

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146721.7 - phase: 0
(188 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
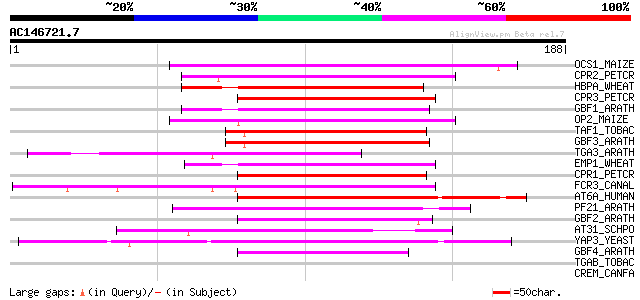
Sequences producing significant alignments: (bits) Value
OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1) 70 2e-12
CPR2_PETCR (Q99090) Light-inducible protein CPRF-2 69 8e-12
HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specifi... 57 3e-08
CPR3_PETCR (Q99091) Light-inducible protein CPRF-3 56 4e-08
GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41) 55 1e-07
OP2_MAIZE (P12959) Opaque-2 regulatory protein 54 2e-07
TAF1_TOBAC (Q99142) Transcriptional activator TAF-1 (Fragment) 53 4e-07
GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55) 52 1e-06
TGA3_ARATH (Q39234) Transcription factor TGA3 (AtbZIP22) 49 7e-06
EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein 49 7e-06
CPR1_PETCR (Q99089) Common plant regulatory factor CPRF-1 48 2e-05
FCR3_CANAL (Q8X229) Fluconazole resistance protein 3 47 3e-05
AT6A_HUMAN (P18850) Cyclic-AMP-dependent transcription factor AT... 46 4e-05
PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59) 45 1e-04
GBF2_ARATH (P42775) G-box binding factor 2 (AtbZIP54) 44 2e-04
AT31_SCHPO (Q09771) Transcription factor atf31 44 3e-04
YAP3_YEAST (P38749) AP-1 like transcription factor YAP3 43 4e-04
GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40) 42 6e-04
TGAB_TOBAC (P14233) TGACG-sequence specific DNA-binding protein ... 40 0.002
CREM_CANFA (P79145) cAMP responsive element modulator 40 0.002
>OCS1_MAIZE (P24068) Ocs-element binding factor 1 (OCSBF-1)
Length = 151
Score = 70.5 bits (171), Expect = 2e-12
Identities = 42/128 (32%), Positives = 67/128 (51%), Gaps = 10/128 (7%)
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNE 114
SS +S + + + + R+ +R +SNRESARRSR+RKQ+HLDEL +V L+ +
Sbjct: 3 SSSLSPTAGRTSGSDGDSAADTHRREKRRLSNRESARRSRLRKQQHLDELVQEVARLQAD 62
Query: 115 NHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFS---------- 164
N ++ + ++ + +V QEN L+ A EL ++ + L+ FS
Sbjct: 63 NARVAARARDIASQYTRVEQENTVLRARAAELGDRLRSVNEVLRLVEEFSGVAMDIQEEM 122
Query: 165 PLDDTYLR 172
P DD LR
Sbjct: 123 PADDPLLR 130
>CPR2_PETCR (Q99090) Light-inducible protein CPRF-2
Length = 401
Score = 68.6 bits (166), Expect = 8e-12
Identities = 40/101 (39%), Positives = 59/101 (57%), Gaps = 8/101 (7%)
Query: 59 SSNSTSDEADE--------QNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLW 110
SS SD+ DE +N + ++ RRM+SNRESARRSR RKQ H+ EL +QV
Sbjct: 173 SSRDHSDDDDELEGETETTRNGDPSDAKRVRRMLSNRESARRSRRRKQAHMTELETQVSQ 232
Query: 111 LRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
LR EN L+++L +S+ ++ +N LK + +R +K
Sbjct: 233 LRVENSSLLKRLTDISQRYNDAAVDNRVLKADIETMRAKVK 273
>HBPA_WHEAT (P23922) Transcription factor HBP-1a (Histone-specific
transcription factor HBP1)
Length = 349
Score = 57.0 bits (136), Expect = 3e-08
Identities = 29/82 (35%), Positives = 52/82 (63%), Gaps = 5/82 (6%)
Query: 59 SSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
S ++ ++ DE+ L +K +R +SNRESARRSR+RKQ +EL + L++EN L
Sbjct: 240 SGSARGEQWDEREL-----KKQKRKLSNRESARRSRLRKQAECEELGQRAEALKSENSSL 294
Query: 119 IEKLNHVSENHDQVVQENAQLK 140
+L+ + + +++++ +N LK
Sbjct: 295 RIELDRIKKEYEELLSKNTSLK 316
>CPR3_PETCR (Q99091) Light-inducible protein CPRF-3
Length = 296
Score = 56.2 bits (134), Expect = 4e-08
Identities = 32/67 (47%), Positives = 41/67 (60%)
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ RR SNRESARRSR+RKQ DEL ++ L EN L + L +SE +V EN
Sbjct: 198 KRQRRKQSNRESARRSRLRKQAKSDELQERLDNLSKENRILRKNLQRISEACAEVTSENH 257
Query: 138 QLKEEAL 144
+KEE L
Sbjct: 258 SIKEELL 264
>GBF1_ARATH (P42774) G-box binding factor 1 (AtbZIP41)
Length = 315
Score = 54.7 bits (130), Expect = 1e-07
Identities = 32/84 (38%), Positives = 49/84 (58%), Gaps = 5/84 (5%)
Query: 59 SSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
SS + DE+ L ++ +R SNRESARRSR+RKQ ++L +V L NEN L
Sbjct: 210 SSQAGVPVKDEREL-----KRQKRKQSNRESARRSRLRKQAECEQLQQRVESLSNENQSL 264
Query: 119 IEKLNHVSENHDQVVQENAQLKEE 142
++L +S D++ EN +++E
Sbjct: 265 RDELQRLSSECDKLKSENNSIQDE 288
>OP2_MAIZE (P12959) Opaque-2 regulatory protein
Length = 453
Score = 54.3 bits (129), Expect = 2e-07
Identities = 35/102 (34%), Positives = 54/102 (52%), Gaps = 5/102 (4%)
Query: 55 SSCISSNSTSDEADEQNLSLIN-----ERKHRRMISNRESARRSRMRKQKHLDELWSQVL 109
SS S SDE + + ++ E + R+ SNRESARRSR RK HL EL QV
Sbjct: 199 SSSSRDPSPSDEDMDGEVEILGFKMPTEERVRKKESNRESARRSRYRKAAHLKELEDQVA 258
Query: 110 WLRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
L+ EN L+ ++ +++ ++ +N L+ + LR +K
Sbjct: 259 QLKAENSCLLRRIAALNQKYNDANVDNRVLRADMETLRAKVK 300
>TAF1_TOBAC (Q99142) Transcriptional activator TAF-1 (Fragment)
Length = 265
Score = 53.1 bits (126), Expect = 4e-07
Identities = 33/71 (46%), Positives = 44/71 (61%), Gaps = 3/71 (4%)
Query: 74 LINER---KHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHD 130
L NER + +R SNRESARRSR+RKQ +EL +V L EN L ++N + EN +
Sbjct: 189 LQNERELKREKRKQSNRESARRSRLRKQAEAEELAIRVQSLTAENMTLKSEINKLMENSE 248
Query: 131 QVVQENAQLKE 141
++ ENA L E
Sbjct: 249 KLKLENAALME 259
>GBF3_ARATH (P42776) G-box binding factor 3 (AtbZIP55)
Length = 382
Score = 51.6 bits (122), Expect = 1e-06
Identities = 33/72 (45%), Positives = 44/72 (60%), Gaps = 3/72 (4%)
Query: 74 LINER---KHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHD 130
L NER + RR SNRESARRSR+RKQ +EL +V L EN L +LN ++E D
Sbjct: 254 LQNERELKRERRKQSNRESARRSRLRKQAETEELARKVEALTAENMALRSELNQLNEKSD 313
Query: 131 QVVQENAQLKEE 142
++ NA L ++
Sbjct: 314 KLRGANATLLDK 325
>TGA3_ARATH (Q39234) Transcription factor TGA3 (AtbZIP22)
Length = 384
Score = 48.9 bits (115), Expect = 7e-06
Identities = 33/116 (28%), Positives = 55/116 (46%), Gaps = 12/116 (10%)
Query: 7 SPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISSNSTSDE 66
SP+ + NN+ N N +NQ L + D N + + +NS E
Sbjct: 33 SPFKSDINNITSNQN---------NNQSSSTTLEVDARPEADDNNRVNYTSVYNNSLEAE 83
Query: 67 A---DEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
++Q+ IN++ RR+ NRE+AR+SR+RK+ H+ +L L L +L+
Sbjct: 84 PSSNNDQDEDRINDKMKRRLAQNREAARKSRLRKKAHVQQLEESRLKLSQLEQELV 139
>EMP1_WHEAT (P25032) DNA-binding EMBP-1 protein
Length = 354
Score = 48.9 bits (115), Expect = 7e-06
Identities = 31/85 (36%), Positives = 49/85 (57%), Gaps = 5/85 (5%)
Query: 60 SNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
SN++ + DE+ L ++ RR SNRESARRSR+RKQ+ +EL +V L N L
Sbjct: 239 SNASLSQMDEREL-----KRERRKQSNRESARRSRLRKQQECEELAQKVSELTAANGTLR 293
Query: 120 EKLNHVSENHDQVVQENAQLKEEAL 144
+L+ + ++ + EN +L + L
Sbjct: 294 SELDQLKKDCKTMETENKKLMGKIL 318
>CPR1_PETCR (Q99089) Common plant regulatory factor CPRF-1
Length = 411
Score = 47.8 bits (112), Expect = 2e-05
Identities = 26/64 (40%), Positives = 41/64 (63%)
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ RR SNRESARRSR+RKQ +EL +V L EN L ++N ++ +++ +N+
Sbjct: 271 KRERRKQSNRESARRSRLRKQAEAEELAIKVDSLTAENMALKAEINRLTLTAEKLTNDNS 330
Query: 138 QLKE 141
+L E
Sbjct: 331 RLLE 334
>FCR3_CANAL (Q8X229) Fluconazole resistance protein 3
Length = 399
Score = 47.0 bits (110), Expect = 3e-05
Identities = 42/181 (23%), Positives = 78/181 (42%), Gaps = 38/181 (20%)
Query: 2 LPSNTSPYLQNYNNMIQ------------------------NNNISSYQLQKFSNQIY-- 35
+P+ PY+ N NN + +NN +S +K SN+ +
Sbjct: 98 MPTTQHPYITNTNNHLSYSNSSEEFSPIGNNMSPDSTGGANSNNFTSGNKRKASNESFSP 157
Query: 36 --GNYLNTPHQQFPDFNYSPQSSCISSNSTSDEA-DEQNLSLI---------NERKHRRM 83
G++ T + N + +SS SS+ + + DE++ +LI E + +R
Sbjct: 158 LSGHHYGTESGNNNNNNGTSRSSQSSSHKSRKKLLDEKDAALIARDDSELTEEELQMKRK 217
Query: 84 ISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENAQLKEEA 143
NR + R R RK+ L EL +++L E +L+++L + + + + EN LK
Sbjct: 218 AQNRAAQRAFRERKESKLKELEAKLLASEEERQKLLDELEQIKKQNISIATENEILKHNG 277
Query: 144 L 144
+
Sbjct: 278 M 278
>AT6A_HUMAN (P18850) Cyclic-AMP-dependent transcription factor ATF-6
alpha (Activating transcription factor 6 alpha)
(ATF6-alpha)
Length = 670
Score = 46.2 bits (108), Expect = 4e-05
Identities = 31/98 (31%), Positives = 60/98 (60%), Gaps = 3/98 (3%)
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
R+ +RMI NRESA +SR +K++++ L +++ +EN QL ++ + D+VV EN
Sbjct: 308 RRQQRMIKNRESACQSRKKKKEYMLGLEARLKAALSENEQLKKENGTLKRQLDEVVSENQ 367
Query: 138 QLKEEALELRQMIKDMQIHSPLIPSFSPLDDTYLRDDS 175
+LK + + R+++ M + + +I ++ P+ + L DS
Sbjct: 368 RLKVPSPK-RRVVCVMIVLAFIILNYGPM--SMLEQDS 402
>PF21_ARATH (Q04088) Possible transcription factor PosF21 (AtbZIP59)
Length = 398
Score = 44.7 bits (104), Expect = 1e-04
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 5/101 (4%)
Query: 56 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
S I + + L+LI+ ++ +R+ +NR+SA RS+ RK +++ EL +V L+ E
Sbjct: 181 SAIDAKKSMSATKLAELALIDPKRAKRIWANRQSAARSKERKTRYIFELERKVQTLQTEA 240
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIH 156
L +L + + + + EN +LK LR + Q+H
Sbjct: 241 TTLSAQLTLLQRDTNGLTVENNELK-----LRLQTMEQQVH 276
>GBF2_ARATH (P42775) G-box binding factor 2 (AtbZIP54)
Length = 360
Score = 43.9 bits (102), Expect = 2e-04
Identities = 28/70 (40%), Positives = 41/70 (58%), Gaps = 4/70 (5%)
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENA 137
++ +R SNRESARRSR+RKQ ++L +V L EN L KL ++ +++ EN
Sbjct: 251 KREKRKQSNRESARRSRLRKQAETEQLSVKVDALVAENMSLRSKLGQLNNESEKLRLENE 310
Query: 138 ----QLKEEA 143
QLK +A
Sbjct: 311 AILDQLKAQA 320
>AT31_SCHPO (Q09771) Transcription factor atf31
Length = 209
Score = 43.5 bits (101), Expect = 3e-04
Identities = 29/121 (23%), Positives = 53/121 (42%), Gaps = 21/121 (17%)
Query: 37 NYLNTPHQQFPDFNYSPQSSCIS-------SNSTSDEADEQNLSLINERKHRRMISNRES 89
N + + ++ S S IS S D DE L ++ NR++
Sbjct: 75 NIRRSDFDELSEYTASKSPSIISEASHNSPSRELDDSGDENTSKLTGTKQSMLKARNRQA 134
Query: 90 ARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQM 149
A++ R++K+K+L L QV + +EN +L++ N L+EE ++LR +
Sbjct: 135 AQKCRIKKKKYLQTLQDQVNYYTSENKELLQSAN--------------DLREEIIKLRTL 180
Query: 150 I 150
+
Sbjct: 181 V 181
>YAP3_YEAST (P38749) AP-1 like transcription factor YAP3
Length = 330
Score = 43.1 bits (100), Expect = 4e-04
Identities = 37/172 (21%), Positives = 77/172 (44%), Gaps = 9/172 (5%)
Query: 4 SNTSPYLQNYNNMIQNNNISSYQLQKFSNQIYGNYL-----NTPHQQFPDFNYSPQSSCI 58
SN LQN + + +++ + QL S Y Y+ N + + + NY +++ I
Sbjct: 69 SNNDLLLQNNISFPKGSDLQAIQLTPISGD-YSTYVMADNNNNDNDSYSNTNYFSKNNGI 127
Query: 59 SSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQL 118
S +S S N ++ ++ K ++ NR + + R RK+ + EL ++L L
Sbjct: 128 SPSSRSPSV-AHNENVPDDSKAKKKAQNRAAQKAFRERKEARMKELQDKLLESERNRQSL 186
Query: 119 IEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDDTY 170
++++ + + + ++ EN L E KD++ + SF D+ +
Sbjct: 187 LKEIEELRKANTEINAENRLLLRSGNE--NFSKDIEDDTNYKYSFPTKDEFF 236
>GBF4_ARATH (P42777) G-box binding factor 4 (AtbZIP40)
Length = 270
Score = 42.4 bits (98), Expect = 6e-04
Identities = 22/58 (37%), Positives = 35/58 (59%)
Query: 78 RKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQE 135
++ +RMI NRESA RSR RKQ + EL + L EN QL++++ ++ + + E
Sbjct: 189 QRQKRMIKNRESAARSRERKQAYQVELETLAAKLEEENEQLLKEIEESTKERYKKLME 246
>TGAB_TOBAC (P14233) TGACG-sequence specific DNA-binding protein
TGA-1B (TGA1b) (HSBF) (Fragment)
Length = 242
Score = 40.4 bits (93), Expect = 0.002
Identities = 28/85 (32%), Positives = 45/85 (52%), Gaps = 16/85 (18%)
Query: 58 ISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQ 117
+S N +DE +E+K R++ NRESA+ SR RK+ +++EL +V + H
Sbjct: 174 LSDNVNNDE---------DEKKRARLVRNRESAQLSRQRKKHYVEELEDKVRIM----HS 220
Query: 118 LIEKLNHVSENHDQVVQENAQLKEE 142
I+ LN ++ ENA LK +
Sbjct: 221 TIQDLN---AKVAYIIAENATLKTQ 242
>CREM_CANFA (P79145) cAMP responsive element modulator
Length = 344
Score = 40.4 bits (93), Expect = 0.002
Identities = 21/75 (28%), Positives = 43/75 (57%)
Query: 53 PQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLR 112
PQ ++++ S + +Q ++ R++ NRE+AR R +K++++ L ++V L
Sbjct: 263 PQGVVMAASPGSLHSPQQLAEEATRKRELRLMKNREAARECRRKKKEYVKCLENRVAVLE 322
Query: 113 NENHQLIEKLNHVSE 127
N+N LIE+L + +
Sbjct: 323 NQNKTLIEELKALKD 337
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.126 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,798,643
Number of Sequences: 164201
Number of extensions: 858204
Number of successful extensions: 4020
Number of sequences better than 10.0: 264
Number of HSP's better than 10.0 without gapping: 97
Number of HSP's successfully gapped in prelim test: 169
Number of HSP's that attempted gapping in prelim test: 3789
Number of HSP's gapped (non-prelim): 400
length of query: 188
length of database: 59,974,054
effective HSP length: 104
effective length of query: 84
effective length of database: 42,897,150
effective search space: 3603360600
effective search space used: 3603360600
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146721.7


