

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
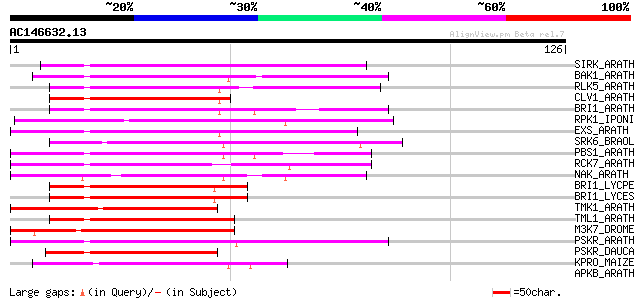
Score E
Sequences producing significant alignments: (bits) Value
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 53 1e-07
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 52 2e-07
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 52 3e-07
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 52 3e-07
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 50 9e-07
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 50 1e-06
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 47 7e-06
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 46 1e-05
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 45 4e-05
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 44 6e-05
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 44 6e-05
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 43 1e-04
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 43 1e-04
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 42 2e-04
TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor 42 3e-04
M3K7_DROME (P83104) Putative mitogen-activated protein kinase ki... 42 3e-04
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 41 5e-04
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 40 7e-04
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 40 7e-04
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 40 0.001
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 52.8 bits (125), Expect = 1e-07
Identities = 26/74 (35%), Positives = 44/74 (59%), Gaps = 1/74 (1%)
Query: 8 ALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHG 67
++GY+ PE ++NEK DVY GV++LE++TG+ + + + ++DHVR +L +G
Sbjct: 737 SIGYLDPEYY-STRQMNEKSDVYSLGVVLLEVITGQPAIASSKTEKVHISDHVRSILANG 795
Query: 68 NALECVDPSLMSEY 81
+ VD L Y
Sbjct: 796 DIRGIVDQRLRERY 809
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 52.0 bits (123), Expect = 2e-07
Identities = 30/85 (35%), Positives = 51/85 (59%), Gaps = 6/85 (7%)
Query: 6 QSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLILNDHVR 61
+ +G++APE + +EK DV+G+GVM+LE++TG+R + +D+V++L D V+
Sbjct: 453 RGTIGHIAPEYL-STGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLL-DWVK 510
Query: 62 VLLEHGNALECVDPSLMSEYPEDEV 86
LL+ VD L Y ++EV
Sbjct: 511 GLLKEKKLEALVDVDLQGNYKDEEV 535
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 51.6 bits (122), Expect = 3e-07
Identities = 31/77 (40%), Positives = 45/77 (58%), Gaps = 6/77 (7%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV--EYGEDNVLILNDHVRVLLEHG 67
GY+APE LRVNEK D+Y FGV++LE+VTGK+P E G+ + + V L+
Sbjct: 863 GYIAPEYV-YTLRVNEKSDIYSFGVVLLELVTGKQPTDSELGDKD---MAKWVCTALDKC 918
Query: 68 NALECVDPSLMSEYPED 84
+DP L ++ E+
Sbjct: 919 GLEPVIDPKLDLKFKEE 935
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 51.6 bits (122), Expect = 3e-07
Identities = 25/42 (59%), Positives = 34/42 (80%), Gaps = 2/42 (4%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV-EYGE 50
GY+APE A L+V+EK DVY FGV++LE++ GK+PV E+GE
Sbjct: 859 GYIAPEYA-YTLKVDEKSDVYSFGVVLLELIAGKKPVGEFGE 899
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 50.1 bits (118), Expect = 9e-07
Identities = 30/81 (37%), Positives = 48/81 (59%), Gaps = 10/81 (12%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV---EYGEDNVL-ILNDHVRVLLE 65
GYV PE Q R + K DVY +GV++LE++TGKRP ++G++N++ + H ++ +
Sbjct: 1051 GYVPPEYY-QSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNLVGWVKQHAKLRIS 1109
Query: 66 HGNALECVDPSLMSEYPEDEV 86
+ DP LM E P E+
Sbjct: 1110 -----DVFDPELMKEDPALEI 1125
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 49.7 bits (117), Expect = 1e-06
Identities = 30/87 (34%), Positives = 49/87 (55%), Gaps = 2/87 (2%)
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVR 61
SN Q +GY+APE A ++ E DVY +GV++LE++T K+ ++ + + VR
Sbjct: 976 SNTVQGTIGYMAPENAFTTVKSRES-DVYSYGVVLLELITRKKALDPSFNGETDIVGWVR 1034
Query: 62 -VLLEHGNALECVDPSLMSEYPEDEVL 87
V + G + VDPSL+ E + V+
Sbjct: 1035 SVWTQTGEIQKIVDPSLLDELIDSSVM 1061
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 47.0 bits (110), Expect = 7e-06
Identities = 29/81 (35%), Positives = 41/81 (49%), Gaps = 3/81 (3%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPV--EYGEDNVLILND 58
+S GY+ PE Q R K DVY FGV++LE+VTGK P ++ E L
Sbjct: 1075 VSTVIAGTFGYIPPEYG-QSARATTKGDVYSFGVILLELVTGKEPTGPDFKESEGGNLVG 1133
Query: 59 HVRVLLEHGNALECVDPSLMS 79
+ G A++ +DP L+S
Sbjct: 1134 WAIQKINQGKAVDVIDPLLVS 1154
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 46.2 bits (108), Expect = 1e-05
Identities = 34/88 (38%), Positives = 48/88 (53%), Gaps = 9/88 (10%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE-YGEDNVLILNDHVRVLLEHGN 68
GY++PE A + +EK DV+ FGV++LEIV+GK+ Y D L +V + G
Sbjct: 695 GYMSPEYAMYGI-FSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYENDLLSYVWSRWKEGR 753
Query: 69 ALECVDPSLM-------SEYPEDEVLPC 89
ALE VDP ++ S + EVL C
Sbjct: 754 ALEIVDPVIVDSLSSQPSIFQPQEVLKC 781
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 44.7 bits (104), Expect = 4e-05
Identities = 28/91 (30%), Positives = 46/91 (49%), Gaps = 17/91 (18%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE----YGEDNVL-- 54
+S R GY APE A ++ K DVY FGV+ LE++TG++ ++ +GE N++
Sbjct: 246 VSTRVMGTYGYCAPEYA-MTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAW 304
Query: 55 ---ILNDHVRVLLEHGNALECVDPSLMSEYP 82
+ ND + ++ DP L +P
Sbjct: 305 ARPLFNDRRKF-------IKLADPRLKGRFP 328
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 43.9 bits (102), Expect = 6e-05
Identities = 28/88 (31%), Positives = 43/88 (48%), Gaps = 11/88 (12%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S R GY AP+ A ++ K D+Y FGV++LE++TG++ + DN D
Sbjct: 263 VSTRVMGTYGYCAPDYA-MTGQLTFKSDIYSFGVVLLELITGRKAI----DNTKTRKDQN 317
Query: 61 RV------LLEHGNALECVDPSLMSEYP 82
V + N + VDP L +YP
Sbjct: 318 LVGWARPLFKDRRNFPKMVDPLLQGQYP 345
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 43.9 bits (102), Expect = 6e-05
Identities = 33/87 (37%), Positives = 46/87 (51%), Gaps = 11/87 (12%)
Query: 1 MSNRFQSALGYVAPE-LACQILRVNEKCDVYGFGVMILEIVTGKRPVE----YGEDNVLI 55
+S R GY APE LA L V K DVY FGV++LE+++G+R ++ GE N++
Sbjct: 236 VSTRVMGTQGYAAPEYLATGHLSV--KSDVYSFGVVLLELLSGRRAIDKNQPVGEHNLV- 292
Query: 56 LNDHVR-VLLEHGNALECVDPSLMSEY 81
D R L L +DP L +Y
Sbjct: 293 --DWARPYLTNKRRLLRVMDPRLQGQY 317
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 42.7 bits (99), Expect = 1e-04
Identities = 21/48 (43%), Positives = 34/48 (70%), Gaps = 4/48 (8%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRP---VEYGEDNVL 54
GYV PE Q R + K DVY +GV++LE++TGK+P ++G++N++
Sbjct: 1056 GYVPPEYY-QSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLV 1102
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 42.7 bits (99), Expect = 1e-04
Identities = 21/48 (43%), Positives = 34/48 (70%), Gaps = 4/48 (8%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRP---VEYGEDNVL 54
GYV PE Q R + K DVY +GV++LE++TGK+P ++G++N++
Sbjct: 1056 GYVPPEYY-QSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLV 1102
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 42.0 bits (97), Expect = 2e-04
Identities = 19/47 (40%), Positives = 30/47 (63%), Gaps = 1/47 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE 47
+ R GY+APE A RV K DVY FGV+++E++TG++ ++
Sbjct: 749 IETRIAGTFGYLAPEYAVTG-RVTTKVDVYSFGVILMELITGRKSLD 794
>TML1_ARATH (P33543) Putative kinase-like protein TMKL1 precursor
Length = 674
Score = 41.6 bits (96), Expect = 3e-04
Identities = 19/42 (45%), Positives = 30/42 (71%), Gaps = 1/42 (2%)
Query: 10 GYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED 51
GY APEL ++ + N + DVY FG+++LEI+ GK+P + G +
Sbjct: 541 GYKAPELH-KMKKCNPRSDVYAFGILLLEILMGKKPGKSGRN 581
>M3K7_DROME (P83104) Putative mitogen-activated protein kinase
kinase kinase 7 (EC 2.7.1.37)
Length = 393
Score = 41.6 bits (96), Expect = 3e-04
Identities = 23/54 (42%), Positives = 33/54 (60%), Gaps = 4/54 (7%)
Query: 1 MSNR---FQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED 51
MSN Q L Y+APE A + L+ KCDVY FG+M+ E++T + P + E+
Sbjct: 159 MSNNKTDMQGTLRYMAPE-AIKHLKYTAKCDVYSFGIMLWELMTRQLPYSHLEN 211
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 40.8 bits (94), Expect = 5e-04
Identities = 29/87 (33%), Positives = 42/87 (47%), Gaps = 2/87 (2%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE-DNVLILNDH 59
+S LGY+ PE Q K DVY FGV++LE++T KRPV+ + L
Sbjct: 892 VSTDLVGTLGYIPPEYG-QASVATYKGDVYSFGVVLLELLTDKRPVDMCKPKGCRDLISW 950
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEV 86
V + A E DP + S+ + E+
Sbjct: 951 VVKMKHESRASEVFDPLIYSKENDKEM 977
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 40.4 bits (93), Expect = 7e-04
Identities = 19/39 (48%), Positives = 27/39 (68%), Gaps = 1/39 (2%)
Query: 9 LGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE 47
LGY+ PE Q K DVY FGV++LE++TG+RP++
Sbjct: 909 LGYIPPEYG-QASVATYKGDVYSFGVVLLELLTGRRPMD 946
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 40.4 bits (93), Expect = 7e-04
Identities = 26/61 (42%), Positives = 36/61 (58%), Gaps = 4/61 (6%)
Query: 6 QSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY--GEDNV-LILNDHVRV 62
+ LGY+APE L + K DVY +GV++LE++TG R E G D V +L VR+
Sbjct: 696 RGTLGYIAPEWVSS-LPITAKVDVYSYGVVLLELLTGTRVSELVGGTDEVHSMLRKLVRM 754
Query: 63 L 63
L
Sbjct: 755 L 755
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 40.0 bits (92), Expect = 0.001
Identities = 28/90 (31%), Positives = 45/90 (49%), Gaps = 9/90 (10%)
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLIL 56
+S R GY APE + K DVY +GV++LE+++G+R V+ GE ++
Sbjct: 237 VSTRIMGTYGYAAPEYLATG-HLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQKLV-- 293
Query: 57 NDHVRVLLEHGNAL-ECVDPSLMSEYPEDE 85
+ R LL + L +D L +Y +E
Sbjct: 294 -EWARPLLANKRKLFRVIDNRLQDQYSMEE 322
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,150,173
Number of Sequences: 164201
Number of extensions: 391915
Number of successful extensions: 1615
Number of sequences better than 10.0: 299
Number of HSP's better than 10.0 without gapping: 117
Number of HSP's successfully gapped in prelim test: 182
Number of HSP's that attempted gapping in prelim test: 1488
Number of HSP's gapped (non-prelim): 312
length of query: 126
length of database: 59,974,054
effective HSP length: 102
effective length of query: 24
effective length of database: 43,225,552
effective search space: 1037413248
effective search space used: 1037413248
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146632.13


