

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146651.7 - phase: 0 /pseudo
(394 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
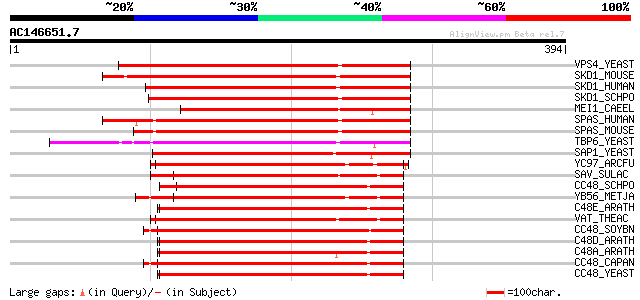
Score E
Sequences producing significant alignments: (bits) Value
VPS4_YEAST (P52917) Vacuolar protein sorting-associated protein ... 221 3e-57
SKD1_MOUSE (P46467) SKD1 protein (Vacuolar sorting protein 4b) 216 1e-55
SKD1_HUMAN (O75351) SKD1 protein (Vacuolar sorting protein 4b) 214 4e-55
SKD1_SCHPO (Q09803) Suppressor protein of bem1/bed5 double mutants 211 3e-54
MEI1_CAEEL (P34808) Meiotic spindle formation protein mei-1 194 4e-49
SPAS_HUMAN (Q9UBP0) Spastin 188 2e-47
SPAS_MOUSE (Q9QYY8) Spastin 187 5e-47
TBP6_YEAST (P40328) Probable 26S protease subunit YTA6 (TAT-bind... 185 2e-46
SAP1_YEAST (P39955) SAP1 protein 175 2e-43
YC97_ARCFU (O28972) Cell division cycle protein 48 homolog AF1297 162 2e-39
SAV_SULAC (Q07590) SAV protein 153 6e-37
CC48_SCHPO (Q9P3A7) Cell division cycle protein 48 homolog 151 3e-36
YB56_METJA (Q58556) Cell division cycle protein 48 homolog MJ1156 149 2e-35
C48E_ARATH (Q9LZF6) Cell division control protein 48 homolog E (... 147 4e-35
VAT_THEAC (O05209) VCP-like ATPase 147 5e-35
CC48_SOYBN (P54774) Cell division cycle protein 48 homolog (Valo... 147 6e-35
C48D_ARATH (Q9SCN8) Putative cell division control protein 48 ho... 146 8e-35
C48A_ARATH (P54609) Cell division control protein 48 homolog A (... 145 1e-34
CC48_CAPAN (Q96372) Cell division cycle protein 48 homolog 145 2e-34
CC48_YEAST (P25694) Cell division control protein 48 141 3e-33
>VPS4_YEAST (P52917) Vacuolar protein sorting-associated protein
VPS4 (END13 protein)
Length = 437
Score = 221 bits (563), Expect = 3e-57
Identities = 111/207 (53%), Positives = 147/207 (70%), Gaps = 2/207 (0%)
Query: 78 GFGPNGVDEKPKKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEA 137
G G NG ++K + + + L +LS I+ PNVKWE + GLE AK LKEA
Sbjct: 89 GSGSNGGNKKISQEEGEDNGGEDNKKLRGALSSAILSEKPNVKWEDVAGLEGAKEALKEA 148
Query: 138 VVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRG 197
V++P+K+P F G P GILL+GPPGTGK+ LAKAVATE N+TFF++S+S +VSKW G
Sbjct: 149 VILPVKFPHLFKGNRKPTSGILLYGPPGTGKSYLAKAVATEANSTFFSVSSSDLVSKWMG 208
Query: 198 DSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLART 257
+SEKLVK LF +AR + P+ IF+DE+DA+ RGEG E EASRR+KTELL+QM+G+
Sbjct: 209 ESEKLVKQLFAMARENKPSIIFIDEVDALTGTRGEG--ESEASRRIKTELLVQMNGVGND 266
Query: 258 DELVFVLAATNLPWELDAAMLRRLEKR 284
+ V VL ATN+PW+LD+A+ RR E+R
Sbjct: 267 SQGVLVLGATNIPWQLDSAIRRRFERR 293
>SKD1_MOUSE (P46467) SKD1 protein (Vacuolar sorting protein 4b)
Length = 444
Score = 216 bits (549), Expect = 1e-55
Identities = 116/220 (52%), Positives = 150/220 (67%), Gaps = 6/220 (2%)
Query: 67 QSQGQNPTHTNGFGPNGVDEKPKKSL-LPPFESAEMRTLAESLSRDIIRGSPNVKWESIK 125
+ + Q P GP VDEK S + E + L L I+ PNVKW +
Sbjct: 80 EKKPQKPVKEEQSGP--VDEKGNDSDGEAESDDPEKKKLQNQLQGAIVIERPNVKWSDVA 137
Query: 126 GLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATEC-NTTFF 184
GLE AK LKEAV++PIK+P FTG +PW+GILLFGPPGTGK+ LAKAVATE N+TFF
Sbjct: 138 GLEGAKEALKEAVILPIKFPHLFTGKRTPWRGILLFGPPGTGKSYLAKAVATEANNSTFF 197
Query: 185 NISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLK 244
+IS+S +VSKW G+SEKLVK LF+LAR + P+ IF+DEID++ R E +E EA+RR+K
Sbjct: 198 SISSSDLVSKWLGESEKLVKNLFQLARENKPSIIFIDEIDSLCGSRSE--NESEAARRIK 255
Query: 245 TELLIQMDGLARTDELVFVLAATNLPWELDAAMLRRLEKR 284
TE L+QM G+ ++ + VL ATN+PW LD+A+ RR EKR
Sbjct: 256 TEFLVQMQGVGVDNDGILVLGATNIPWVLDSAIRRRFEKR 295
>SKD1_HUMAN (O75351) SKD1 protein (Vacuolar sorting protein 4b)
Length = 444
Score = 214 bits (544), Expect = 4e-55
Identities = 107/189 (56%), Positives = 139/189 (72%), Gaps = 3/189 (1%)
Query: 97 ESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 156
+ E + L L I+ PNVKW + GLE AK LKEAV++PIK+P FTG +PW+
Sbjct: 109 DDPEKKKLQNQLQGAIVIERPNVKWSDVAGLEGAKEALKEAVILPIKFPHLFTGKRTPWR 168
Query: 157 GILLFGPPGTGKTMLAKAVATEC-NTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAP 215
GILLFGPPGTGK+ LAKAVATE N+TFF+IS+S +VSKW G+SEKLVK LF+LAR + P
Sbjct: 169 GILLFGPPGTGKSYLAKAVATEANNSTFFSISSSDLVSKWLGESEKLVKNLFQLARENKP 228
Query: 216 ATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDA 275
+ IF+DEID++ R E +E EA+RR+KTE L+QM G+ ++ + VL ATN+PW LD+
Sbjct: 229 SIIFIDEIDSLCGSRSE--NESEAARRIKTEFLVQMQGVGVDNDGILVLGATNIPWVLDS 286
Query: 276 AMLRRLEKR 284
A+ RR EKR
Sbjct: 287 AIRRRFEKR 295
>SKD1_SCHPO (Q09803) Suppressor protein of bem1/bed5 double mutants
Length = 432
Score = 211 bits (536), Expect = 3e-54
Identities = 102/186 (54%), Positives = 138/186 (73%), Gaps = 2/186 (1%)
Query: 99 AEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGI 158
++ + L +L+ I+ PNV+W+ I GLENAK LKE V++PIK P+ F+ PW GI
Sbjct: 106 SDAKKLRSALTSAILVEKPNVRWDDIAGLENAKEALKETVLLPIKLPQLFSHGRKPWSGI 165
Query: 159 LLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATI 218
LL+GPPGTGK+ LAKAVATE +TFF+IS+S +VSKW G+SE+LV+ LFE+AR P+ I
Sbjct: 166 LLYGPPGTGKSYLAKAVATEAGSTFFSISSSDLVSKWMGESERLVRQLFEMAREQKPSII 225
Query: 219 FLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAML 278
F+DEID++ R EG E E+SRR+KTE L+QM+G+ + + V VL ATN+PW LD+A+
Sbjct: 226 FIDEIDSLCGSRSEG--ESESSRRIKTEFLVQMNGVGKDESGVLVLGATNIPWTLDSAIR 283
Query: 279 RRLEKR 284
RR EKR
Sbjct: 284 RRFEKR 289
>MEI1_CAEEL (P34808) Meiotic spindle formation protein mei-1
Length = 472
Score = 194 bits (492), Expect = 4e-49
Identities = 93/165 (56%), Positives = 127/165 (76%), Gaps = 3/165 (1%)
Query: 122 ESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECNT 181
+ I G+ + K++L EAV +P+ P++F GL SPWK ++L GPPGTGKT++A+A+A+E ++
Sbjct: 193 DDIIGMHDVKQVLHEAVTLPLLVPEFFQGLRSPWKAMVLAGPPGTGKTLIARAIASESSS 252
Query: 182 TFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHEASR 241
TFF +S++ + SKWRGDSEK+V++LFELAR +AP+ IF+DEID + QRG EHEASR
Sbjct: 253 TFFTVSSTDLSSKWRGDSEKIVRLLFELARFYAPSIIFIDEIDTLGGQRGNS-GEHEASR 311
Query: 242 RLKTELLIQMDGLAR--TDELVFVLAATNLPWELDAAMLRRLEKR 284
R+K+E L+QMDG VFVLAATN+PWELD A+ RR EKR
Sbjct: 312 RVKSEFLVQMDGSQNKFDSRRVFVLAATNIPWELDEALRRRFEKR 356
>SPAS_HUMAN (Q9UBP0) Spastin
Length = 616
Score = 188 bits (477), Expect = 2e-47
Identities = 108/227 (47%), Positives = 144/227 (62%), Gaps = 12/227 (5%)
Query: 67 QSQGQNPTHTNGFGPNGVDEKP--------KKSLLPPFESAEMRTLAESLSRDIIRGSPN 118
Q G PT G KP KK L F + + LA + +I+
Sbjct: 280 QGSGPAPTTHKGTPKTNRTNKPSTPTTATRKKKDLKNFRNVDSN-LANLIMNEIVDNGTA 338
Query: 119 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATE 178
VK++ I G + AK+ L+E V++P P+ FTGL +P +G+LLFGPPG GKTMLAKAVA E
Sbjct: 339 VKFDDIAGQDLAKQALQEIVILPSLRPELFTGLRAPARGLLLFGPPGNGKTMLAKAVAAE 398
Query: 179 CNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEHE 238
N TFFNISA+S+ SK+ G+ EKLV+ LF +AR P+ IF+DE+D+++ +R EG EH+
Sbjct: 399 SNATFFNISAASLTSKYVGEGEKLVRALFAVARELQPSIIFIDEVDSLLCERREG--EHD 456
Query: 239 ASRRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLRRLEKR 284
ASRRLKTE LI+ DG+ + D+ V V+ ATN P ELD A+LRR KR
Sbjct: 457 ASRRLKTEFLIEFDGVQSAGDDRVLVMGATNRPQELDEAVLRRFIKR 503
>SPAS_MOUSE (Q9QYY8) Spastin
Length = 614
Score = 187 bits (474), Expect = 5e-47
Identities = 102/197 (51%), Positives = 137/197 (68%), Gaps = 4/197 (2%)
Query: 89 KKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYF 148
KK L F + + LA + +I+ VK++ I G E AK+ L+E V++P P+ F
Sbjct: 308 KKKDLKNFRNVDSN-LANLIMNEIVDNGTAVKFDDIAGQELAKQALQEIVILPSLRPELF 366
Query: 149 TGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFE 208
TGL +P +G+LLFGPPG GKTMLAKAVA E N TFFNISA+S+ SK+ G+ EKLV+ LF
Sbjct: 367 TGLRAPARGLLLFGPPGNGKTMLAKAVAAESNATFFNISAASLTSKYVGEGEKLVRALFA 426
Query: 209 LARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLAAT 267
+AR P+ IF+DE+D+++ +R EG EH+ASRRLKTE LI+ DG+ + D+ V V+ AT
Sbjct: 427 VARELQPSIIFIDEVDSLLCERREG--EHDASRRLKTEFLIEFDGVQSAGDDRVLVMGAT 484
Query: 268 NLPWELDAAMLRRLEKR 284
N P ELD A+LRR KR
Sbjct: 485 NRPQELDEAVLRRFIKR 501
>TBP6_YEAST (P40328) Probable 26S protease subunit YTA6 (TAT-binding
homolog 6)
Length = 754
Score = 185 bits (470), Expect = 2e-46
Identities = 110/267 (41%), Positives = 157/267 (58%), Gaps = 16/267 (5%)
Query: 29 KLVENGENAVSNGNSSGIASNGNSHGKVTSDRAIYDQFQSQGQNPTHTNGFGPNGVDEKP 88
K ++ N VS N + + T+ A +S+ P + + +D +
Sbjct: 381 KASKSNTNKVSRRNEQNLEPSSPVLVSATAVPAESKPMRSKSGTPDKESS-ASSSLDSR- 438
Query: 89 KKSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYF 148
K+ +L + + R E + +I+ V WE I GL NAK LKEAVV P P F
Sbjct: 439 KEDILKSVQGVD-RNACEQILNEILVTDEKVYWEDIAGLRNAKNSLKEAVVYPFLRPDLF 497
Query: 149 TGLLSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFE 208
GL P +G+LLFGPPGTGKTM+AKAVATE N+TFF++SASS++SK+ G+SEKLV+ LF
Sbjct: 498 KGLREPVRGMLLFGPPGTGKTMIAKAVATESNSTFFSVSASSLLSKYLGESEKLVRALFY 557
Query: 209 LARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLART----------- 257
+A+ +P+ IF+DEID++++ R + +E+E+SRR+KTELLIQ L+
Sbjct: 558 MAKKLSPSIIFIDEIDSMLTARSD--NENESSRRIKTELLIQWSSLSSATAQSEDRNNTL 615
Query: 258 DELVFVLAATNLPWELDAAMLRRLEKR 284
D V VL ATNLPW +D A RR ++
Sbjct: 616 DSRVLVLGATNLPWAIDDAARRRFSRK 642
>SAP1_YEAST (P39955) SAP1 protein
Length = 897
Score = 175 bits (444), Expect = 2e-43
Identities = 97/202 (48%), Positives = 128/202 (63%), Gaps = 20/202 (9%)
Query: 102 RTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLF 161
R A+ + +I+ V W+ I GLE+AK LKEAVV P P F GL P +G+LLF
Sbjct: 585 RQAAKQIFAEIVVHGDEVHWDDIAGLESAKYSLKEAVVYPFLRPDLFRGLREPVRGMLLF 644
Query: 162 GPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLD 221
GPPGTGKTMLA+AVATE ++TFF+ISASS+ SK+ G+SEKLV+ LF +A+ +P+ IF+D
Sbjct: 645 GPPGTGKTMLARAVATESHSTFFSISASSLTSKYLGESEKLVRALFAIAKKLSPSIIFVD 704
Query: 222 EIDAIISQRGEGRSEHEASRRLKTELLIQMDGLA-------------------RTDELVF 262
EID+I+ R +E+E+SRR+K E L+Q L+ D V
Sbjct: 705 EIDSIMGSR-NNENENESSRRIKNEFLVQWSSLSSAAAGSNKSNTNNSDTNGDEDDTRVL 763
Query: 263 VLAATNLPWELDAAMLRRLEKR 284
VLAATNLPW +D A RR +R
Sbjct: 764 VLAATNLPWSIDEAARRRFVRR 785
>YC97_ARCFU (O28972) Cell division cycle protein 48 homolog AF1297
Length = 733
Score = 162 bits (409), Expect = 2e-39
Identities = 88/186 (47%), Positives = 124/186 (66%), Gaps = 5/186 (2%)
Query: 101 MRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGIL 159
++ + S R+++ PNVKWE I GLE+AK+ L EAV P+KYP+ F + P +GIL
Sbjct: 434 LKNIEPSAMREVLVEVPNVKWEDIGGLEHAKQELMEAVEWPLKYPEVFRAANIKPPRGIL 493
Query: 160 LFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIF 219
LFGPPGTGKT+LAKAVA E N F ++ ++SKW G+SEK V+ +F AR AP IF
Sbjct: 494 LFGPPGTGKTLLAKAVANESNANFISVKGPELLSKWVGESEKHVREMFRKARQVAPCVIF 553
Query: 220 LDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
DEID++ +RG G + + R+ ++LL ++DGL ++V V+AATN P +D A+LR
Sbjct: 554 FDEIDSLAPRRG-GIGDSHVTERVVSQLLTELDGLEELKDVV-VIAATNRPDMIDPALLR 611
Query: 280 --RLEK 283
RLE+
Sbjct: 612 PGRLER 617
Score = 141 bits (356), Expect = 3e-33
Identities = 82/178 (46%), Positives = 119/178 (66%), Gaps = 6/178 (3%)
Query: 104 LAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFG 162
L E + ++ R P+V +E I GL+ RL++E + +P+K+P+ F L + P KG+LL+G
Sbjct: 164 LKEKPAEEVKRAVPDVTYEDIGGLKRELRLVREMIELPLKHPELFQRLGIEPPKGVLLYG 223
Query: 163 PPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDE 222
PPGTGKT++AKAVA E + F IS I+SK+ G+SE+ ++ +FE A+ +AP+ IF+DE
Sbjct: 224 PPGTGKTLIAKAVANEVDAHFIPISGPEIMSKYYGESEQRLREIFEEAKENAPSIIFIDE 283
Query: 223 IDAIISQRGEGRSEHEASRRLKTELLIQMDGL-ARTDELVFVLAATNLPWELDAAMLR 279
ID+I +R E E E RR+ +LL MDGL AR D V V+AATN P +D A+ R
Sbjct: 284 IDSIAPKREEVTGEVE--RRVVAQLLALMDGLEARGD--VIVIAATNRPDAIDPALRR 337
>SAV_SULAC (Q07590) SAV protein
Length = 780
Score = 153 bits (387), Expect = 6e-37
Identities = 83/180 (46%), Positives = 118/180 (65%), Gaps = 3/180 (1%)
Query: 101 MRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGIL 159
++++ SL R++ P V W I GL+N K+ L+EAV P+++P+ FT ++P KGIL
Sbjct: 466 LKSIQPSLLREVYVEVPKVNWNDIGGLDNVKQQLREAVEWPLRFPELFTKSGVTPPKGIL 525
Query: 160 LFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIF 219
LFGPPGTGKTMLAKAVATE F + I+SKW G+SEK ++ +F AR AP IF
Sbjct: 526 LFGPPGTGKTMLAKAVATESGANFIAVRGPEILSKWVGESEKAIREIFRKARQAAPTVIF 585
Query: 220 LDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
DEID+I RG ++ + R+ +LL +MDG+ +++V ++AATN P LD A+LR
Sbjct: 586 FDEIDSIAPIRGLS-TDSGVTERIVNQLLAEMDGIVPLNKVV-IIAATNRPDILDPALLR 643
Score = 127 bits (318), Expect = 6e-29
Identities = 70/164 (42%), Positives = 104/164 (62%), Gaps = 4/164 (2%)
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 175
P V WE I LE AK+ ++E V P+++P+ F L + P KGILL+GPPGTGKT+LA+A+
Sbjct: 207 PKVSWEDIGDLEEAKQKIREIVEWPMRHPELFQRLGIDPPKGILLYGPPGTGKTLLARAL 266
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRS 235
E F ++ I+SK+ G+SE+ ++ +F+ A +AP+ IF+DEIDAI +R +
Sbjct: 267 RNEIGAYFITVNGPEIMSKFYGESEQRIREIFKEAEENAPSIIFIDEIDAIAPKREDVTG 326
Query: 236 EHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
E E +R+ +LL MDG+ V V+ ATN P +D A+ R
Sbjct: 327 EVE--KRVVAQLLTLMDGIKGRGR-VIVIGATNRPDAIDPALRR 367
>CC48_SCHPO (Q9P3A7) Cell division cycle protein 48 homolog
Length = 815
Score = 151 bits (381), Expect = 3e-36
Identities = 76/174 (43%), Positives = 111/174 (63%), Gaps = 2/174 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV+WE I GLE KR L+E V MP+ Y + F ++P KG+L FGPPG
Sbjct: 482 SALRETVVEVPNVRWEDIGGLEEVKRELRETVQMPVMYAEKFLRFGVTPSKGVLFFGPPG 541
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKA+A EC+ F ++ ++S W G+SE V+ +F+ AR AP +FLDE+D+
Sbjct: 542 TGKTLLAKAIANECSANFISVKGPELLSMWFGESESNVRDIFDKARAAAPCVVFLDELDS 601
Query: 226 IISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I RG + R+ +LL +MDG+ + + VFV+ ATN P ++D A++R
Sbjct: 602 IAKARGASAGDSGGGDRVVNQLLTEMDGV-NSKKNVFVIGATNRPDQIDPALMR 654
Score = 118 bits (295), Expect = 3e-26
Identities = 65/162 (40%), Positives = 102/162 (62%), Gaps = 4/162 (2%)
Query: 119 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVAT 177
V ++ I G ++E V +P+++P+ F + + P +GIL++GPPGTGKT++A+AVA
Sbjct: 221 VGYDDIGGCRRQMAQIRELVELPLRHPQLFKSIGIKPPRGILMYGPPGTGKTLMARAVAN 280
Query: 178 ECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRSEH 237
E FF I+ I+SK G+SE ++ FE A ++PA IF+DEID+I +R ++
Sbjct: 281 ETGAFFFLINGPEIMSKMAGESESNLRKAFEEAEKNSPAIIFIDEIDSIAPKR--EKTNG 338
Query: 238 EASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
E RR+ ++LL MDG+ +V V+AATN P +D A+ R
Sbjct: 339 EVERRVVSQLLTLMDGMKARSNVV-VMAATNRPNSIDPALRR 379
>YB56_METJA (Q58556) Cell division cycle protein 48 homolog MJ1156
Length = 903
Score = 149 bits (375), Expect = 2e-35
Identities = 84/191 (43%), Positives = 119/191 (61%), Gaps = 4/191 (2%)
Query: 90 KSLLPPFESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFT 149
K + F+ A ++ + S R+++ PNVKWE I GLE K+ L+EAV P+K + F
Sbjct: 421 KVTMDDFKEA-LKDVEPSAMREVLVEVPNVKWEDIGGLEEVKQELREAVEWPLKAKEVFE 479
Query: 150 GL-LSPWKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFE 208
+ + P KG+LLFGPPGTGKT+LAKAVA E F ++ I SKW G+SEK ++ +F
Sbjct: 480 KIGVRPPKGVLLFGPPGTGKTLLAKAVANESGANFISVKGPEIFSKWVGESEKAIREIFR 539
Query: 209 LARHHAPATIFLDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATN 268
AR AP IF DEIDAI +RG S + ++ +LL ++DG+ ++V V+AATN
Sbjct: 540 KARQSAPCIIFFDEIDAIAPKRGRDLSS-AVTDKVVNQLLTELDGMEEPKDVV-VIAATN 597
Query: 269 LPWELDAAMLR 279
P +D A+LR
Sbjct: 598 RPDIIDPALLR 608
Score = 131 bits (330), Expect = 3e-30
Identities = 74/164 (45%), Positives = 108/164 (65%), Gaps = 4/164 (2%)
Query: 117 PNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 175
P+V +E I GL+ + ++E + +P+++P+ F L + P KG+LL GPPGTGKT+LAKAV
Sbjct: 174 PDVTYEDIGGLKEEVKKVREMIELPMRHPELFEKLGIEPPKGVLLVGPPGTGKTLLAKAV 233
Query: 176 ATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDAIISQRGEGRS 235
A E F+ I+ I+SK+ G++E+ ++ +FE A +AP+ IF+DEIDAI +R E
Sbjct: 234 ANEAGANFYVINGPEIMSKYVGETEENLRKIFEEAEENAPSIIFIDEIDAIAPKRDEATG 293
Query: 236 EHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
E E RRL +LL MDGL ++V V+ ATN P LD A+ R
Sbjct: 294 EVE--RRLVAQLLTLMDGLKGRGQVV-VIGATNRPNALDPALRR 334
>C48E_ARATH (Q9LZF6) Cell division control protein 48 homolog E
(AtCDC48e) (Transitional endoplasmic reticulum ATPase E)
Length = 810
Score = 147 bits (372), Expect = 4e-35
Identities = 75/175 (42%), Positives = 112/175 (63%), Gaps = 3/175 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QRG + A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR
Sbjct: 585 IATQRGNSAGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLR 638
Score = 117 bits (294), Expect = 4e-26
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 311 SIAPKR--EKTNGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 362
>VAT_THEAC (O05209) VCP-like ATPase
Length = 745
Score = 147 bits (371), Expect = 5e-35
Identities = 79/180 (43%), Positives = 117/180 (64%), Gaps = 3/180 (1%)
Query: 101 MRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGIL 159
++++ S R+++ PNV W+ I GLE+ KR +KE V +P+ P F L + P KG L
Sbjct: 446 LKSIEPSSLREVMVEVPNVHWDDIGGLEDVKREIKETVELPLLKPDVFKRLGIRPSKGFL 505
Query: 160 LFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIF 219
L+GPPG GKT+LAKAVATE N F +I ++SKW G+SEK ++ +F+ A+ APA +F
Sbjct: 506 LYGPPGVGKTLLAKAVATESNANFISIKGPEVLSKWVGESEKAIREIFKKAKQVAPAIVF 565
Query: 220 LDEIDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
LDEID+I +RG S+ + R+ +LL +DG+ + +V V+ ATN P +D A+LR
Sbjct: 566 LDEIDSIAPRRGT-TSDSGVTERIVNQLLTSLDGIEVMNGVV-VIGATNRPDIMDPALLR 623
Score = 119 bits (298), Expect = 1e-26
Identities = 66/177 (37%), Positives = 107/177 (60%), Gaps = 4/177 (2%)
Query: 104 LAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFG 162
+ E + +++ + +E I GL ++E + +P+K+P+ F L ++P KG++L+G
Sbjct: 172 IREEPASEVLEEVSRISYEDIGGLSEQLGKIREMIELPLKHPELFERLGITPPKGVILYG 231
Query: 163 PPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDE 222
PPGTGKT++A+AVA E F +I+ I+SK+ G SE+ ++ +F A AP+ IF+DE
Sbjct: 232 PPGTGKTLIARAVANESGANFLSINGPEIMSKYYGQSEQKLREIFSKAEETAPSIIFIDE 291
Query: 223 IDAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
ID+I +R E + E E RR+ +LL MDG+ V V+ ATN +D A+ R
Sbjct: 292 IDSIAPKREEVQGEVE--RRVVAQLLTLMDGMKERGH-VIVIGATNRIDAIDPALRR 345
>CC48_SOYBN (P54774) Cell division cycle protein 48 homolog (Valosin
containing protein homolog) (VCP)
Length = 807
Score = 147 bits (370), Expect = 6e-35
Identities = 78/186 (41%), Positives = 118/186 (62%), Gaps = 4/186 (2%)
Query: 96 FESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSP 154
F++A + T S R+ + PNV WE I GLEN KR L+E V P+++P+ F +SP
Sbjct: 456 FQTA-LGTSNPSALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSP 514
Query: 155 WKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHA 214
KG+L +GPPG GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR A
Sbjct: 515 SKGVLFYGPPGCGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSA 574
Query: 215 PATIFLDEIDAIISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWEL 273
P +F DE+D+I +QRG + A+ R+ +LL +MDG++ + VF++ ATN P +
Sbjct: 575 PCVLFFDELDSIATQRGSSVGDAGGAADRVLNQLLTEMDGMS-AKKTVFIIGATNRPDII 633
Query: 274 DAAMLR 279
D A+LR
Sbjct: 634 DPALLR 639
Score = 119 bits (297), Expect = 2e-26
Identities = 68/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E L R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 192 EPLKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 251
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 252 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 311
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 312 SIAPKR--EKTHGEVERRIVSQLLTLMDGL-KSRAHVIVIGATNRPNSIDPALRR 363
>C48D_ARATH (Q9SCN8) Putative cell division control protein 48
homolog D (AtCDC48d) (Transitional endoplasmic reticulum
ATPase D)
Length = 815
Score = 146 bits (369), Expect = 8e-35
Identities = 76/175 (43%), Positives = 111/175 (63%), Gaps = 3/175 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV WE I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 466 SALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 525
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F +I +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 526 CGKTLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 585
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QRG + A+ R+ +LL +MDG+ + VF++ ATN P +D A+LR
Sbjct: 586 IATQRGNSVGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDPALLR 639
Score = 118 bits (295), Expect = 3e-26
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 192 EPIKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 251
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 252 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 311
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 312 SIAPKR--EKTHGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 363
>C48A_ARATH (P54609) Cell division control protein 48 homolog A
(AtCDC48a)
Length = 809
Score = 145 bits (367), Expect = 1e-34
Identities = 75/176 (42%), Positives = 111/176 (62%), Gaps = 4/176 (2%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + PNV W I GLEN KR L+E V P+++P+ F +SP KG+L +GPPG
Sbjct: 465 SALRETVVEVPNVSWNDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPG 524
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR AP +F DE+D+
Sbjct: 525 CGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDS 584
Query: 226 IISQR--GEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I +QR G G A+ R+ +LL +MDG+ + VF++ ATN P +D+A+LR
Sbjct: 585 IATQRGGGSGGDGGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDIIDSALLR 639
Score = 118 bits (295), Expect = 3e-26
Identities = 67/175 (38%), Positives = 108/175 (61%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ +V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 191 EPVKREDEERLDDVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 250
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 251 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 310
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 311 SIAPKR--EKTNGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 362
>CC48_CAPAN (Q96372) Cell division cycle protein 48 homolog
Length = 805
Score = 145 bits (366), Expect = 2e-34
Identities = 78/186 (41%), Positives = 116/186 (61%), Gaps = 4/186 (2%)
Query: 96 FESAEMRTLAESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSP 154
F++A + T S R+ + PNV WE I GLEN KR L+E V P++ P+ F +SP
Sbjct: 456 FQTA-LGTSNPSALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEPPEKFEKFGMSP 514
Query: 155 WKGILLFGPPGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHA 214
KG+L +GPPG GKT+LAKA+A EC F ++ +++ W G+SE V+ +F+ AR A
Sbjct: 515 SKGVLFYGPPGCGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSA 574
Query: 215 PATIFLDEIDAIISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWEL 273
P +F DE+D+I +QRG + A+ R+ +LL +MDG+ + VF++ ATN P +
Sbjct: 575 PCVLFFDELDSIATQRGSSSGDAGGAADRVLNQLLTEMDGM-NAKKTVFIIGATNRPDII 633
Query: 274 DAAMLR 279
D A+LR
Sbjct: 634 DPALLR 639
Score = 117 bits (294), Expect = 4e-26
Identities = 67/175 (38%), Positives = 107/175 (60%), Gaps = 4/175 (2%)
Query: 106 ESLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 164
E + R+ V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 192 EPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 251
Query: 165 GTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEID 224
G+GKT++A+AVA E FF I+ I+SK G+SE ++ FE A +AP+ IF+DEID
Sbjct: 252 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 311
Query: 225 AIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
+I +R ++ E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R
Sbjct: 312 SIAPKR--EKTHGEVERRIVSQLLTLMDGL-KSRAHVIVMGATNRPNSIDPALRR 363
>CC48_YEAST (P25694) Cell division control protein 48
Length = 835
Score = 141 bits (356), Expect = 3e-33
Identities = 76/175 (43%), Positives = 110/175 (62%), Gaps = 3/175 (1%)
Query: 107 SLSRDIIRGSPNVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 165
S R+ + S NV W+ + GL+ K LKE V P+ +P +T LSP KG+L +GPPG
Sbjct: 472 SALRETVVESVNVTWDDVGGLDEIKEELKETVEYPVLHPDQYTKFGLSPSKGVLFYGPPG 531
Query: 166 TGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEIDA 225
TGKT+LAKAVATE + F ++ ++S W G+SE ++ +F+ AR AP +FLDE+D+
Sbjct: 532 TGKTLLAKAVATEVSANFISVKGPELLSMWYGESESNIRDIFDKARAAAPTVVFLDELDS 591
Query: 226 IISQRGEGRSE-HEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
I RG + AS R+ +LL +MDG+ + VFV+ ATN P ++D A+LR
Sbjct: 592 IAKARGGSLGDAGGASDRVVNQLLTEMDGM-NAKKNVFVIGATNRPDQIDPAILR 645
Score = 119 bits (298), Expect = 1e-26
Identities = 67/176 (38%), Positives = 109/176 (61%), Gaps = 5/176 (2%)
Query: 106 ESLSRDIIRGSPN-VKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGP 163
E ++R+ + N V ++ I G ++E V +P+++P+ F + + P +G+L++GP
Sbjct: 197 EPINREDEENNMNEVGYDDIGGCRKQMAQIREMVELPLRHPQLFKAIGIKPPRGVLMYGP 256
Query: 164 PGTGKTMLAKAVATECNTTFFNISASSIVSKWRGDSEKLVKVLFELARHHAPATIFLDEI 223
PGTGKT++A+AVA E FF I+ ++SK G+SE ++ FE A +APA IF+DEI
Sbjct: 257 PGTGKTLMARAVANETGAFFFLINGPEVMSKMAGESESNLRKAFEEAEKNAPAIIFIDEI 316
Query: 224 DAIISQRGEGRSEHEASRRLKTELLIQMDGLARTDELVFVLAATNLPWELDAAMLR 279
D+I +R ++ E RR+ ++LL MDG+ +V V+AATN P +D A+ R
Sbjct: 317 DSIAPKR--DKTNGEVERRVVSQLLTLMDGMKARSNVV-VIAATNRPNSIDPALRR 369
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,856,211
Number of Sequences: 164201
Number of extensions: 1939829
Number of successful extensions: 7966
Number of sequences better than 10.0: 779
Number of HSP's better than 10.0 without gapping: 519
Number of HSP's successfully gapped in prelim test: 260
Number of HSP's that attempted gapping in prelim test: 6864
Number of HSP's gapped (non-prelim): 1017
length of query: 394
length of database: 59,974,054
effective HSP length: 112
effective length of query: 282
effective length of database: 41,583,542
effective search space: 11726558844
effective search space used: 11726558844
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146651.7


