

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146747.6 + phase: 0
(498 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
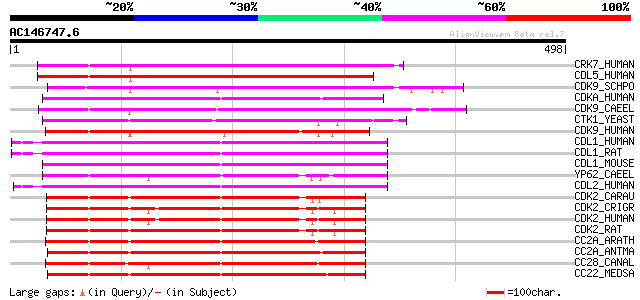
Score E
Sequences producing significant alignments: (bits) Value
CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase... 278 2e-74
CDL5_HUMAN (Q14004) Cell division cycle 2-like protein kinase 5 ... 267 5e-71
CDK9_SCHPO (Q96WV9) Serine/threonine-protein kinase cdk9 (EC 2.7... 261 4e-69
CDKA_HUMAN (Q15131) Cell division protein kinase 10 (EC 2.7.1.37... 244 4e-64
CDK9_CAEEL (P46551) Putative cell division protein kinase 9 homo... 244 4e-64
CTK1_YEAST (Q03957) CTD kinase alpha subunit (EC 2.7.1.-) (CTD k... 236 1e-61
CDK9_HUMAN (P50750) Cell division protein kinase 9 (EC 2.7.1.37)... 236 1e-61
CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2... 227 5e-59
CDL1_RAT (P46892) PITSLRE serine/threonine-protein kinase CDC2L1... 226 1e-58
CDL1_MOUSE (P24788) PITSLRE serine/threonine-protein kinase CDC2... 223 9e-58
YP62_CAEEL (Q09437) Putative serine/threonine-protein kinase B04... 223 1e-57
CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2... 223 1e-57
CDK2_CARAU (P43450) Cell division protein kinase 2 (EC 2.7.1.37) 223 1e-57
CDK2_CRIGR (O55076) Cell division protein kinase 2 (EC 2.7.1.37) 222 2e-57
CDK2_HUMAN (P24941) Cell division protein kinase 2 (EC 2.7.1.37)... 221 3e-57
CDK2_RAT (Q63699) Cell division protein kinase 2 (EC 2.7.1.37) 221 3e-57
CC2A_ARATH (P24100) Cell division control protein 2 homolog A (E... 221 4e-57
CC2A_ANTMA (Q38772) Cell division control protein 2 homolog A (E... 221 4e-57
CC28_CANAL (P43063) Cell division control protein 28 (EC 2.7.1.37) 221 4e-57
CC22_MEDSA (Q05006) Cell division control protein 2 homolog 2 (E... 221 4e-57
>CRK7_HUMAN (Q9NYV4) Cell division cycle 2-related protein kinase 7
(EC 2.7.1.37) (CDC2-related protein kinase 7) (CrkRS)
Length = 1490
Score = 278 bits (711), Expect = 2e-74
Identities = 155/339 (45%), Positives = 202/339 (58%), Gaps = 16/339 (4%)
Query: 26 DWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REIL 84
DW R + F+ + IG+GTY VYKA+D TG++VALKKVR DN E E A REI
Sbjct: 718 DWGKRCVDKFDIIGIIGEGTYGQVYKARDKDTGELVALKKVRLDN-EKEGFPITAIREIK 776
Query: 85 VLRKLDHPNVVKLEGLVTSRMSC--------SLYLVFEYMEHDLAGLSAGQGVKFTEPQV 136
+LR+L H +VV ++ +VT + + YLVFEYM+HDL GL V F+E +
Sbjct: 777 ILRQLIHRSVVNMKEIVTDKQDALDFKKDKGAFYLVFEYMDHDLMGLLESGLVHFSEDHI 836
Query: 137 KCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRV 196
K FMKQL+ GLE+CH + LHRDIK SN+L++N G +K+ADFGLA YN + + T++V
Sbjct: 837 KSFMKQLMEGLEYCHKKNFLHRDIKCSNILLNNSGQIKLADFGLARLYNSEESRPYTNKV 896
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
+TLWYRPPELLLG Y ID+WS GCIL EL KPI E+ QL I +LCGSP
Sbjct: 897 ITLWYRPPELLLGEERYTPAIDVWSCGCILGELFTKKPIFQANLELAQLELISRLCGSPC 956
Query: 257 EEYWRK-HKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALN 315
W KLP KP++ Y+R + E F P ++L L+D +L +DP +R TA L
Sbjct: 957 PAVWPDVIKLPYFNTMKPKKQYRRRLREEFSFIPSAALDLLDHMLTLDPSKRCTAEQTLQ 1016
Query: 316 HEFF-TTEPYACEPSSLPKYPPSKELDVKMRDEEARRQK 353
+F E P LP + EL K R RRQ+
Sbjct: 1017 SDFLKDVELSKMAPPDLPHWQDCHELWSKKR----RRQR 1051
>CDL5_HUMAN (Q14004) Cell division cycle 2-like protein kinase 5 (EC
2.7.1.37) (Cholinesterase-related cell division
controller) (CDC2-related protein kinase 5)
Length = 418
Score = 267 bits (682), Expect = 5e-71
Identities = 143/312 (45%), Positives = 191/312 (60%), Gaps = 12/312 (3%)
Query: 26 DWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REIL 84
DW + F+ + IG+GTY VYKA+D TG++VALKKVR DN E E A REI
Sbjct: 82 DWGKLCVDKFDIIGIIGEGTYGQVYKARDKDTGEMVALKKVRLDN-EKEGFPITAIREIK 140
Query: 85 VLRKLDHPNVVKLEGLVTSRMSC--------SLYLVFEYMEHDLAGLSAGQGVKFTEPQV 136
+LR+L H +++ ++ +VT + + YLVFEYM+HDL GL V F E +
Sbjct: 141 ILRQLTHQSIINMKEIVTDKEDALDFKKDKGAFYLVFEYMDHDLMGLLESGLVHFYENHI 200
Query: 137 KCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRV 196
K FM+QL+ GL++CH + LHRDIK SN+L++N G +K+ADFGLA Y+ + + T++V
Sbjct: 201 KSFMRQLMEGLDYCHKKNFLHRDIKCSNILLNNRGQIKLADFGLARLYSSEESRPYTNKV 260
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
+TLWYRPPELLLG Y ID+WS GCIL EL KPI E+ QL I ++CGSP
Sbjct: 261 ITLWYRPPELLLGEERYTPAIDVWSCGCILGELFTKKPIFQANQELAQLELISRICGSPC 320
Query: 257 EEYWRK-HKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALN 315
W KLP KP++ Y+R + E F P ++L L D +LA+DP +R TA AL
Sbjct: 321 PAVWPDVIKLPYFNTMKPKKQYRRKLREEFVFIPAAALDLFDYMLALDPSKRCTAEQALQ 380
Query: 316 HEFF-TTEPYAC 326
EF EP C
Sbjct: 381 CEFLRDVEPSKC 392
>CDK9_SCHPO (Q96WV9) Serine/threonine-protein kinase cdk9 (EC
2.7.1.37) (Cyclin-dependent kinase cdk9)
Length = 591
Score = 261 bits (666), Expect = 4e-69
Identities = 155/410 (37%), Positives = 221/410 (53%), Gaps = 42/410 (10%)
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+ + K+G+GT+ VYK++ GK+ ALK++ + E E A REI +L+ + H N
Sbjct: 36 YHLMEKLGEGTFGEVYKSQRRKDGKVYALKRILM-HTEKEGFPITAIREIKILKSIKHEN 94
Query: 94 VVKLEGLVTSRMSC------SLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGL 147
++ L + R S+Y+V YM+HDL+GL VKFTEPQ+KC+MKQL +G
Sbjct: 95 IIPLSDMTVVRADKKHRRRGSIYMVTPYMDHDLSGLLENPSVKFTEPQIKCYMKQLFAGT 154
Query: 148 EHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYN------------PNKKQSMTSR 195
++ H + +LHRD+K +NLLIDN GILKIADFGLA P ++ T
Sbjct: 155 KYLHDQLILHRDLKAANLLIDNHGILKIADFGLARVITEESYANKNPGLPPPNRREYTGC 214
Query: 196 VVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSP 255
VVT WYR PELLLG Y ID+WS GCI+AE+ G+PI+ G ++++QL KIF+LCGSP
Sbjct: 215 VVTRWYRSPELLLGERRYTTAIDMWSVGCIMAEMYKGRPILQGSSDLDQLDKIFRLCGSP 274
Query: 256 AEEYWRK-HKLPNATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAAL 314
+ KLP + + R + F F L ++L ++PD R +AS AL
Sbjct: 275 TQATMPNWEKLPGCEGVRSFPSHPRTLETAFFTFGKEMTSLCGAILTLNPDERLSASMAL 334
Query: 315 NHEFFTTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKA--NAVDGAKRVRARE 372
HE+FTT PY PS L Y S E D + + R Q+ N A +G ++ R
Sbjct: 335 EHEYFTTPPYPANPSELQSYSASHEYDKRRK----REQRDANSHAFEQTANGKRQFRFMT 390
Query: 373 RGRAIP-----APEANAEIQ----------TNLDRWRVVTHANAKSKSEK 407
RG + P P N++ Q N+DR R V + + ++ K
Sbjct: 391 RGPSDPWYGIRRPNYNSQPQYQRGSYNREGGNMDRSRNVNYQPKRQQNFK 440
>CDKA_HUMAN (Q15131) Cell division protein kinase 10 (EC 2.7.1.37)
(Serine/threonine-protein kinase PISSLRE)
Length = 360
Score = 244 bits (623), Expect = 4e-64
Identities = 135/308 (43%), Positives = 183/308 (58%), Gaps = 4/308 (1%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKL 89
R FEKL +IG+GTY VY+A+D T +IVALKKVR D + REI +L +L
Sbjct: 34 RSVKEFEKLNRIGEGTYGIVYRARDTQTDEIVALKKVRMDKEKDGIPISSLREITLLLRL 93
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
HPN+V+L+ +V S++LV Y E DLA L F+E QVKC + Q+L GL++
Sbjct: 94 RHPNIVELKEVVVGNHLESIFLVMGYCEQDLASLLENMPTPFSEAQVKCIVLQVLRGLQY 153
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
H ++HRD+K SNLL+ ++G +K ADFGLA Y K MT +VVTLWYR PELLLG
Sbjct: 154 LHRNFIIHRDLKVSNLLMTDKGCVKTADFGLARAYGVPVK-PMTPKVVTLWYRAPELLLG 212
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRK-HKLPNA 268
T ID+W+ GCILAELLA +P++PG +E+ Q+ I +L G+P+E W KLP
Sbjct: 213 TTTQTTSIDMWAVGCILAELLAHRPLLPGTSEIHQIDLIVQLLGTPSENIWPGFSKLPLV 272
Query: 269 TIFK-PQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACE 327
+ +QPY + F + L L+ L DP +R TA L +F +P CE
Sbjct: 273 GQYSLRKQPYNN-LKHKFPWLSEAGLRLLHFLFMYDPKKRATAGDCLESSYFKEKPLPCE 331
Query: 328 PSSLPKYP 335
P +P +P
Sbjct: 332 PELMPTFP 339
>CDK9_CAEEL (P46551) Putative cell division protein kinase 9 homolog
(EC 2.7.1.37)
Length = 730
Score = 244 bits (623), Expect = 4e-64
Identities = 151/397 (38%), Positives = 222/397 (55%), Gaps = 19/397 (4%)
Query: 27 WTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILV 85
W + L +IG+GTY VYKA + +TG+ VALK+VR +N E E A REI +
Sbjct: 303 WYKTNLTHYTMLDQIGEGTYGQVYKAVNNLTGEQVALKRVRLEN-EKEGFPITAIREIKI 361
Query: 86 LRKLDHPNVVKLEGLVTSRMS--------CSLYLVFEYMEHDLAGL-SAGQGVKFTEPQV 136
LR+L H N+V+L +V +S + YLVFEY++HDL GL + + V F + Q+
Sbjct: 362 LRQLHHKNIVRLMDIVIDDISMDELKRTRANFYLVFEYVDHDLIGLLESKELVDFNKDQI 421
Query: 137 KCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRV 196
KQLL GL + H+ G LHRDIK SN+L++N+G LKIAD GLA + + + T+RV
Sbjct: 422 CSLFKQLLEGLAYIHNTGFLHRDIKCSNILVNNKGELKIADLGLARLWE-KESRLYTNRV 480
Query: 197 VTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPA 256
+TLWYRPPELLLG YG ID+WS GC+L EL KP+ G E QL I K+CGSP
Sbjct: 481 ITLWYRPPELLLGDERYGPAIDVWSTGCMLGELFTRKPLFNGNNEFGQLELISKVCGSPN 540
Query: 257 EEYWRK-HKLPNATIFKPQQPYKRCISETFKD-FPPSSLPLIDSLLAIDPDRRGTASAAL 314
+ W + +L F+ ++ Y+R I E F+ P ++ L+D +L ++P++R +A AL
Sbjct: 541 VDNWPELTELVGWNTFRMKRTYQRRIREEFEHIMPREAVDLLDKMLTLNPEKRISAKEAL 600
Query: 315 NHEFF-TTEPYACEPSSLPKYPPSKELDVKMRDEEARRQKALNGKANAVDGAKRVRARER 373
NH + + E +P LP++ E+ K + + AR + G + + +RA
Sbjct: 601 NHPWIRSLEHTTVQPLKLPQHQDCHEMWSKKQKKSARLGRQAEGSSGS---GHSIRATSH 657
Query: 374 GRAIPAPEANAEIQTNLDRWRVVTHANAKSKSEKFPP 410
RA P + ++N H ++ + PP
Sbjct: 658 PRA-PTQPSTTTTKSNGSSNHHHHHHHSHHHASSLPP 693
>CTK1_YEAST (Q03957) CTD kinase alpha subunit (EC 2.7.1.-) (CTD
kinase 58 kDa subunit) (CTDK-I alpha subunit)
Length = 528
Score = 236 bits (602), Expect = 1e-61
Identities = 138/332 (41%), Positives = 192/332 (57%), Gaps = 13/332 (3%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKL 89
R + + ++ ++G+GTY VYKAK+ T K+VALKK+R REI +L+
Sbjct: 178 RSTSVYLRIMQVGEGTYGKVYKAKNTNTEKLVALKKLRLQGEREGFPITSIREIKLLQSF 237
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
DHPNV ++ ++ ++Y++FEY ++DL+GL + V+ + Q K KQLL G+E+
Sbjct: 238 DHPNVSTIKEIMVESQK-TVYMIFEYADNDLSGLLLNKEVQISHSQCKHLFKQLLLGMEY 296
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
H +LHRD+KGSN+LIDN+G LKI DFGLA N + T+RV+TLWYRPPELLLG
Sbjct: 297 LHDNKILHRDVKGSNILIDNQGNLKITDFGLAR--KMNSRADYTNRVITLWYRPPELLLG 354
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPNA 268
T YG +D+W GC+L EL I G E+EQ+ IFK+ G+P W + +P
Sbjct: 355 TTNYGTEVDMWGCGCLLVELFNKTAIFQGSNELEQIESIFKIMGTPTINSWPTLYDMPWF 414
Query: 269 TIFKPQQ--PYKRCISETFKDFPPSS--LPLIDSLLAIDPDRRGTASAALNHEFFTTEPY 324
+ PQQ Y SE FK PSS L L +LL D +R +A+ AL ++F EP
Sbjct: 415 FMIMPQQTTKYVNNFSEKFKSVLPSSKCLQLAINLLCYDQTKRFSATEALQSDYFKEEPK 474
Query: 325 ACEPSSLPKYPPSKELDVKMRDEEARRQKALN 356
EP L E +VK+ AR+QK N
Sbjct: 475 P-EPLVLDGLVSCHEYEVKL----ARKQKRPN 501
>CDK9_HUMAN (P50750) Cell division protein kinase 9 (EC 2.7.1.37)
(Serine/threonine-protein kinase PITALRE) (C-2K)
Length = 372
Score = 236 bits (602), Expect = 1e-61
Identities = 141/306 (46%), Positives = 190/306 (62%), Gaps = 18/306 (5%)
Query: 33 NSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDH 91
+ +EKLAKIGQGT+ V+KA+ TG+ VALKKV +N E E A REI +L+ L H
Sbjct: 17 SKYEKLAKIGQGTFGEVFKARHRKTGQKVALKKVLMEN-EKEGFPITALREIKILQLLKH 75
Query: 92 PNVVKLEGLVTSRMS----C--SLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLS 145
NVV L + ++ S C S+YLVF++ EHDLAGL + VKFT ++K M+ LL+
Sbjct: 76 ENVVNLIEICRTKASPYNRCKGSIYLVFDFCEHDLAGLLSNVLVKFTLSEIKRVMQMLLN 135
Query: 146 GLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQS---MTSRVVTLWYR 202
GL + H +LHRD+K +N+LI +G+LK+ADFGLA ++ K T+RVVTLWYR
Sbjct: 136 GLYYIHRNKILHRDMKAANVLITRDGVLKLADFGLARAFSLAKNSQPNRYTNRVVTLWYR 195
Query: 203 PPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRK 262
PPELLLG YG IDLW AGCI+AE+ PIM TE QL I +LCGS E W
Sbjct: 196 PPELLLGERDYGPPIDLWGAGCIMAEMWTRSPIMQANTEQHQLALISQLCGSITPEVW-- 253
Query: 263 HKLPNATIFKPQQ---PYKRCISETFKDF--PPSSLPLIDSLLAIDPDRRGTASAALNHE 317
+ N +++ + KR + + K + P +L LID LL +DP +R + ALNH+
Sbjct: 254 PNVDNYELYEKLELVKGQKRKVKDRLKAYVRDPYALDLIDKLLVLDPAQRIDSDDALNHD 313
Query: 318 FFTTEP 323
FF ++P
Sbjct: 314 FFWSDP 319
>CDL1_HUMAN (P21127) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1) (CLK-1)
(CDK11) (p58 CLK-1)
Length = 795
Score = 227 bits (579), Expect = 5e-59
Identities = 130/340 (38%), Positives = 184/340 (53%), Gaps = 11/340 (3%)
Query: 2 LDPLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIV 61
L P+ + Q+ P +L A+ G R F+ L +I +GTY VY+AKD T +IV
Sbjct: 413 LSPIELKQE-LPKYLPALQG-------CRSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIV 464
Query: 62 ALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLA 121
ALK+++ + + REI + K HPN+V + +V +Y+V Y+EHDL
Sbjct: 465 ALKRLKMEKEKEGFPITSLREINTILKAQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLK 524
Query: 122 GLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLA 181
L F +VK M QLL G++H H +LHRD+K SNLL+ + GILK+ DFGLA
Sbjct: 525 SLMETMKQPFLPGEVKTLMIQLLRGVKHLHDNWILHRDLKTSNLLLSHAGILKVGDFGLA 584
Query: 182 TFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTE 241
Y K + T VVTLWYR PELLLGA Y +D+WS GCI ELL KP+ PG++E
Sbjct: 585 REYGSPLK-AYTPVVVTLWYRAPELLLGAKEYSTAVDMWSVGCIFGELLTQKPLFPGKSE 643
Query: 242 VEQLHKIFKLCGSPAEEYWRKH-KLPNA-TIFKPQQPYKRCISETFKDFPPSSLPLIDSL 299
++Q++K+FK G+P+E+ W + +LP + + PY L++
Sbjct: 644 IDQINKVFKDLGTPSEKIWPGYSELPAVKKMTFSEHPYNNLRKRFGALLSDQGFDLMNKF 703
Query: 300 LAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKE 339
L P RR +A L HE+F P +PS P +P E
Sbjct: 704 LTYFPGRRISAEDGLKHEYFRETPLPIDPSMFPTWPAKSE 743
>CDL1_RAT (P46892) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1)
Length = 436
Score = 226 bits (575), Expect = 1e-58
Identities = 131/340 (38%), Positives = 183/340 (53%), Gaps = 11/340 (3%)
Query: 2 LDPLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIV 61
L P+ + Q+ P +L A+ G R F+ L +I +GTY VY+AKD T +IV
Sbjct: 54 LSPIELKQE-LPKYLPALQG-------CRSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIV 105
Query: 62 ALKKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLA 121
ALK+++ + + REI + K HPN+V + +V +Y+V Y+EHDL
Sbjct: 106 ALKRLKMEKEKEGFPLTSIREINTILKAQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLK 165
Query: 122 GLSAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLA 181
L F +VK M QLLSG++H H +LHRD+K SNLL+ + GILK+ DFGLA
Sbjct: 166 SLMETMKQPFLPGEVKTLMIQLLSGVKHLHDNWILHRDLKTSNLLLTHAGILKVGDFGLA 225
Query: 182 TFYNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTE 241
Y K + T VVTLWYR PELLLGA Y D+WS GCI ELL KP+ PG+++
Sbjct: 226 REYGSPLK-AYTPVVVTLWYRAPELLLGAKEYSTACDMWSVGCIFGELLTQKPLFPGKSD 284
Query: 242 VEQLHKIFKLCGSPAEEYWRKH-KLPNA-TIFKPQQPYKRCISETFKDFPPSSLPLIDSL 299
++Q++KIFK G+P+E+ W + +LP + + PY L++
Sbjct: 285 IDQINKIFKDIGTPSEKIWPGYSELPAVKKMTFSELPYNNLRKRFGALLSDQGFDLMNKF 344
Query: 300 LAIDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKE 339
L P RR A L HE+F P +PS P +P E
Sbjct: 345 LTYYPGRRINAEDGLKHEYFRETPLPIDPSMFPTWPAKSE 384
>CDL1_MOUSE (P24788) PITSLRE serine/threonine-protein kinase CDC2L1
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 1)
Length = 434
Score = 223 bits (568), Expect = 9e-58
Identities = 125/312 (40%), Positives = 170/312 (54%), Gaps = 3/312 (0%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMAREILVLRKL 89
R F+ L +I +GTY VY+AKD T +IVALK+++ + + REI + K
Sbjct: 72 RSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVALKRLKMEKEKEGFPITSLREINTILKA 131
Query: 90 DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQVKCFMKQLLSGLEH 149
HPN+V + +V +Y+V Y+EHDL L F +VK M QLLSG++H
Sbjct: 132 QHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSLMETMKQPFLPGEVKTLMIQLLSGVKH 191
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
H +LHRD+K SNLL+ + GILK+ DFGLA Y K + T VVTLWYR PELLLG
Sbjct: 192 LHDNWILHRDLKTSNLLLTHAGILKVGDFGLAREYGSPLK-AYTPVVVTLWYRAPELLLG 250
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHK-LPNA 268
A Y D+WS GCI ELL KP+ PG+++++Q++KIFK GSP+E+ W + LP
Sbjct: 251 AKEYSTACDMWSVGCIFGELLTQKPLFPGKSDIDQINKIFKDLGSPSEKIWPGYNDLPAV 310
Query: 269 -TIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFFTTEPYACE 327
+ + PY L++ L P RR A L HE+F P +
Sbjct: 311 KKMTFSEIPYNNLRKRFGALLSDQGFDLMNKFLTYYPGRRINAEDGLKHEYFRETPLPID 370
Query: 328 PSSLPKYPPSKE 339
PS P +P E
Sbjct: 371 PSMFPTWPAKSE 382
>YP62_CAEEL (Q09437) Putative serine/threonine-protein kinase
B0495.2 (EC 2.7.1.37)
Length = 719
Score = 223 bits (567), Expect = 1e-57
Identities = 131/327 (40%), Positives = 192/327 (58%), Gaps = 27/327 (8%)
Query: 30 RRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRK 88
R + +E + ++ +GT+ VY+ KD T +IVALK+++ + E E A REI +L K
Sbjct: 351 RNIDEYECVNRVDEGTFGVVYRGKDKRTDEIVALKRLKMEK-EKEGFPITALREINMLLK 409
Query: 89 L-DHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL---SAGQGVKFTEPQVKCFMKQLL 144
+HPN+V ++ ++ +Y+ E++EHD+ L + + +F+ + K ++QLL
Sbjct: 410 AGNHPNIVNVKEILLGSNMDKIYMAMEFVEHDMKSLLDTMSRRNKRFSIGEQKTLLQQLL 469
Query: 145 SGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFY-NPNKKQSMTSRVVTLWYRP 203
SG+EH H +LHRD+K SNLL+ ++GILKIADFGLA Y +P KK TS VVTLWYR
Sbjct: 470 SGIEHMHKLWILHRDLKTSNLLMSHKGILKIADFGLAREYGDPLKK--FTSIVVTLWYRS 527
Query: 204 PELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKH 263
PELLLG Y +D+WS GCI+AE + KP+ PGR E+EQ+ KIF G+P E W
Sbjct: 528 PELLLGTRLYSTPVDMWSVGCIMAEFILLKPLFPGRGELEQIKKIFMEMGTPTESIW--- 584
Query: 264 KLPNAT-------IFKPQQPY----KRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASA 312
P T + + PY KR ++ + + L++ LL +DP R +A+
Sbjct: 585 --PGVTELDGWKALTFEKYPYNQLRKRFLAGRLLN--DTGFKLLNGLLTLDPKNRFSATQ 640
Query: 313 ALNHEFFTTEPYACEPSSLPKYPPSKE 339
AL+HE+FT EPY P P +P E
Sbjct: 641 ALDHEWFTEEPYPVPPEEFPTFPAKSE 667
>CDL2_HUMAN (Q9UQ88) PITSLRE serine/threonine-protein kinase CDC2L2
(EC 2.7.1.37) (Galactosyltransferase associated protein
kinase p58/GTA) (Cell division cycle 2-like 2) (CDK11)
Length = 780
Score = 223 bits (567), Expect = 1e-57
Identities = 128/338 (37%), Positives = 181/338 (52%), Gaps = 11/338 (3%)
Query: 4 PLSVNQQGWPSWLMAVAGEAIGDWTPRRANSFEKLAKIGQGTYSNVYKAKDLVTGKIVAL 63
P+ + Q+ P +L A+ G R F+ L +I +GTY VY+AKD T +IVAL
Sbjct: 400 PIELKQE-LPKYLPALQG-------CRSVEEFQCLNRIEEGTYGVVYRAKDKKTDEIVAL 451
Query: 64 KKVRFDNLEPESVKFMAREILVLRKLDHPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL 123
K+++ + + REI + K HPN+V + +V +Y+V Y+EHDL L
Sbjct: 452 KRLKMEKEKEGFPITSLREINTILKAQHPNIVTVREIVVGSNMDKIYIVMNYVEHDLKSL 511
Query: 124 SAGQGVKFTEPQVKCFMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATF 183
F +VK M QLL G++H H +LHRD+K SNLL+ + GILK+ DFGLA
Sbjct: 512 METMKQPFLPGEVKTLMIQLLRGVKHLHDNWILHRDLKTSNLLLSHAGILKVGDFGLARE 571
Query: 184 YNPNKKQSMTSRVVTLWYRPPELLLGATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVE 243
Y K + T VVT WYR PELLLGA Y +D+WS GCI ELL KP+ PG +E++
Sbjct: 572 YGSPLK-AYTPVVVTQWYRAPELLLGAKEYSTAVDMWSVGCIFGELLTQKPLFPGNSEID 630
Query: 244 QLHKIFKLCGSPAEEYWRKH-KLPNA-TIFKPQQPYKRCISETFKDFPPSSLPLIDSLLA 301
Q++K+FK G+P+E+ W + +LP + + PY L++ L
Sbjct: 631 QINKVFKELGTPSEKIWPGYSELPVVKKMTFSEHPYNNLRKRFGALLSDQGFDLMNKFLT 690
Query: 302 IDPDRRGTASAALNHEFFTTEPYACEPSSLPKYPPSKE 339
P RR +A L HE+F P +PS P +P E
Sbjct: 691 YFPGRRISAEDGLKHEYFRETPLPIDPSMFPTWPAKSE 728
>CDK2_CARAU (P43450) Cell division protein kinase 2 (EC 2.7.1.37)
Length = 298
Score = 223 bits (567), Expect = 1e-57
Identities = 128/293 (43%), Positives = 178/293 (60%), Gaps = 16/293 (5%)
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHP 92
SF+K+ KIG+GTY VYKAK+ VTG+ VALKK+R D E E V A REI +L++L+HP
Sbjct: 3 SFQKVEKIGEGTYGVVYKAKNKVTGETVALKKIRLDT-ETEGVPSTAIREISLLKELNHP 61
Query: 93 NVVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVK-FTEPQVKCFMKQLLSGLEHCH 151
N+VKL ++ + LYLVFE++ DL V + P VK ++ QLL GL CH
Sbjct: 62 NIVKLHDVIHTENK--LYLVFEFLHQDLKRFMDSSTVTGISLPLVKSYLFQLLQGLAFCH 119
Query: 152 SRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
S VLHRD+K NLLI+ +G +K+ADFGLA + + + T VVTLWYR PE+LLG
Sbjct: 120 SHRVLHRDLKPQNLLINAQGEIKLADFGLARAFGVPVR-TYTHEVVTLWYRAPEILLGCK 178
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATI- 270
+Y +D+WS GCI AE++ K + PG +E++QL +IF+ G+P E W P T
Sbjct: 179 YYSTAVDIWSLGCIFAEMITRKALFPGDSEIDQLFRIFRTLGTPDESIW-----PGVTSM 233
Query: 271 --FKPQQP--YKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
+KP P ++ +S+ L+ +L DP++R +A AL H FF
Sbjct: 234 PDYKPSFPKWARQDLSKVVPPLDEDGRDLLGQMLIYDPNKRISAKNALVHRFF 286
>CDK2_CRIGR (O55076) Cell division protein kinase 2 (EC 2.7.1.37)
Length = 298
Score = 222 bits (565), Expect = 2e-57
Identities = 131/296 (44%), Positives = 182/296 (61%), Gaps = 22/296 (7%)
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHP 92
+F+K+ KIG+GTY VYKAK+ +TG++VALKK+R D E E V A REI +L++L+HP
Sbjct: 3 NFQKVEKIGEGTYGVVYKAKNKLTGEVVALKKIRLDT-ETEGVPSTAIREISLLKELNHP 61
Query: 93 NVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL---SAGQGVKFTEPQVKCFMKQLLSGLEH 149
N+VKL ++ + LYLVFE++ DL SA G+ P +K ++ QLL GL
Sbjct: 62 NIVKLLDVIHTENK--LYLVFEFLHQDLKKFMDASAVTGIPL--PLIKSYLFQLLQGLAF 117
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
CHS VLHRD+K NLLI+ EG +K+ADFGLA + + + T VVTLWYR PE+LLG
Sbjct: 118 CHSHRVLHRDLKPQNLLINAEGSIKLADFGLARAFGVPVR-TYTHEVVTLWYRAPEILLG 176
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNAT 269
+Y +D+WS GCI AE++ + + PG +E++QL +IF+ G+P E W P T
Sbjct: 177 CKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVW-----PGVT 231
Query: 270 I---FKPQQPYKRCISETFKDFPP---SSLPLIDSLLAIDPDRRGTASAALNHEFF 319
+KP P K + K PP L+ +L DP++R +A AAL H FF
Sbjct: 232 SMPDYKPSFP-KWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFF 286
>CDK2_HUMAN (P24941) Cell division protein kinase 2 (EC 2.7.1.37)
(p33 protein kinase)
Length = 298
Score = 221 bits (564), Expect = 3e-57
Identities = 130/296 (43%), Positives = 182/296 (60%), Gaps = 22/296 (7%)
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHP 92
+F+K+ KIG+GTY VYKA++ +TG++VALKK+R D E E V A REI +L++L+HP
Sbjct: 3 NFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDT-ETEGVPSTAIREISLLKELNHP 61
Query: 93 NVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL---SAGQGVKFTEPQVKCFMKQLLSGLEH 149
N+VKL ++ + LYLVFE++ DL SA G+ P +K ++ QLL GL
Sbjct: 62 NIVKLLDVIHTENK--LYLVFEFLHQDLKKFMDASALTGIPL--PLIKSYLFQLLQGLAF 117
Query: 150 CHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
CHS VLHRD+K NLLI+ EG +K+ADFGLA + + + T VVTLWYR PE+LLG
Sbjct: 118 CHSHRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVR-TYTHEVVTLWYRAPEILLG 176
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNAT 269
+Y +D+WS GCI AE++ + + PG +E++QL +IF+ G+P E W P T
Sbjct: 177 CKYYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVW-----PGVT 231
Query: 270 I---FKPQQPYKRCISETFKDFPP---SSLPLIDSLLAIDPDRRGTASAALNHEFF 319
+KP P K + K PP L+ +L DP++R +A AAL H FF
Sbjct: 232 SMPDYKPSFP-KWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFF 286
>CDK2_RAT (Q63699) Cell division protein kinase 2 (EC 2.7.1.37)
Length = 298
Score = 221 bits (563), Expect = 3e-57
Identities = 129/294 (43%), Positives = 180/294 (60%), Gaps = 18/294 (6%)
Query: 34 SFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHP 92
+F+K+ KIG+GTY VYKAK+ +TG++VALKK+R D E E V A REI +L++L+HP
Sbjct: 3 NFQKVEKIGEGTYGVVYKAKNKLTGEVVALKKIRLDT-ETEGVPSTAIREISLLKELNHP 61
Query: 93 NVVKLEGLVTSRMSCSLYLVFEYMEHDLAG-LSAGQGVKFTEPQVKCFMKQLLSGLEHCH 151
N+VKL ++ + LYLVFE++ DL + A P +K ++ QLL GL CH
Sbjct: 62 NIVKLLDVIHTENK--LYLVFEFLHQDLKKFMDASALTGLPLPLIKSYLFQLLQGLAFCH 119
Query: 152 SRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
S VLHRD+K NLLI+ EG +K+ADFGLA + + + T VVTLWYR PE+LLG
Sbjct: 120 SHRVLHRDLKPQNLLINAEGSIKLADFGLARAFGVPVR-TYTHEVVTLWYRAPEILLGCK 178
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRKHKLPNATI- 270
+Y +D+WS GCI AE++ + + PG +E++QL +IF+ G+P E W P T
Sbjct: 179 YYSTAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVW-----PGVTSM 233
Query: 271 --FKPQQPYKRCISETFKDFPP---SSLPLIDSLLAIDPDRRGTASAALNHEFF 319
+KP P K + K PP L+ +L DP++R +A AAL H FF
Sbjct: 234 PDYKPSFP-KWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFF 286
>CC2A_ARATH (P24100) Cell division control protein 2 homolog A (EC
2.7.1.37)
Length = 294
Score = 221 bits (562), Expect = 4e-57
Identities = 127/292 (43%), Positives = 183/292 (62%), Gaps = 11/292 (3%)
Query: 33 NSFEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDH 91
+ +EK+ KIG+GTY VYKA+D VT + +ALKK+R + E E V A REI +L+++ H
Sbjct: 2 DQYEKVEKIGEGTYGVVYKARDKVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 60
Query: 92 PNVVKLEGLVTSRMSCSLYLVFEYMEHDLAG-LSAGQGVKFTEPQVKCFMKQLLSGLEHC 150
N+VKL+ +V S LYLVFEY++ DL + + +K ++ Q+L G+ +C
Sbjct: 61 SNIVKLQDVVHSEKR--LYLVFEYLDLDLKKHMDSTPDFSKDLHMIKTYLYQILRGIAYC 118
Query: 151 HSRGVLHRDIKGSNLLIDNE-GILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLG 209
HS VLHRD+K NLLID LK+ADFGLA + + + T VVTLWYR PE+LLG
Sbjct: 119 HSHRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLG 177
Query: 210 ATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWR-KHKLPNA 268
+ Y +D+WS GCI AE+++ KP+ PG +E++QL KIF++ G+P E+ WR LP+
Sbjct: 178 SHHYSTPVDIWSVGCIFAEMISQKPLFPGDSEIDQLFKIFRIMGTPYEDTWRGVTSLPDY 237
Query: 269 TIFKPQQPYKRCISETF-KDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
P+ +K ETF + P + L+ +L +DP +R A AAL HE+F
Sbjct: 238 KSAFPK--WKPTDLETFVPNLDPDGVDLLSKMLLMDPTKRINARAALEHEYF 287
>CC2A_ANTMA (Q38772) Cell division control protein 2 homolog A (EC
2.7.1.37)
Length = 294
Score = 221 bits (562), Expect = 4e-57
Identities = 127/289 (43%), Positives = 181/289 (61%), Gaps = 9/289 (3%)
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+EK+ KIG+GTY VYKA+D VT + +ALKK+R + E E V A REI +L+++ H N
Sbjct: 4 YEKVEKIGEGTYGVVYKARDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQHGN 62
Query: 94 VVKLEGLVTSRMSCSLYLVFEYMEHDLAGLSAGQGVKFTEPQ-VKCFMKQLLSGLEHCHS 152
+V+L+ +V S LYLVFEY++ DL +P+ VK F+ Q+L G+ +CHS
Sbjct: 63 IVRLQDVVHSEKR--LYLVFEYLDLDLKKHMDSCPEFSQDPRLVKMFLYQILRGIAYCHS 120
Query: 153 RGVLHRDIKGSNLLIDNE-GILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
VLHRD+K NLLID LK+ADFGLA + + + T VVTLWYR PE+LLG+
Sbjct: 121 HRVLHRDLKPQNLLIDRRTNALKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLGSR 179
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPNATI 270
Y +D+WS GCI AE++ +P+ PG +E+++L KIF++ G+P EE W LP+
Sbjct: 180 HYSTPVDVWSVGCIFAEMVNQRPLFPGDSEIDELFKIFRVMGTPNEETWPGVTSLPDFKS 239
Query: 271 FKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
P+ P K ++ + S L L+D +L +DP +R TA AL HE+F
Sbjct: 240 AFPKWPAKE-LAAVVPNLDASGLDLLDKMLRLDPSKRITARNALQHEYF 287
>CC28_CANAL (P43063) Cell division control protein 28 (EC 2.7.1.37)
Length = 317
Score = 221 bits (562), Expect = 4e-57
Identities = 126/292 (43%), Positives = 181/292 (61%), Gaps = 9/292 (3%)
Query: 33 NSFEKLAKIGQGTYSNVYKAKDLV-TGKIVALKKVRFDNLEPESVKFMA-REILVLRKLD 90
+ +++ K+G+GTY VYKA D ++VALKK+R ++ E E V A REI +L+++
Sbjct: 5 SDYQRQEKVGEGTYGVVYKALDTKHNNRVVALKKIRLES-EDEGVPSTAIREISLLKEMK 63
Query: 91 HPNVVKLEGLVTSRMSCSLYLVFEYMEHDLAGL--SAGQGVKFTEPQVKCFMKQLLSGLE 148
N+V+L ++ S S LYLVFE+++ DL S QGV +K FM QL+ G++
Sbjct: 64 DDNIVRLYDIIHSD-SHKLYLVFEFLDLDLKKYMESIPQGVGLGANMIKRFMNQLIRGIK 122
Query: 149 HCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLL 208
HCHS VLHRD+K NLLID EG LK+ADFGLA + + + T VVTLWYR PE+LL
Sbjct: 123 HCHSHRVLHRDLKPQNLLIDKEGNLKLADFGLARAFGVPLR-AYTHEVVTLWYRAPEILL 181
Query: 209 GATFYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYWRK-HKLPN 267
G Y G+D+WS GCI AE+ KP+ PG +E++++ +IF++ G+P EE W + LP+
Sbjct: 182 GGKQYSTGVDMWSVGCIFAEMCNRKPLFPGDSEIDEIFRIFRILGTPNEEIWPDVNYLPD 241
Query: 268 ATIFKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
PQ K+ +SE + + L+D +L DP RR +A AL H +F
Sbjct: 242 FKSSFPQWK-KKPLSEAVPSLDANGIDLLDQMLVYDPSRRISAKRALIHPYF 292
>CC22_MEDSA (Q05006) Cell division control protein 2 homolog 2 (EC
2.7.1.37)
Length = 294
Score = 221 bits (562), Expect = 4e-57
Identities = 124/289 (42%), Positives = 181/289 (61%), Gaps = 9/289 (3%)
Query: 35 FEKLAKIGQGTYSNVYKAKDLVTGKIVALKKVRFDNLEPESVKFMA-REILVLRKLDHPN 93
+EK+ KIG+GTY VYKA+D T + +ALKK+R + E E V A REI +L+++ H N
Sbjct: 4 YEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQHRN 62
Query: 94 VVKLEGLVTSRMSCSLYLVFEYMEHDLAG-LSAGQGVKFTEPQVKCFMKQLLSGLEHCHS 152
+V+L+ +V S LYLVFEY++ DL + + + Q+K F+ Q+L G+ +CHS
Sbjct: 63 IVRLQDVVHSEKR--LYLVFEYLDLDLKKFMDSSPEFAKDQRQIKMFLYQILCGIAYCHS 120
Query: 153 RGVLHRDIKGSNLLID-NEGILKIADFGLATFYNPNKKQSMTSRVVTLWYRPPELLLGAT 211
VLHRD+K NLLID + +K+ADFGLA + + + T VVTLWYR PE+LLG+
Sbjct: 121 HRVLHRDLKPQNLLIDRSSNAVKLADFGLARAFGIPVR-TFTHEVVTLWYRAPEILLGSR 179
Query: 212 FYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPAEEYW-RKHKLPNATI 270
Y +D+WS GCI AE++ +P+ PG +E+++L KIF++ G+P EE W LP+
Sbjct: 180 HYSTPVDVWSVGCIFAEMINQRPLFPGDSEIDELFKIFRITGTPNEETWPGVTSLPDFKS 239
Query: 271 FKPQQPYKRCISETFKDFPPSSLPLIDSLLAIDPDRRGTASAALNHEFF 319
P+ P K ++ + P+ L L+ S +DP RR TA AL HE+F
Sbjct: 240 AFPKWPAKDLATQV-PNLEPAGLDLLSSTCRLDPTRRITARGALEHEYF 287
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 62,360,427
Number of Sequences: 164201
Number of extensions: 2777131
Number of successful extensions: 11172
Number of sequences better than 10.0: 1687
Number of HSP's better than 10.0 without gapping: 1551
Number of HSP's successfully gapped in prelim test: 136
Number of HSP's that attempted gapping in prelim test: 6466
Number of HSP's gapped (non-prelim): 2000
length of query: 498
length of database: 59,974,054
effective HSP length: 114
effective length of query: 384
effective length of database: 41,255,140
effective search space: 15841973760
effective search space used: 15841973760
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146747.6


