNITVISLEK-MELTGTISPEFG-------------------------AIKS 387
Query: 125 LQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNL 170
LQ + LG N+L+G IP L L LK +D+S+NKL G +P N+
Sbjct: 388 LQRIILGINNLTGMIPQELTTLPNLKTLDVSSNKLFGKVPGFRSNV 433
Score = 58.5 bits (140), Expect = 1e-08
Identities = 47/152 (30%), Positives = 72/152 (46%), Gaps = 26/152 (17%)
Query: 86 LSGTLPAEFG--NLPYLSMLSLA------EMP----------------KVTGPIPNSFSK 121
+SG+LP G P LS+L LA E+P K+TG I
Sbjct: 172 VSGSLPGFLGPDEFPGLSILHLAFNNLEGELPMSLAGSQVQSLWLNGQKLTGDI-TVLQN 230
Query: 122 LQRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIPASLGNLQSLSQFNVSFN 181
+ L+ + L SN SGP+P F G LK L+ + L +N +G +PASL +L+SL N++ N
Sbjct: 231 MTGLKEVWLHSNKFSGPLPDFSG-LKELESLSLRDNSFTGPVPASLLSLESLKVVNLTNN 289
Query: 182 QLCGAIPAGLKKFNKNVFEHNKCLCGAPLAPC 213
L G +P + ++ + + C + C
Sbjct: 290 HLQGPVPVFKSSVSVDLDKDSNSFCLSSPGEC 321
Score = 45.8 bits (107), Expect = 7e-05
Identities = 55/214 (25%), Positives = 81/214 (37%), Gaps = 64/214 (29%)
Query: 2 FLLLHLTLLFLSSIYFSKSNASDDFFCNADDKAALLKIRDHFGGPKGRLDDWDNNTECCD 61
FLL T L L S+ S A D D +A+L ++ P W ++ + C
Sbjct: 7 FLLFSFTFLLLLSL----SKADSD-----GDLSAMLSLKKSLNPPSSF--GW-SDPDPCK 54
Query: 62 WSFVGCGRPYPGRVTVVTISRGWGLSGTLPAEFGNLPYLSMLSLAEMPKVTGPIPNSFSK 121
W+ + C RVT + I GL GTL + NL
Sbjct: 55 WTHIVCTGTK--RVTRIQIGHS-GLQGTLSPDLRNL------------------------ 87
Query: 122 LQRLQNLDLGSNSLSGPIPSFLG-----------------------KLKRLKEVDLSNNK 158
L+ L+L N++SGP+PS G L L+ V++ NN
Sbjct: 88 -SELERLELQWNNISGPVPSLSGLASLQVLMLSNNNFDSIPSDVFQGLTSLQSVEIDNNP 146
Query: 159 L-SGTIPASLGNLQSLSQFNVSFNQLCGAIPAGL 191
S IP SL N +L F+ + + G++P L
Sbjct: 147 FKSWEIPESLRNASALQNFSANSANVSGSLPGFL 180
Score = 35.8 bits (81), Expect = 0.076
Identities = 21/82 (25%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Query: 123 QRLQNLDLGSNSLSGPIPSFLGKLKRLKEVDLSNNKLSGTIP--ASLGNLQSLSQFNVSF 180
+R+ + +G + L G + L L L+ ++L N +SG +P + L +LQ L N +F
Sbjct: 64 KRVTRIQIGHSGLQGTLSPDLRNLSELERLELQWNNISGPVPSLSGLASLQVLMLSNNNF 123
Query: 181 NQLCGAIPAGLKKFNKNVFEHN 202
+ + + GL ++N
Sbjct: 124 DSIPSDVFQGLTSLQSVEIDNN 145
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,695,599
Number of Sequences: 164201
Number of extensions: 1162570
Number of successful extensions: 3817
Number of sequences better than 10.0: 285
Number of HSP's better than 10.0 without gapping: 154
Number of HSP's successfully gapped in prelim test: 131
Number of HSP's that attempted gapping in prelim test: 2491
Number of HSP's gapped (non-prelim): 1001
length of query: 214
length of database: 59,974,054
effective HSP length: 106
effective length of query: 108
effective length of database: 42,568,748
effective search space: 4597424784
effective search space used: 4597424784
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146750.9


