

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.10 - phase: 0
(146 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
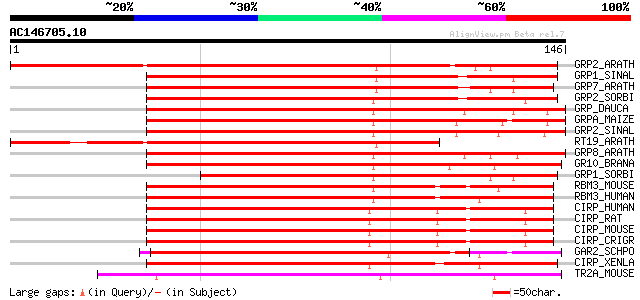
Score E
Sequences producing significant alignments: (bits) Value
GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2, mitochon... 187 9e-48
GRP1_SINAL (P49310) Glycine-rich RNA-binding protein GRP1A 129 2e-30
GRP7_ARATH (Q03250) Glycine-rich RNA-binding protein 7 126 1e-29
GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2 126 2e-29
GRP_DAUCA (Q03878) Glycine-rich RNA-binding protein 125 4e-29
GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic acid-indu... 124 5e-29
GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A 124 5e-29
RT19_ARATH (P39697) 40S ribosomal protein S19, mitochondrial pre... 120 8e-28
GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1 pro... 120 1e-27
GR10_BRANA (Q05966) Glycine-rich RNA-binding protein 10 115 3e-26
GRP1_SORBI (Q99069) Glycine-rich RNA-binding protein 1 (Fragment) 113 2e-25
RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding ... 103 1e-22
RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding ... 103 2e-22
CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein (Glycine-... 96 3e-20
CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-ri... 95 6e-20
CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein (Glycine-... 95 6e-20
CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein (Glycine-... 95 6e-20
GAR2_SCHPO (P41891) Protein gar2 94 1e-19
CIRP_XENLA (O93235) Cold-inducible RNA-binding protein (Glycine-... 93 2e-19
TR2A_MOUSE (Q6PFR5) Transformer-2 protein homolog (TRA-2 alpha) 92 4e-19
>GRP2_ARATH (Q9SVM8) Glycine-rich RNA-binding protein 2,
mitochondrial precursor (AtGRP2)
Length = 158
Score = 187 bits (474), Expect = 9e-48
Identities = 100/149 (67%), Positives = 117/149 (78%), Gaps = 7/149 (4%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAFCNK+G LLRQ + + PV+SML LR MS+ KLFIGGLS+ DD SLRDAF +G
Sbjct: 1 MAFCNKLGGLLRQNISSNGNVPVTSMLGSLRLMST-KLFIGGLSWGTDDASLRDAFAHFG 59
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSALS-MDGQDLNGRNIRVSYANDRQAGGP 119
DVV+A+VI DRETGRSRGFGFVNF E +AT+A+S MDG++LNGR+IRV+ ANDR + P
Sbjct: 60 DVVDAKVIVDRETGRSRGFGFVNFNDEGAATAAISEMDGKELNGRHIRVNPANDRPS-AP 118
Query: 120 RP---GGGY-GGGGGYSGGGGGYSGGGGG 144
R GGGY GGGGGY GGGGGY GGGGG
Sbjct: 119 RAYGGGGGYSGGGGGYGGGGGGYGGGGGG 147
>GRP1_SINAL (P49310) Glycine-rich RNA-binding protein GRP1A
Length = 166
Score = 129 bits (325), Expect = 2e-30
Identities = 64/110 (58%), Positives = 84/110 (76%), Gaps = 4/110 (3%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS- 95
+ F+GGL++ DD++L AF+ YG+V+++++I DRETGRSRGFGFV F E+S A+
Sbjct: 9 RCFVGGLAWATDDRALETAFSQYGEVLDSKIINDRETGRSRGFGFVTFKDEKSMKDAIEG 68
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGY-SGGGGGYSGGGGG 144
M+GQDL+GR+I V+ A R +GG GGG GGGGGY SGGGGGY GGGGG
Sbjct: 69 MNGQDLDGRSITVNEAQSRGSGG--GGGGRGGGGGYRSGGGGGYGGGGGG 116
>GRP7_ARATH (Q03250) Glycine-rich RNA-binding protein 7
Length = 176
Score = 126 bits (317), Expect = 1e-29
Identities = 64/111 (57%), Positives = 84/111 (75%), Gaps = 7/111 (6%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALS- 95
+ F+GGL++ DD++L AF YGDV+++++I DRETGRSRGFGFV F E++ A+
Sbjct: 9 RCFVGGLAWATDDRALETAFAQYGDVIDSKIINDRETGRSRGFGFVTFKDEKAMKDAIEG 68
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGY--GGGGGY-SGGGGGYSGGGG 143
M+GQDL+GR+I V+ A R +GG GGG+ GGGGGY SGGGGGYSGGGG
Sbjct: 69 MNGQDLDGRSITVNEAQSRGSGG---GGGHRGGGGGGYRSGGGGGYSGGGG 116
Score = 39.3 bits (90), Expect = 0.003
Identities = 18/30 (60%), Positives = 20/30 (66%)
Query: 109 SYANDRQAGGPRPGGGYGGGGGYSGGGGGY 138
SY R+ GG GGG GGG G SGGGGG+
Sbjct: 147 SYGGGRREGGGGYGGGEGGGYGGSGGGGGW 176
>GRP2_SORBI (Q99070) Glycine-rich RNA-binding protein 2
Length = 168
Score = 126 bits (316), Expect = 2e-29
Identities = 62/111 (55%), Positives = 84/111 (74%), Gaps = 5/111 (4%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ ++++L AF +G V++++VITDRETGRSRGFGFV F+SE+S A+ +
Sbjct: 9 RCFVGGLAWATNNETLEQAFANFGQVIDSKVITDRETGRSRGFGFVTFSSEQSMLDAIEN 68
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGYGGGGGYSGG--GGGYSGGGGG 144
M+G++L+GRNI V+ A R GG GGGYGGGGG GG GGGY GGGGG
Sbjct: 69 MNGKELDGRNITVNQAQSRGGGG--GGGGYGGGGGGYGGREGGGYGGGGGG 117
>GRP_DAUCA (Q03878) Glycine-rich RNA-binding protein
Length = 157
Score = 125 bits (313), Expect = 4e-29
Identities = 64/124 (51%), Positives = 85/124 (67%), Gaps = 14/124 (11%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ +D+SL AF+ +GD+ ++++I DRETGRSRGFGFV F E+S A+
Sbjct: 7 RCFVGGLAWATNDESLEQAFSQFGDITDSKIINDRETGRSRGFGFVTFKDEKSMRDAIEG 66
Query: 96 MDGQDLNGRNIRVSYANDRQAGG-----PRPGGGYGGGGGY-----SGGGGGYSG---GG 142
M+GQ+L+GRNI V+ A R +GG GGGYGGGGGY GGGGGY G GG
Sbjct: 67 MNGQELDGRNITVNEAQSRGSGGGGGRREGGGGGYGGGGGYGGRREGGGGGGYGGRREGG 126
Query: 143 GGGW 146
GGG+
Sbjct: 127 GGGY 130
>GRPA_MAIZE (P10979) Glycine-rich RNA-binding, abscisic
acid-inducible protein
Length = 157
Score = 124 bits (312), Expect = 5e-29
Identities = 66/124 (53%), Positives = 89/124 (71%), Gaps = 15/124 (12%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ ++SL +AF +YG++++++VITDRETGRSRGFGFV F+SE S A+ +
Sbjct: 9 RCFVGGLAWATSNESLENAFASYGEILDSKVITDRETGRSRGFGFVTFSSENSMLDAIEN 68
Query: 96 MDGQDLNGRNIRVSYANDRQA-------GGPRPGGGYGGG---GGYSGGGGGYSG---GG 142
M+G++L+GRNI V+ A R GG R GGGYGGG GGY GGGGGY G GG
Sbjct: 69 MNGKELDGRNITVNQAQSRGGGGGGGGYGGGRGGGGYGGGRRDGGY-GGGGGYGGRREGG 127
Query: 143 GGGW 146
GGG+
Sbjct: 128 GGGY 131
>GRP2_SINAL (P49311) Glycine-rich RNA-binding protein GRP2A
Length = 169
Score = 124 bits (312), Expect = 5e-29
Identities = 67/134 (50%), Positives = 89/134 (66%), Gaps = 24/134 (17%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ D++SL AF+ +G++V++++I DRETGRSRGFGFV F E+S A+
Sbjct: 9 RCFVGGLAWATDERSLETAFSQFGELVDSKIINDRETGRSRGFGFVTFKDEKSMKDAIEG 68
Query: 96 MDGQDLNGRNIRVSYANDRQA----------GGPRPGGGYGG-----------GGGYSGG 134
M+GQDL+GR+I V+ A R + GG R GGGYGG GGGYSGG
Sbjct: 69 MNGQDLDGRSITVNEAQSRGSGAGGGGRGGGGGYRGGGGYGGGGGGYGGGRREGGGYSGG 128
Query: 135 GGGYS--GGGGGGW 146
GGGYS GGGGGG+
Sbjct: 129 GGGYSSRGGGGGGY 142
>RT19_ARATH (P39697) 40S ribosomal protein S19, mitochondrial
precursor
Length = 212
Score = 120 bits (302), Expect = 8e-28
Identities = 66/114 (57%), Positives = 85/114 (73%), Gaps = 6/114 (5%)
Query: 1 MAFCNKIGNLLRQGATQSTQAPVSSMLNYLRHMSSSKLFIGGLSYNVDDQSLRDAFTTYG 60
MAFC K+G +QG PVSSML LR+MS+ KL+IGGLS D+ SL+DAF+++
Sbjct: 1 MAFCTKLGGHWKQGVN----VPVSSMLGSLRYMST-KLYIGGLSPGTDEHSLKDAFSSFN 55
Query: 61 DVVEARVITDRETGRSRGFGFVNFTSEESATSALS-MDGQDLNGRNIRVSYAND 113
V EARV+T++ TGRSRG+GFVNF SE+SA SA+S M+GQ+LNG NI V+ A D
Sbjct: 56 GVTEARVMTNKVTGRSRGYGFVNFISEDSANSAISAMNGQELNGFNISVNVAKD 109
>GRP8_ARATH (Q03251) Glycine-rich RNA-binding protein 8 (CCR1
protein)
Length = 169
Score = 120 bits (301), Expect = 1e-27
Identities = 61/119 (51%), Positives = 86/119 (72%), Gaps = 9/119 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ +D+ L+ F+ +GDV+++++I DRE+GRSRGFGFV F E++ A+
Sbjct: 7 RCFVGGLAWATNDEDLQRTFSQFGDVIDSKIINDRESGRSRGFGFVTFKDEKAMRDAIEE 66
Query: 96 MDGQDLNGRNIRVSYANDRQAGG-----PRPGGGY--GGGGGYS-GGGGGYSGGGGGGW 146
M+G++L+GR I V+ A R +GG GGGY GGGGGYS GGGGGYSGGGGGG+
Sbjct: 67 MNGKELDGRVITVNEAQSRGSGGGGGGRGGSGGGYRSGGGGGYSGGGGGGYSGGGGGGY 125
>GR10_BRANA (Q05966) Glycine-rich RNA-binding protein 10
Length = 169
Score = 115 bits (288), Expect = 3e-26
Identities = 58/114 (50%), Positives = 77/114 (66%), Gaps = 5/114 (4%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
+ F+GGL++ D L F+ +G+V+++++I DRETGRSRGFGFV F E+S A+
Sbjct: 7 RCFVGGLAWATGDAELERTFSQFGEVIDSKIINDRETGRSRGFGFVTFKDEKSMKDAIDE 66
Query: 96 MDGQDLNGRNIRVSYANDR--QAGGPRPGGGYG--GGGGYSGGGGGYSGGGGGG 145
M+G++L+GR I V+ A R GG R GGGYG GGGGY GGGGGY GGG
Sbjct: 67 MNGKELDGRTITVNEAQSRGGGGGGGRGGGGYGGRGGGGYGGGGGGYGDRRGGG 120
>GRP1_SORBI (Q99069) Glycine-rich RNA-binding protein 1 (Fragment)
Length = 142
Score = 113 bits (282), Expect = 2e-25
Identities = 59/102 (57%), Positives = 74/102 (71%), Gaps = 8/102 (7%)
Query: 51 SLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-SMDGQDLNGRNIRVS 109
SL AF+TYG+V+E+++I DRET RSRGFGFV F++EE+ SA+ M+G++L+GRNI V+
Sbjct: 2 SLHSAFSTYGEVLESKIILDRETQRSRGFGFVTFSTEEAMRSAIEGMNGKELDGRNITVN 61
Query: 110 YANDRQAGGPRPGGGY----GGGGGY---SGGGGGYSGGGGG 144
A R G GGGY GGGGGY GGGGGY GGGGG
Sbjct: 62 EAQSRGGRGGGGGGGYGGGRGGGGGYGRRDGGGGGYGGGGGG 103
>RBM3_MOUSE (O89086) Putative RNA-binding protein 3 (RNA binding
motif protein 3)
Length = 153
Score = 103 bits (258), Expect = 1e-22
Identities = 55/109 (50%), Positives = 76/109 (69%), Gaps = 4/109 (3%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
KLF+GGL++N D+Q+L D F+++G + E V+ DRET RSRGFGF+ FT+ E A+ A+ +
Sbjct: 7 KLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHASDAMRA 66
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPGGGYGG-GGGYSGGGGGYSGGGG 143
M+G+ L+GR IRV +A + A G R GG +GG G YS GGG G G
Sbjct: 67 MNGESLDGRQIRVDHAG-KSARGSR-GGAFGGRGRSYSRGGGDQGYGSG 113
>RBM3_HUMAN (P98179) Putative RNA-binding protein 3 (RNA binding
motif protein 3) (RNPL)
Length = 157
Score = 103 bits (256), Expect = 2e-22
Identities = 54/110 (49%), Positives = 75/110 (68%), Gaps = 4/110 (3%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL-S 95
KLF+GGL++N D+Q+L D F+++G + E V+ DRET RSRGFGF+ FT+ E A+ A+ +
Sbjct: 7 KLFVGGLNFNTDEQALEDHFSSFGPISEVVVVKDRETQRSRGFGFITFTNPEHASVAMRA 66
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPG--GGYGGGGGYSGGGGGYSGGGG 143
M+G+ L+GR IRV +A + A G R G G +G G YS GGG G G
Sbjct: 67 MNGESLDGRQIRVDHAG-KSARGTRGGGFGAHGRGRSYSRGGGDQGYGSG 115
>CIRP_HUMAN (Q14011) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 95.9 bits (237), Expect = 3e-20
Identities = 54/114 (47%), Positives = 75/114 (65%), Gaps = 8/114 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA-LS 95
KLF+GGLS++ ++QSL F+ YG + E V+ DRET RSRGFGFV F + + A A ++
Sbjct: 7 KLFVGGLSFDTNEQSLEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 96 MDGQDLNGRNIRVSYA---NDRQAGGPRPGGGYGGGGGYSGG---GGGYSGGGG 143
M+G+ ++GR IRV A +D ++ G R GG GG G + GG G G+S GGG
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGYR-GGSAGGRGFFRGGRGRGRGFSRGGG 119
>CIRP_RAT (P60825) Cold-inducible RNA-binding protein (Glycine-rich
RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 94.7 bits (234), Expect = 6e-20
Identities = 53/114 (46%), Positives = 75/114 (65%), Gaps = 8/114 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA-LS 95
KLF+GGLS++ ++Q+L F+ YG + E V+ DRET RSRGFGFV F + + A A ++
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 96 MDGQDLNGRNIRVSYA---NDRQAGGPRPGGGYGGGGGYSGG---GGGYSGGGG 143
M+G+ ++GR IRV A +D ++ G R GG GG G + GG G G+S GGG
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGYR-GGSAGGRGFFRGGRSRGRGFSRGGG 119
>CIRP_MOUSE (P60824) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 94.7 bits (234), Expect = 6e-20
Identities = 53/114 (46%), Positives = 75/114 (65%), Gaps = 8/114 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA-LS 95
KLF+GGLS++ ++Q+L F+ YG + E V+ DRET RSRGFGFV F + + A A ++
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 96 MDGQDLNGRNIRVSYA---NDRQAGGPRPGGGYGGGGGYSGG---GGGYSGGGG 143
M+G+ ++GR IRV A +D ++ G R GG GG G + GG G G+S GGG
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGYR-GGSAGGRGFFRGGRSRGRGFSRGGG 119
>CIRP_CRIGR (P60826) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (A18 hnRNP)
Length = 172
Score = 94.7 bits (234), Expect = 6e-20
Identities = 53/114 (46%), Positives = 75/114 (65%), Gaps = 8/114 (7%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA-LS 95
KLF+GGLS++ ++Q+L F+ YG + E V+ DRET RSRGFGFV F + + A A ++
Sbjct: 7 KLFVGGLSFDTNEQALEQVFSKYGQISEVVVVKDRETQRSRGFGFVTFENIDDAKDAMMA 66
Query: 96 MDGQDLNGRNIRVSYA---NDRQAGGPRPGGGYGGGGGYSGG---GGGYSGGGG 143
M+G+ ++GR IRV A +D ++ G R GG GG G + GG G G+S GGG
Sbjct: 67 MNGKSVDGRQIRVDQAGKSSDNRSRGYR-GGSAGGRGFFRGGRSRGRGFSRGGG 119
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 93.6 bits (231), Expect = 1e-19
Identities = 47/112 (41%), Positives = 68/112 (59%), Gaps = 2/112 (1%)
Query: 35 SSKLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSAL 94
S +F+G LS+N + L AF GD+ R+ TD ++GR +GFG+V F+ +SA +
Sbjct: 365 SDTVFVGNLSFNATEDDLSTAFGGCGDIQSIRLPTDPQSGRLKGFGYVTFSDIDSAKKCV 424
Query: 95 SMDGQDLNGRNIRVSYANDRQAGGPRPG-GGYGGGGGYSGGGGGYSGGGGGG 145
M+G + GR R+ ++ R GG R G GG+GG GG+ GG GG+ GG G G
Sbjct: 425 EMNGHFIAGRPCRLDFSTPRTGGGSRGGRGGFGGRGGF-GGRGGFGGGRGRG 475
Score = 75.1 bits (183), Expect = 5e-14
Identities = 37/85 (43%), Positives = 61/85 (71%), Gaps = 2/85 (2%)
Query: 38 LFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSALSMD 97
+F+G LS+NVDDQ L F YG +V ARVI D ++GRS+G+G+V+F + E+A +A++ +
Sbjct: 265 VFVGRLSWNVDDQWLGQEFEEYGTIVGARVIMDGQSGRSKGYGYVDFETPEAAKAAVAAN 324
Query: 98 G-QDLNGRNIRVSYANDRQAGGPRP 121
G ++++GR + + +N R A P+P
Sbjct: 325 GTKEIDGRMVNLDLSNPRPA-NPQP 348
>CIRP_XENLA (O93235) Cold-inducible RNA-binding protein
(Glycine-rich RNA-binding protein CIRP) (XCIRP)
Length = 163
Score = 92.8 bits (229), Expect = 2e-19
Identities = 52/112 (46%), Positives = 69/112 (61%), Gaps = 6/112 (5%)
Query: 37 KLFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRSRGFGFVNFTSEESATSA-LS 95
KLFIGGL++ ++ L AFT YG + E V+ DRET RSRGFGFV F + + A A ++
Sbjct: 6 KLFIGGLNFETNEDCLEQAFTKYGRISEVVVVKDRETKRSRGFGFVTFENVDDAKDAMMA 65
Query: 96 MDGQDLNGRNIRVSYANDRQAGGPRPG---GGYGGGGGYSGGGGGYSGGGGG 144
M+G+ ++GR IRV A ++ G R G GG GG G+ GG G GG G
Sbjct: 66 MNGKSVDGRQIRVDQAG--KSSGERRGGYRGGSSGGRGFFRGGRGRGGGDRG 115
>TR2A_MOUSE (Q6PFR5) Transformer-2 protein homolog (TRA-2 alpha)
Length = 281
Score = 92.0 bits (227), Expect = 4e-19
Identities = 56/135 (41%), Positives = 73/135 (53%), Gaps = 13/135 (9%)
Query: 24 SSMLNYLRHMSSSK-------LFIGGLSYNVDDQSLRDAFTTYGDVVEARVITDRETGRS 76
S M N RH S L + GLS ++ LR+ F+ YG + V+ D+ TGRS
Sbjct: 98 SPMSNRRRHTGSRANPDPNTCLGVFGLSLYTTERDLREVFSRYGPLSGVNVVYDQRTGRS 157
Query: 77 RGFGFVNFTSEESATSALSM-DGQDLNGRNIRVSYANDRQAGGPRPGGGYG-----GGGG 130
RGF FV F + + A+ +G +L+GR IRV Y+ ++A P PG G GGGG
Sbjct: 158 RGFAFVYFERIDDSKEAMERANGMELDGRRIRVDYSITKRAHTPTPGIYMGRPTHSGGGG 217
Query: 131 YSGGGGGYSGGGGGG 145
GGGGG GGGGGG
Sbjct: 218 GGGGGGGGGGGGGGG 232
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,668,064
Number of Sequences: 164201
Number of extensions: 957617
Number of successful extensions: 42890
Number of sequences better than 10.0: 1279
Number of HSP's better than 10.0 without gapping: 1029
Number of HSP's successfully gapped in prelim test: 253
Number of HSP's that attempted gapping in prelim test: 6508
Number of HSP's gapped (non-prelim): 13830
length of query: 146
length of database: 59,974,054
effective HSP length: 100
effective length of query: 46
effective length of database: 43,553,954
effective search space: 2003481884
effective search space used: 2003481884
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146705.10


