

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.5 - phase: 0 /pseudo
(637 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
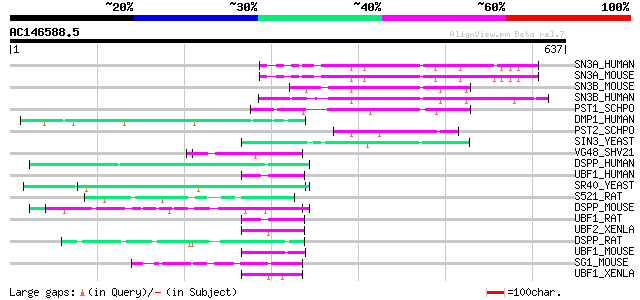
Score E
Sequences producing significant alignments: (bits) Value
SN3A_HUMAN (Q96ST3) Paired amphipathic helix protein Sin3a 101 7e-21
SN3A_MOUSE (Q60520) Paired amphipathic helix protein Sin3a 98 8e-20
SN3B_MOUSE (Q62141) Paired amphipathic helix protein Sin3b 69 5e-11
SN3B_HUMAN (O75182) Paired amphipathic helix protein Sin3b 67 1e-10
PST1_SCHPO (Q09750) Paired amphipathic helix protein pst1 (SIN3 ... 62 5e-09
DMP1_HUMAN (Q13316) Dentin matrix acidic phosphoprotein 1 precur... 58 7e-08
PST2_SCHPO (O13919) Paired amphipathic helix protein pst2 (SIN3 ... 54 1e-06
SIN3_YEAST (P22579) Paired amphipathic helix protein SIN3 53 3e-06
VG48_SHV21 (Q01033) Hypothetical gene 48 protein 51 8e-06
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 50 1e-05
UBF1_HUMAN (P17480) Nucleolar transcription factor 1 (Upstream b... 49 3e-05
SR40_YEAST (P32583) Suppressor protein SRP40 49 4e-05
S521_RAT (Q9QY02) Putative splicing factor YT521 (RA301-binding ... 49 4e-05
DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin... 49 5e-05
UBF1_RAT (P25977) Nucleolar transcription factor 1 (Upstream bin... 48 7e-05
UBF2_XENLA (P25980) Nucleolar transcription factor 2 (Upstream b... 47 1e-04
DSPP_RAT (Q62598) Dentin sialophosphoprotein precursor [Contains... 47 1e-04
UBF1_MOUSE (P25976) Nucleolar transcription factor 1 (Upstream b... 47 2e-04
SG1_MOUSE (P16014) Secretogranin I precursor (SgI) (Chromogranin... 47 2e-04
UBF1_XENLA (P25979) Nucleolar transcription factor 1 (Upstream b... 45 4e-04
>SN3A_HUMAN (Q96ST3) Paired amphipathic helix protein Sin3a
Length = 1273
Score = 101 bits (251), Expect = 7e-21
Identities = 98/379 (25%), Positives = 162/379 (41%), Gaps = 86/379 (22%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 826 AQRGDLSDVE---------EEEEEEMDVD-------EATGAVKKHN--GVGGSPPKSKLL 867
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 868 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 921
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + SLLDG+ D+S++ED
Sbjct: 922 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRSLLDGNIDSSQYED 981
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E +
Sbjct: 982 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEICVQVTDLYLAENNNGATGGQLNT 1039
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHDKPE--VTAVSID 543
+ S S + Y A L+ DEN +++ +L+I+L+D + + V A
Sbjct: 1040 QNSRSLLESTYQRKAEQLMSDENCFKLMFIQSQGQVQLTIELLDTEEENSDDPVEAERWS 1099
Query: 544 PNFSTYLHNDFLSVVPDSKE---KSGIFLKRNK---LKCARSDEL---------SSQVMD 588
Y+++D S P+ +E + +FL RN KC R E S + M+
Sbjct: 1100 DYVERYMNSDTTS--PELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGNSKKTME 1157
Query: 589 GIQVINGLECKIACGSSKV 607
+ ++ LEC+ S K+
Sbjct: 1158 NVDSLDKLECRFKLNSYKM 1176
>SN3A_MOUSE (Q60520) Paired amphipathic helix protein Sin3a
Length = 1282
Score = 97.8 bits (242), Expect = 8e-20
Identities = 95/385 (24%), Positives = 163/385 (41%), Gaps = 90/385 (23%)
Query: 287 SENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERF 346
++ GD+S E EE EE+ D D EA G ++ G G S P S+
Sbjct: 827 AQRGDLSDVE---------EEEEEEMDVD-------EATGAPKKHN--GVGGSPPKSKLL 868
Query: 347 LLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------RSAKINSSS 400
+ G +E +FY N+++Y+ RLHQ L R+ +I +
Sbjct: 869 FSNT------AAQKLRGMDEVYNLFYVNNNWYIFMRLHQILCLRLLRICSQAERQIEEEN 922
Query: 401 AEKKW---------------------RASNDTSSTDRYARFMNSLYSLLDGSSDNSKFED 439
E++W + D D Y F++ + LLDG+ D+S++ED
Sbjct: 923 REREWEREVLGIKRDKSDSPAIQLRLKEPMDVDVEDYYPAFLDMVRQLLDGNIDSSQYED 982
Query: 440 DCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYE----------QS 489
R + +Y+ FT+DKLI +V+QLQ + S DE+ ++ LY E S
Sbjct: 983 SLREMFTIHAYIAFTMDKLIQSIVRQLQHIVS--DEVCVQVTDLYLAENNNGATGGQLNS 1040
Query: 490 RKSGSFFDVVYHENARVLLHDENIYRI----ECSPTRLSIQLMDYGHD-KPEVTAVSIDP 544
+ S S + Y A L+ DEN +++ +L+++L+D + + +
Sbjct: 1041 QTSRSLVESAYQRKAEQLMSDENCFKLMFIQSQGQVQLTVELLDTEEENSDDPVEAEVWT 1100
Query: 545 NFSTYLHNDFL-------SVVPDSKE---KSGIFLKRNK---LKCARSDEL--------- 582
+S Y +D++ + P+ +E + +FL RN KC R E
Sbjct: 1101 RWSDYRWSDYVERYMSSDTTSPELREHLAQKPVFLPRNLRRIRKCQRGREQQEKEGKEGN 1160
Query: 583 SSQVMDGIQVINGLECKIACGSSKV 607
S + M+ ++ ++ LEC+ S K+
Sbjct: 1161 SKKTMENVESLDKLECRFKLNSYKM 1185
>SN3B_MOUSE (Q62141) Paired amphipathic helix protein Sin3b
Length = 954
Score = 68.6 bits (166), Expect = 5e-11
Identities = 60/253 (23%), Positives = 109/253 (42%), Gaps = 53/253 (20%)
Query: 322 GEAEGMADANDVEGDGAS----LPYSERFLLTAKPLVKYVSPVFHGKEENVQIFYGNDSF 377
G ++ AD D + D A P E+ A P V ++ +F+ N+++
Sbjct: 665 GTSDDSADERDRDRDSAEPERRRPTDEKPPADASPEPPKVL------DDVYSLFFANNNW 718
Query: 378 YVLFRLHQTLYERI---------------------------RSAKINSSSAEKKWRASND 410
Y RLHQTL R+ R K + E + + ++
Sbjct: 719 YFFLRLHQTLCARLLKIYRQAQKQLLEHRREQEREQLLCEGRREKAADPAMELRLKQPSE 778
Query: 411 TSSTDRYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVA 470
+ Y F++ + SLL+GS D +++ED R + +Y+ FT+DKL+ + +QL +
Sbjct: 779 VELEEYYPAFLDMVRSLLEGSIDPTQYEDTLREMFTIHAYIGFTMDKLVQNIARQLHHLV 838
Query: 471 SGTDEMDNKLLQLYAYEQSRKSG----------SFFDVVYHENARVLLHDENIYRIECSP 520
S D++ K+++LY EQ R + + + Y A + DEN +++
Sbjct: 839 S--DDVCLKVVELYLNEQQRGAAGGNLSSRCVRAARETSYQWKAERCMADENCFKVMFLQ 896
Query: 521 TR----LSIQLMD 529
R ++I+L+D
Sbjct: 897 RRGQVIMTIELLD 909
>SN3B_HUMAN (O75182) Paired amphipathic helix protein Sin3b
Length = 1130
Score = 67.4 bits (163), Expect = 1e-10
Identities = 86/387 (22%), Positives = 158/387 (40%), Gaps = 77/387 (19%)
Query: 286 ASENGDVSGTESADGEECSREEHEED--GDHDNKVESEGEAEGMADANDVEGDGASLPYS 343
ASE +S G+ E ++ G H + E +G A G A A + P+
Sbjct: 676 ASEESADEDRDSPQGQTTDPSERKKPAPGPHSSPPEEKG-AFGDAPATEQPPLPPPAPH- 733
Query: 344 ERFLLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERI------------ 391
KPL ++ +F+ N+++Y RLHQTL R+
Sbjct: 734 -------KPL-----------DDVYSLFFANNNWYFFLRLHQTLCSRLLKIYRQAQKQLL 775
Query: 392 ---------------RSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSSDNSK 436
R K + + E + + ++ + Y F++ + SLL+GS D ++
Sbjct: 776 EYRTEKEREKLLCEGRREKGSDPAMELRLKQPSEVELEEYYPAFLDMVRSLLEGSIDPTQ 835
Query: 437 FEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSG--- 493
+ED R + +YV FT+DKL+ + +QL + S D++ K+++LY E+ R +
Sbjct: 836 YEDTLREMFTIHAYVGFTMDKLVQNIARQLHHLVS--DDVCLKVVELYLNEKKRGAAGGN 893
Query: 494 -------SFFDVVYHENARVLLHDENIYRIECSPTR----LSIQLMDYGHDKPE--VTAV 540
+ + Y A + DEN +++ + ++I+L+D + E V
Sbjct: 894 LSSRCVRAARETSYQWKAERCMADENCFKVMFLQRKGQVIMTIELLDTEEAQTEDPVEVQ 953
Query: 541 SIDPNFSTYLHNDFLSVVP-DSKEKSGIFLKRNKLKCAR------SDELSSQVMDGIQVI 593
+ Y+ + S P + +FL+RN K R + L + + +
Sbjct: 954 HLARYVEQYVGTEGASSSPTEGFLLKPVFLQRNLKKFRRRWQSEQARALRGEARSSWKRL 1013
Query: 594 NGLE--CKIACGSSKVCNHLLLHIVKN 618
G+E C + C K+ H ++ IV +
Sbjct: 1014 VGVESACDVDC-RFKLSTHKMVFIVNS 1039
>PST1_SCHPO (Q09750) Paired amphipathic helix protein pst1 (SIN3
homolog 1)
Length = 1522
Score = 62.0 bits (149), Expect = 5e-09
Identities = 72/284 (25%), Positives = 115/284 (40%), Gaps = 62/284 (21%)
Query: 277 HRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEG- 335
+RSS + SE+ + + S E S EHEE+ D + V S GE D +G
Sbjct: 931 NRSSVKEDYVSESTERTPDASEIDEHIS--EHEENDDESSSVFSTGEVWVNCKFTDTDGS 988
Query: 336 ---DGASLPYSERFLLTAKPLVKYVSPVFHGKEENVQIFYGNDSFYVLFRLHQTLYERIR 392
DG L + +V +GN S Y FRL TLY R+
Sbjct: 989 LLDDGTKL-----------------------SDRSVYNLFGNMSLYCFFRLFHTLYSRLE 1025
Query: 393 SAKINSSSAEKKWR--ASNDTS-------------------STDRYARFMNSLYSLLDGS 431
K A K SN + + Y + + L++G
Sbjct: 1026 EIKNLEQMAYSKQHDVKSNPVAVELGLVRHPSERLGFALPTADTVYEQAIQLCERLMEGE 1085
Query: 432 SDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQ--- 488
D + FED R + G ++ L+T++KL+ ++KQL +V + + +L Q++ Y +
Sbjct: 1086 IDQNGFEDALRCLYGIHAFRLYTVEKLVTSIIKQLHSVTT-----NRRLAQVFMYYEKDR 1140
Query: 489 -SRKSGSFFDVVYH-ENARVLLHDENIYRIEC-SPTRLS-IQLM 528
R++ ++Y + DEN+ I+ S TR S I+LM
Sbjct: 1141 VQRRTSPRQQIMYRIQTETAFGPDENLCCIDWNSQTRQSAIRLM 1184
>DMP1_HUMAN (Q13316) Dentin matrix acidic phosphoprotein 1 precursor
(Dentin matrix protein-1) (DMP-1)
Length = 513
Score = 58.2 bits (139), Expect = 7e-08
Identities = 77/355 (21%), Positives = 142/355 (39%), Gaps = 36/355 (10%)
Query: 13 GVPSRLRVPEDTEDAVKAKKDSAKTG-------TASIAKGDSSPGVGATVMSPKNTFEQS 65
G SRL ED++D ++A ++SA G T+ + D+ V + +T ++S
Sbjct: 129 GGNSRLGSDEDSDDTIQASEESAPQGQDSAQDTTSESRELDNEDRVDSKPEGGDST-QES 187
Query: 66 NSCKEW--------QTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVS 117
S + W ++G G +D+ ++SD G+S + + +
Sbjct: 188 ESEEHWVGGGSDGESSHGDGSELDDEGMQSDDPESIRSERGNSRMNSAGMKSKESGENSE 247
Query: 118 RVNKQDHSIEQLV---NANVSMSSRVEQSNGRTNI--NNASGLAATLSRPGYVYREGGLD 172
+ N QD QL+ + + SR+ + + R+ + NN + S R+ GL
Sbjct: 248 QANTQDSGGSQLLEHPSRKIFRKSRISEEDDRSELDDNNTMEEVKSDSTENSNSRDTGLS 307
Query: 173 LPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFK-------SEREEGELSPNGDFE 225
P + ++ D+ N + E ES SE +E +S +
Sbjct: 308 QPRRDSKGDSQEDSKENLSQEESQNVDGPSSESSQEANLSSQENSSESQEEVVSESRGDN 367
Query: 226 EDNFAVYANAGLEAVHKGKDCN-TSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSE 284
D Y ++ +D + T H ++ E+ E+ + S+ SP S +D
Sbjct: 368 PDPTTSYVEDQEDSDSSEEDSSHTLSHSKSESREEQADSESSESLNFSEESPE-SPEDEN 426
Query: 285 NASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
++S+ G S + SA+ + S E H E+ D D S+ + D+N E +S
Sbjct: 427 SSSQEGLQSHSSSAESQ--SEESHSEEDDSD----SQDSSRSKEDSNSTESKSSS 475
>PST2_SCHPO (O13919) Paired amphipathic helix protein pst2 (SIN3
homolog 2)
Length = 1075
Score = 53.9 bits (128), Expect = 1e-06
Identities = 48/166 (28%), Positives = 76/166 (44%), Gaps = 24/166 (14%)
Query: 372 YGNDSFYVLFRLHQTLYERI---------------RSAKINSSSAEKKWRAS-NDTSST- 414
+G S YV FRL LYER+ R S +K WR ND S
Sbjct: 744 FGYSSMYVFFRLFNLLYERLYELQRLEDQVSIIQQRIIPNPVSQKQKIWRDRWNDLSDVP 803
Query: 415 DRYARFMNS---LYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVAS 471
D + N+ + L+ G D S FED R G ++Y ++T+DKL++ KQ+ + S
Sbjct: 804 DEKTHYENTYVMILRLIYGIVDQSAFEDYLRFYYGNKAYKIYTIDKLVWSAAKQVHHIVS 863
Query: 472 -GTDEMDNKLLQLYAYEQSRKSGSFFDVVYH-ENARVLLHDENIYR 515
G + L++ + +K ++ D +Y E ++L DE ++R
Sbjct: 864 DGKYKFVTSLVEQNSSASPKK--NYDDFLYRLEIEKLLNPDEILFR 907
>SIN3_YEAST (P22579) Paired amphipathic helix protein SIN3
Length = 1536
Score = 52.8 bits (125), Expect = 3e-06
Identities = 61/287 (21%), Positives = 114/287 (39%), Gaps = 36/287 (12%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSR--EEHEEDGDHDNKVESEGEA 324
+N ES GS SS S ++S + + ++EDG E E
Sbjct: 1038 QNVSESSGSDDGSSIASRKRPYQQEMSLLDILHRSRYQKLKRSNDEDGKVPQLSEPPEEE 1097
Query: 325 EGMADANDVEGDGASLPYSERFLLTAKPLVKYVSPVFHGKEENVQIF--YGNDSFYVLFR 382
+ ++ + A P+ LT + + S G +N IF + N + Y+ FR
Sbjct: 1098 PNTIEEEELIDEEAKNPW-----LTGNLVEEANS---QGIIQNRSIFNLFANTNIYIFFR 1149
Query: 383 LHQTLYERIRSAKINSSSAEKKWRASN--------------------DTSSTDRYARFMN 422
T+YER+ K + K+ + D D Y + +
Sbjct: 1150 HWTTIYERLLEIKQMNERVTKEINTRSTVTFAKDLDLLSSQLSEMGLDFVGEDAYKQVLR 1209
Query: 423 SLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQ 482
L++G ++ FE+ R +++ L+T+DK+ LVK + TD +++
Sbjct: 1210 LSRRLINGDLEHQWFEESLRQAYNNKAFKLYTIDKVTQSLVKHAHTLM--TDAKTAEIMA 1267
Query: 483 LYAYEQSRKSGSFFD-VVYHENARV-LLHDENIYRIECSPTRLSIQL 527
L+ +++ + S D ++Y R + + EN++RIE L + +
Sbjct: 1268 LFVKDRNASTTSAKDQIIYRLQVRSHMSNTENMFRIEFDKRTLHVSI 1314
>VG48_SHV21 (Q01033) Hypothetical gene 48 protein
Length = 797
Score = 51.2 bits (121), Expect = 8e-06
Identities = 34/127 (26%), Positives = 53/127 (40%), Gaps = 9/127 (7%)
Query: 210 KSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEND 269
K E +EG+ GD ED + + G + +G+D + E G E +
Sbjct: 517 KDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDE-------GDEGE 569
Query: 270 DESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMAD 329
DE + D+ E+ + G+ G + D E E+ ++GD EGE EG D
Sbjct: 570 DEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEG--D 627
Query: 330 ANDVEGD 336
+ EGD
Sbjct: 628 EGEDEGD 634
Score = 45.8 bits (107), Expect = 3e-04
Identities = 37/137 (27%), Positives = 57/137 (41%), Gaps = 12/137 (8%)
Query: 204 ESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGV 263
E G + + E E GD ED G + +G+D + + E + G
Sbjct: 558 EDEGEDEGDEGEDEGEDEGDEGEDE-------GEDEGDEGEDEGEDEGDEGEDEGEDEGD 610
Query: 264 AGGENDDESDGSPHRSS---DDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVES 320
G E +DE D D+ + + GD E +GE+ +E E++GD ++ E
Sbjct: 611 EGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGED-EGDEGEDEGDEGDEGED 669
Query: 321 EG-EAEGMADANDVEGD 336
EG E E D + EGD
Sbjct: 670 EGDEGEDEGDEGEDEGD 686
Score = 43.1 bits (100), Expect = 0.002
Identities = 42/174 (24%), Positives = 67/174 (38%), Gaps = 11/174 (6%)
Query: 168 EGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEED 227
+ G D EG D D G ED +E G + E +EG+ + E D
Sbjct: 439 DDGEDEGEDEGEDEG--DEGDEGDEGEDEGEDEDDEEDEG--EDEGDEGDEGEDEGDEGD 494
Query: 228 NFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENAS 287
+ G E +G + + + + +E G G E +DE + D+ +
Sbjct: 495 EGEDEGDEGDEGKDEGDEGDEGKDEGDEGDE---GDEGDEGEDEWEDEGDEGEDEGDEGE 551
Query: 288 ENGDVSGTESAD----GEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDG 337
+ GD E D GE+ +E +E D EGE EG + ++ E +G
Sbjct: 552 DEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEG 605
Score = 43.1 bits (100), Expect = 0.002
Identities = 37/137 (27%), Positives = 58/137 (42%), Gaps = 9/137 (6%)
Query: 204 ESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGV 263
E ++ E +EGE GD ED + G + +G+D + + E +
Sbjct: 531 EGEDEWEDEGDEGE--DEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGED--- 585
Query: 264 AGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSRE---EHEEDGDHDNKVES 320
G E +DE + D+ E+ + GD E +GE+ E E +E D ++ E
Sbjct: 586 EGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGED 645
Query: 321 EG-EAEGMADANDVEGD 336
EG E E D + EGD
Sbjct: 646 EGDEGEDEGDEGEDEGD 662
Score = 40.8 bits (94), Expect = 0.011
Identities = 23/72 (31%), Positives = 37/72 (50%), Gaps = 5/72 (6%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDV--SGTESADGEECSREEHEEDGDHDNKVESEGEA 324
E +DE D D+ E+ + GD G + + E+ +E E++GD ++ E EG+
Sbjct: 433 EGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGD- 491
Query: 325 EGMADANDVEGD 336
EG D + EGD
Sbjct: 492 EG--DEGEDEGD 501
Score = 40.4 bits (93), Expect = 0.014
Identities = 26/84 (30%), Positives = 37/84 (43%), Gaps = 6/84 (7%)
Query: 254 NRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGD 313
NR EE+ G + DE + D + + G+ G E +G+E E +ED +
Sbjct: 413 NRDEEEDEGEDEEDEKDEKEEGEDEGDDGEDEGEDEGEDEGDEGDEGDEGEDEGEDEDDE 472
Query: 314 HDNKVESEGEAEG-MADANDVEGD 336
D EGE EG D + EGD
Sbjct: 473 ED-----EGEDEGDEGDEGEDEGD 491
Score = 40.4 bits (93), Expect = 0.014
Identities = 31/125 (24%), Positives = 55/125 (43%), Gaps = 14/125 (11%)
Query: 214 EEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESD 273
+EG+ G E D + G + +G + + ++ E++ G E +DE D
Sbjct: 498 DEGDEGDEGKDEGDE----GDEGKDEGDEGDEGDEGDEGEDEWEDE-----GDEGEDEGD 548
Query: 274 GSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDG-DHDNKVESEGEAEGMADAND 332
D+ + + G+ G E D E +E E++G D ++ E EGE EG ++
Sbjct: 549 ----EGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDE 604
Query: 333 VEGDG 337
E +G
Sbjct: 605 GEDEG 609
Score = 40.4 bits (93), Expect = 0.014
Identities = 36/136 (26%), Positives = 54/136 (39%), Gaps = 12/136 (8%)
Query: 204 ESVGHFKSEREEGELSPNGDFEEDNFAVYANAGL-EAVHKGKDCNTSQHYQNRREEQICG 262
E G + + E E GD ED + G E +G + + + + E++
Sbjct: 569 EDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEGEDE--- 625
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSRE--EHEEDGDHDNKVES 320
G E +DE D D+ + + GD E +GE+ E E E++GD
Sbjct: 626 --GDEGEDEGD----EGEDEGDEGEDEGDEGEDEGDEGEDEGDEGDEGEDEGDEGEDEGD 679
Query: 321 EGEAEGMADANDVEGD 336
EGE EG EGD
Sbjct: 680 EGEDEGDEGDEGDEGD 695
Score = 39.7 bits (91), Expect = 0.024
Identities = 38/138 (27%), Positives = 57/138 (40%), Gaps = 9/138 (6%)
Query: 204 ESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGV 263
E G + + E E GD ED + G E +G D + + E
Sbjct: 580 EDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEG-DEGEDEGDEGEDEGDEGED 638
Query: 264 AGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVES--- 320
G E +DE D D+ E+ + GD E +GE+ +E E++GD ++ +
Sbjct: 639 EGDEGEDEGDEGEDEG-DEGEDEGDEGDEGEDEGDEGED-EGDEGEDEGDEGDEGDEGDE 696
Query: 321 --EGEAEGMADANDVEGD 336
EGE EG + D EGD
Sbjct: 697 GDEGEDEGEDEGED-EGD 713
Score = 39.3 bits (90), Expect = 0.032
Identities = 31/125 (24%), Positives = 52/125 (40%), Gaps = 11/125 (8%)
Query: 214 EEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESD 273
+EG+ G+ E D + G + +G+D + E G E +DE D
Sbjct: 607 DEGDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEGED----EGDEGEDEGD 662
Query: 274 GSPHRSS--DDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADAN 331
D+ E+ + G+ G E +G +E +E + +++ E EGE EG
Sbjct: 663 EGDEGEDEGDEGEDEGDEGEDEGDEGDEG-----DEGDEGDEGEDEGEDEGEDEGDEGTK 717
Query: 332 DVEGD 336
D EG+
Sbjct: 718 DKEGN 722
Score = 35.8 bits (81), Expect = 0.35
Identities = 24/94 (25%), Positives = 41/94 (43%), Gaps = 10/94 (10%)
Query: 244 KDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEEC 303
+D Y+ EE +++E +G +D ++ E G+ G + D E
Sbjct: 397 RDDRDKDEYELENEEY------NRDEEEDEGE---DEEDEKDEKEEGEDEGDDGEDEGED 447
Query: 304 SREEHEEDGDHDNKVESEGEAE-GMADANDVEGD 336
E+ ++GD ++ E EGE E D + EGD
Sbjct: 448 EGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGD 481
Score = 35.0 bits (79), Expect = 0.60
Identities = 43/160 (26%), Positives = 67/160 (41%), Gaps = 41/160 (25%)
Query: 183 RPDTSTNGAIIEDTKAHRCHKESVGHF----KSEREEGELSPNGDFEEDNFAVYANAGLE 238
R D + +E+ + +R +E G K E+EEGE GD ED
Sbjct: 397 RDDRDKDEYELENEEYNRDEEEDEGEDEEDEKDEKEEGE--DEGDDGED----------- 443
Query: 239 AVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESA 298
+G+D E + G G E D+ D +D ++ + G+ G E
Sbjct: 444 ---EGED-----------EGEDEGDEGDEGDEGED-----EGEDEDDEEDEGEDEGDEGD 484
Query: 299 DGEECSRE--EHEEDGDHDNKVESEGEAEGMADANDVEGD 336
+GE+ E E E++GD ++ + EG+ EG D EGD
Sbjct: 485 EGEDEGDEGDEGEDEGDEGDEGKDEGD-EG--DEGKDEGD 521
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 50.4 bits (119), Expect = 1e-05
Identities = 56/324 (17%), Positives = 128/324 (39%), Gaps = 6/324 (1%)
Query: 23 DTEDAVKAKKDSAKTGTASIA-KGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKE 81
D+ D+ + +S+ +S + DSS + + ++ + S+S ++ K
Sbjct: 565 DSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSKS 624
Query: 82 DDCL-KSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRV 140
D +SD S +++ S + + N+ D + N D S + +++ S SS
Sbjct: 625 DSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSDSS-DSSDSSDSSSSSDS 683
Query: 141 EQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHR 200
S+ +N +++S + + + D SS+ +DS+ ++S + + +
Sbjct: 684 SSSSDSSNSSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDS 743
Query: 201 CHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQI 260
+ + + S + D + + + ++ + N+S +
Sbjct: 744 SDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSNSSDSSDSSNSSDS 803
Query: 261 CGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVES 320
+ + +S S +S DS N+S++ D S S+D + S D D N+ +S
Sbjct: 804 SNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSS--NSSDSSDSSDSSDSSDSDSSNRSDS 861
Query: 321 EGEAEGMADANDVEGDGASLPYSE 344
++ +D++D S S+
Sbjct: 862 SNSSDS-SDSSDSSNSSDSSDSSD 884
Score = 44.3 bits (103), Expect = 0.001
Identities = 54/310 (17%), Positives = 118/310 (37%), Gaps = 18/310 (5%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ ++ + DS+ + +S DSS ++ S + S++ + + D
Sbjct: 723 DSSNSNSSDSDSSNSSDSS----DSSDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSD 778
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD + + S + + N+S + S + D S N S SS
Sbjct: 779 SSNSSDSNDSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSS-------NSSDSSNSSD 831
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ +N +++S + + + SS +DS+ S+N + D+
Sbjct: 832 SSDSSNSSDSSDSSDSSDSSD---SDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDSSDS 888
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
ES S+ + S + D + + + ++ + + N+S + + +
Sbjct: 889 NESSN--SSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDN--SNSSDSSNS 944
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
++ D S+ S +S DS N+S++ D + ++S+D S D +
Sbjct: 945 SDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSS 1004
Query: 323 EAEGMADAND 332
+ +D++D
Sbjct: 1005 NSSDSSDSSD 1014
Score = 43.9 bits (102), Expect = 0.001
Identities = 50/310 (16%), Positives = 109/310 (35%), Gaps = 15/310 (4%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ + S + ++ + DSS ++ S + S++ + +
Sbjct: 831 DSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSS----DSS 886
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
D +S S +++ SS + N+S + S + S N+N S SS
Sbjct: 887 DSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSD 946
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ ++ +N+S + G SS+ +DS D+S + + + +
Sbjct: 947 SSDSSDSSNSSDSS-----------NSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSS 995
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
S S + S + ++ ++ D + S N +
Sbjct: 996 DSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSS 1055
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
+ +D SSD S+++ + ++S+D E S D +
Sbjct: 1056 DSSDSSDSSGSSDSSDSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSSDSSDSSDSS 1115
Query: 323 EAEGMADAND 332
++ +D++D
Sbjct: 1116 DSSDSSDSSD 1125
Score = 43.1 bits (100), Expect = 0.002
Identities = 49/310 (15%), Positives = 104/310 (32%), Gaps = 16/310 (5%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ + ++ + S DSS ++ S + S+S ++ D
Sbjct: 811 DSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSN----SSD 866
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD S + S + N N+S + S + D S + + S+ +
Sbjct: 867 SSDSSDSSNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDS 926
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S + +N++ ++ S + SS DS+ S++ + + +
Sbjct: 927 SESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSS 986
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
S S+ + S N D+ D + S N +
Sbjct: 987 DSSDSSDSSDSSDSSDSSNSSDSSDS------------SDSSDSSNSSDSSNSSDSSDSS 1034
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
+ +D + SSD S+++ + ++S+D + S D +
Sbjct: 1035 DSSDSSDSSDSSNSSDSSDSSDSSDSSDSSGSSDSSDSSDSSDSSDSSDSSDSSDSSDSS 1094
Query: 323 EAEGMADAND 332
E+ +D++D
Sbjct: 1095 ESSDSSDSSD 1104
Score = 42.7 bits (99), Expect = 0.003
Identities = 48/312 (15%), Positives = 117/312 (37%), Gaps = 9/312 (2%)
Query: 33 DSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVP 92
DS+ + +S + + + N+ + SNS ++ + S S
Sbjct: 772 DSSDSSDSSNSSDSNDSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSD 831
Query: 93 KTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNA 152
+++ SS + + S + S + +S + +++ S SS S+ ++ N +
Sbjct: 832 SSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSNES 891
Query: 153 SGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSE 212
S + + D SS+ ++S+ S+N + ++ + S S
Sbjct: 892 SNSSDSSDS------SNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSS 945
Query: 213 REEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDES 272
+ + N +N+ + D + S + + + + +D +
Sbjct: 946 DSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSN 1005
Query: 273 DGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADAND 332
SSD S++++ + + ++S+D + S + + D N +S ++ +D++D
Sbjct: 1006 SSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSS--DSSDSSDSSNSSDSSDSSDS-SDSSD 1062
Query: 333 VEGDGASLPYSE 344
G S S+
Sbjct: 1063 SSGSSDSSDSSD 1074
Score = 42.4 bits (98), Expect = 0.004
Identities = 30/123 (24%), Positives = 54/123 (43%), Gaps = 11/123 (8%)
Query: 216 GELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGS 275
G+ S N D +DN + G E D + S N + G G +++D+SD
Sbjct: 489 GDASYNSDESKDNGNGSDSKGAE-----DDDSDSTSDTNNSDSNGNGNNGNDDNDKSDSG 543
Query: 276 PHRS------SDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMAD 329
+S S DS N+S++ D S ++S+D S + + D+ ++ +D
Sbjct: 544 KGKSDSSDSDSSDSSNSSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDSSNSSD 603
Query: 330 AND 332
++D
Sbjct: 604 SSD 606
Score = 40.0 bits (92), Expect = 0.019
Identities = 24/84 (28%), Positives = 41/84 (48%), Gaps = 14/84 (16%)
Query: 270 DESDGSPHRSSDDSENASENGDVSGTESADGEECSR-------EEHEEDGDHDNKVESEG 322
D SDGSP + D + +GD E+ +G++ S ++H ++ DHD+ +
Sbjct: 240 DNSDGSPSGNGADEDEDEGSGDDEDEEAGNGKDSSNNSKGQEGQDHGKEDDHDSSIGQNS 299
Query: 323 EA------EGMADA-NDVEGDGAS 339
++ EG D N+V+GD S
Sbjct: 300 DSKEYYDPEGKEDPHNEVDGDKTS 323
Score = 38.9 bits (89), Expect = 0.042
Identities = 50/306 (16%), Positives = 112/306 (36%), Gaps = 10/306 (3%)
Query: 33 DSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVP 92
D++ + +S + S + N+ + SNS +N + S S
Sbjct: 934 DNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSD 993
Query: 93 KTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNA 152
+++ SS N+S + S + +S + +++ S SS S+ ++ +N+
Sbjct: 994 SSDSSDSSDSS-----NSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNS 1048
Query: 153 SGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTK-AHRCHKESVGHFKS 211
S + + G D SS+ +DS+ S++ + D+ + S S
Sbjct: 1049 SDSSDSSDSSDSSDSSGSSD--SSDSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSS 1106
Query: 212 EREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDE 271
+ + S + D+ ++ D + S + + + +D
Sbjct: 1107 DSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSS 1166
Query: 272 SDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADAN 331
SSD S+++ N ++S+D + S + D + +S ++ +D+
Sbjct: 1167 DSSDSSDSSDSSDSSDSNESSDSSDSSDSSDSSNS--SDSSDSSDSSDSTSDSNDESDSQ 1224
Query: 332 DVEGDG 337
G+G
Sbjct: 1225 SKSGNG 1230
Score = 38.1 bits (87), Expect = 0.071
Identities = 52/331 (15%), Positives = 118/331 (34%), Gaps = 14/331 (4%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ ++ S + +++ + DSS ++ S + S+ +N
Sbjct: 884 DSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDSSE----SSNSSDNSNSS 939
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
D S S +++ SS + + S S + S +++ S SS
Sbjct: 940 DSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDSSNSSDSSDSSDSSDSSD 999
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLP-SSEGADSTRPDTSTNGAIIEDTKAHRC 201
S+ +N +++S + + D SS+ +DS+ S+N + D+
Sbjct: 1000 SSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSD 1059
Query: 202 HKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKG--------KDCNTSQHYQ 253
+S G S + + S + D + + + ++ E+ D + S
Sbjct: 1060 SSDSSGSSDSS-DSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSSDSSDSSDSSDSS 1118
Query: 254 NRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGD 313
+ + + +D SSD S+++ + ++S+D + S D
Sbjct: 1119 DSSDSSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSS 1178
Query: 314 HDNKVESEGEAEGMADANDVEGDGASLPYSE 344
+ ++ +D++D S S+
Sbjct: 1179 DSSDSNESSDSSDSSDSSDSSNSSDSSDSSD 1209
Score = 35.4 bits (80), Expect = 0.46
Identities = 56/318 (17%), Positives = 124/318 (38%), Gaps = 18/318 (5%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ + DS+ +G +S + DSS + N+ + S+S ++ D
Sbjct: 949 DSSDSSNSS-DSSNSGDSSNSS-DSSDSNSSDSSDSSNSSDSSDSSDSSDSSD----SSD 1002
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD S + S++ + ++S + S + S +++ S SS
Sbjct: 1003 SSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSD 1062
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+G ++ +++S + + D SSE +DS+ S++ + D+
Sbjct: 1063 SSGSSDSSDSSDSSDSSDSSDSSDSSDSSD--SSESSDSSDSSDSSDSSDSSDSSDSSDS 1120
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
+S S+ + S + D+ ++ D + S + +
Sbjct: 1121 SDS-----SDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSS 1175
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEED----GDHDNKV 318
+ +D SSD S++++ + ++S+D S +E + ++N
Sbjct: 1176 DSSDSSDSNESSDSSDSSDSSDSSNSSDSSDSSDSSDSTSDSNDESDSQSKSGNGNNNGS 1235
Query: 319 ESEGEAEGMADANDVEGD 336
+S+ ++EG +D+N D
Sbjct: 1236 DSDSDSEG-SDSNHSTSD 1252
Score = 33.5 bits (75), Expect = 1.7
Identities = 74/352 (21%), Positives = 126/352 (35%), Gaps = 57/352 (16%)
Query: 21 PEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSC-KEWQTNGVGGV 79
P EDA D + +G + D G + +NS +E Q +G
Sbjct: 231 PGTGEDAGLDNSDGSPSGNGADEDEDEGSGDDEDEEAGNGKDSSNNSKGQEGQDHG---- 286
Query: 80 KEDDCLKS--------------DRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHS 125
KEDD S + P E G T + + +A IP++
Sbjct: 287 KEDDHDSSIGQNSDSKEYYDPEGKEDPHNEVDGDKTSKSEEN-SAGIPED-----NGSQR 340
Query: 126 IEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDL--PSSEGADSTR 183
IE N S RVE + + +A G + ++ G+++ PSS + T+
Sbjct: 341 IEDTQKLNHRESKRVENRITKESETHAVGKS----------QDKGIEIKGPSSGNRNITK 390
Query: 184 PDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELS-PNGDFEEDNFAVYANAGLEAVHK 242
N ++ K G+ K++ E + P E N ++N G ++
Sbjct: 391 EVGKGNEG--KEDKGQHGMILGKGNVKTQGEVVNIEGPGQKSEPGNKVGHSNTGSDSNSD 448
Query: 243 GKDCNTSQHYQNRREEQICGVAGGENDD---ESD------GSPHRSSDDSENASENGDVS 293
G D + ++ NDD ESD G +SD+S++ D
Sbjct: 449 GYDSYDFDDKSMQGDDPNSSDESNGNDDANSESDNNSSSRGDASYNSDESKDNGNGSDSK 508
Query: 294 GTESADGEECSREEHEE------DGDHDNKVESEGEAEGMADANDVEGDGAS 339
G E D + S + + +G+ DN G +G +D++D + +S
Sbjct: 509 GAEDDDSDSTSDTNNSDSNGNGNNGNDDNDKSDSG--KGKSDSSDSDSSDSS 558
Score = 33.5 bits (75), Expect = 1.7
Identities = 17/74 (22%), Positives = 31/74 (40%)
Query: 266 GENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAE 325
G DD+SD + ++ DS NG+ +S G+ S + D N +S ++
Sbjct: 509 GAEDDDSDSTSDTNNSDSNGNGNNGNDDNDKSDSGKGKSDSSDSDSSDSSNSSDSSDSSD 568
Query: 326 GMADANDVEGDGAS 339
+ ++ D S
Sbjct: 569 SDSSDSNSSSDSDS 582
Score = 32.7 bits (73), Expect = 3.0
Identities = 35/179 (19%), Positives = 68/179 (37%), Gaps = 17/179 (9%)
Query: 175 SSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYAN 234
SS G S D S + D+K ++ S+ + + NG+ D+ +
Sbjct: 486 SSRGDASYNSDESKDNGNGSDSKGA---EDDDSDSTSDTNNSDSNGNGNNGNDDNDKSDS 542
Query: 235 AGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSD-------DSENAS 287
++ D + S + + + + + +SD S SSD DS N+S
Sbjct: 543 GKGKSDSSDSDSSDSSNSSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDSSNSS 602
Query: 288 ENGDVS-------GTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
++ D S ++S+D + S + + D D+K +S + N D ++
Sbjct: 603 DSSDSSDSSDSSDSSDSSDSKSDSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSN 661
Score = 32.7 bits (73), Expect = 3.0
Identities = 49/295 (16%), Positives = 109/295 (36%), Gaps = 15/295 (5%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ + + + S DSS ++ S + S+ + + D
Sbjct: 970 DSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSD 1029
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD S + S++ + ++S + S + S + +++ S SS
Sbjct: 1030 SSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSGSSDSSDSSDSSDSSDSSDSSDSSD 1089
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ + +++S + + + SS+ +DS+ S+N + D+
Sbjct: 1090 SSDSSESSDSSDSSDSSDSS-----DSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDS 1144
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
+S S + + S + D + + + ++ ++ + ++S +
Sbjct: 1145 SDSSDSSDSS-DSSDSSDSSDSSDSSDSSDSSDSSDSSDSNESSDSSDSSDSSDSSN--- 1200
Query: 263 VAGGENDDESDGSPHRSSD----DSENASENGDVSGTESADGEECSREEHEEDGD 313
++ D SD S S DS++ S NG+ +G++S E S H D
Sbjct: 1201 --SSDSSDSSDSSDSTSDSNDESDSQSKSGNGNNNGSDSDSDSEGSDSNHSTSDD 1253
Score = 31.6 bits (70), Expect = 6.6
Identities = 37/198 (18%), Positives = 71/198 (35%), Gaps = 14/198 (7%)
Query: 145 GRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKE 204
G+ N+ G + PG G + G +T D++++G D K
Sbjct: 410 GKGNVKT-QGEVVNIEGPGQKSEPG-----NKVGHSNTGSDSNSDGYDSYDFD----DKS 459
Query: 205 SVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVA 264
G + +E + + + E DN + + KD + ++ +
Sbjct: 460 MQGDDPNSSDESNGNDDANSESDNNSSSRGDASYNSDESKDNGNGSDSKGAEDDDSDSTS 519
Query: 265 GGENDDESDGSPHRSSDD---SENASENGDVSGTESADGEECSREEHEEDGDHDNKVESE 321
N D S+G+ + +DD S++ D S ++S+D S D D + S
Sbjct: 520 DTNNSD-SNGNGNNGNDDNDKSDSGKGKSDSSDSDSSDSSNSSDSSDSSDSDSSDSNSSS 578
Query: 322 GEAEGMADANDVEGDGAS 339
+D++D +S
Sbjct: 579 DSDSSDSDSSDSSDSDSS 596
>UBF1_HUMAN (P17480) Nucleolar transcription factor 1 (Upstream
binding factor 1) (UBF-1) (Autoantigen NOR-90)
Length = 764
Score = 49.3 bits (116), Expect = 3e-05
Identities = 24/72 (33%), Positives = 40/72 (55%), Gaps = 6/72 (8%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEG 326
E DDE+ S D SE++SE+ ES DG+E ++ +ED D D+ + + E+EG
Sbjct: 694 EEDDENGDSSEDGGDSSESSSED------ESEDGDENEEDDEDEDDDEDDDEDEDNESEG 747
Query: 327 MADANDVEGDGA 338
+ ++ GD +
Sbjct: 748 SSSSSSSSGDSS 759
Score = 44.3 bits (103), Expect = 0.001
Identities = 27/76 (35%), Positives = 37/76 (48%), Gaps = 5/76 (6%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGD---HDNKVESEGE 323
E DDE D D+ E ENGD S + D E S E+ EDGD D++ E + E
Sbjct: 678 EEDDEED-EDDEDEDEEEEDDENGD-SSEDGGDSSESSSEDESEDGDENEEDDEDEDDDE 735
Query: 324 AEGMADANDVEGDGAS 339
+ + N+ EG +S
Sbjct: 736 DDDEDEDNESEGSSSS 751
Score = 32.0 bits (71), Expect = 5.1
Identities = 22/93 (23%), Positives = 35/93 (36%), Gaps = 6/93 (6%)
Query: 210 KSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEND 269
KSE EE + D +ED G + G +S ++ ++ E D
Sbjct: 674 KSESEEDDEEDEDDEDEDEEEEDDENGDSSEDGGDSSESSSEDESEDGDE------NEED 727
Query: 270 DESDGSPHRSSDDSENASENGDVSGTESADGEE 302
DE + +D +N SE S + S D +
Sbjct: 728 DEDEDDDEDDDEDEDNESEGSSSSSSSSGDSSD 760
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 48.9 bits (115), Expect = 4e-05
Identities = 61/324 (18%), Positives = 122/324 (36%), Gaps = 28/324 (8%)
Query: 17 RLRVPEDTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGV 76
+++V E + +VK K+ K+ ++S + SS ++ S ++ E S+S ++
Sbjct: 5 KIKVDEVPKLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSS- 63
Query: 77 GGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSM 136
SD S SS+ + ++S E S + S ++ S
Sbjct: 64 ---------SSDSSDSSDSESSSSSSSSSSSSSSSSDSESSSESDSSSS-----GSSSSS 109
Query: 137 SSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDT 196
SS ++S+ + + + A RE + T P++S++
Sbjct: 110 SSSSDESSSESESEDETKKRA---------RESDNEDAKETKKAKTEPESSSSSESSSSG 160
Query: 197 KAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRR 256
+ ES S+ S + + +++ + D ++S +
Sbjct: 161 SSSSSESESGSESDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSD 220
Query: 257 EEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDN 316
+ + +D +SD SS DS+++ + S ++S+ E S + + D D D+
Sbjct: 221 SDSSSSSSSSSSDSDSDSD---SSSDSDSSGSSDSSSSSDSSSDESTSSDSSDSDSDSDS 277
Query: 317 KVESEGEA-EGMADANDVEGDGAS 339
SE E E AD + E AS
Sbjct: 278 GSSSELETKEATADESKAEETPAS 301
Score = 44.7 bits (104), Expect = 8e-04
Identities = 47/277 (16%), Positives = 104/277 (36%), Gaps = 11/277 (3%)
Query: 79 VKEDDCLK---SDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVS 135
+K D+ K ++ + + + SS+ + ++S S + S +++ S
Sbjct: 6 IKVDEVPKLSVKEKEIEEKSSSSSSSSSSSSSSSSSSSSSSSSSGESSSSSSSSSSSSSS 65
Query: 136 MSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIED 195
SS S ++ +++S +++ S SS + S+ ++S+ ++
Sbjct: 66 DSSDSSDSESSSSSSSSSSSSSSSSDSESSSESDSSSSGSSSSSSSSSDESSSESESEDE 125
Query: 196 TK--AHRCHKESVGHFKSEREE------GELSPNGDFEEDNFAVYANAGLEAVHKGKDCN 247
TK A E K + E E S +G + + ++ +
Sbjct: 126 TKKRARESDNEDAKETKKAKTEPESSSSSESSSSGSSSSSESESGSESDSDSSSSSSSSS 185
Query: 248 TSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREE 307
S+ + + ++ +SD S SS DS+++S + S +D + S +
Sbjct: 186 DSESDSESDSQSSSSSSSSDSSSDSDSSSSDSSSDSDSSSSSSSSSSDSDSDSDSSSDSD 245
Query: 308 HEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSE 344
D + +S + +D++D + D S SE
Sbjct: 246 SSGSSDSSSSSDSSSDESTSSDSSDSDSDSDSGSSSE 282
>S521_RAT (Q9QY02) Putative splicing factor YT521 (RA301-binding
protein)
Length = 738
Score = 48.9 bits (115), Expect = 4e-05
Identities = 57/251 (22%), Positives = 93/251 (36%), Gaps = 42/251 (16%)
Query: 87 SDRSVPKTETLGSSTLQGNVH--------INASIPDEVSRVNKQDHSIEQLVNANVSMSS 138
S R + E++ + + ++H +++S+ + V+ + S+ + N S
Sbjct: 45 SKRKSERMESIDTKRQKPSIHSRQLISKPLSSSVSNNKRIVSTKGKSVTEYKNEEYQRSE 104
Query: 139 RVEQSNGRTNINNASGLAATLSRPGYVYREGGLDL---PSSEGADSTRPDTSTNGAIIED 195
R N R + + L+++ SR Y + L S A S PD S + D
Sbjct: 105 R----NKRLDADRKIRLSSSSSREPYKSQPEKPCLRKRDSERRAKSPTPDGSERIGLEVD 160
Query: 196 TKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNR 255
+A R + S +EEG G E G ++ Q
Sbjct: 161 RRASRSSQSS-------KEEGNSEEYGSDHET---------------GSSASSEQGNNTE 198
Query: 256 REEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHD 315
EE+ GGE D E D DD E E+ + E D EE EE EE+ +
Sbjct: 199 NEEE-----GGEEDVEEDEEVDEDGDDDEEVDEDAEEEEDEEEDEEEEDEEEEEEEEEEY 253
Query: 316 NKVESEGEAEG 326
+ E + + EG
Sbjct: 254 EQDERDQKEEG 264
Score = 40.4 bits (93), Expect = 0.014
Identities = 27/85 (31%), Positives = 42/85 (48%), Gaps = 4/85 (4%)
Query: 254 NRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHE--ED 311
+RR + + E + E GS H + S +SE G+ + E GEE E+ E ED
Sbjct: 160 DRRASRSSQSSKEEGNSEEYGSDHETG--SSASSEQGNNTENEEEGGEEDVEEDEEVDED 217
Query: 312 GDHDNKVESEGEAEGMADANDVEGD 336
GD D +V+ + E E + ++ E D
Sbjct: 218 GDDDEEVDEDAEEEEDEEEDEEEED 242
>DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin
matrix protein-3) (DMP-3) [Contains: Dentin
phosphoprotein (Dentin phosphophoryn) (DPP); Dentin
sialoprotein (DSP)]
Length = 934
Score = 48.5 bits (114), Expect = 5e-05
Identities = 70/316 (22%), Positives = 129/316 (40%), Gaps = 43/316 (13%)
Query: 42 IAKGDSSPGV-GATVMSPKNT-------FEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPK 93
+ +GDS+ +SPK+T QS +C ++ G ++ + K ++S+
Sbjct: 320 VEEGDSTQATQDKEKLSPKDTRDAEGGIISQSEACPSGKSQDQG-IETEGPNKGNKSIIT 378
Query: 94 TETLGSSTLQGNVHINASIPDEVSRVN--KQDHSIEQLVNANVSMSSRVEQSNGRTNINN 151
E S L G+ N E+ + N KQ S + A E+S +N+ +
Sbjct: 379 KE---SGKLSGSKDSNGHQGVELDKRNSPKQGESDKPQGTA--------EKSAAHSNLGH 427
Query: 152 ASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKS 211
S + ++ + G+ E D S +G D D S NG+ DT + ++
Sbjct: 428 -SRIGSSSNSDGHDSYE--FDDESMQGDDPKSSDES-NGSDESDTNSESANESG------ 477
Query: 212 EREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHY-QNRREEQICGVAGGENDD 270
R + + + ++DN + H G+D ++ + G + E+ D
Sbjct: 478 SRGDASYTSDESSDDDNDS--------DSHAGEDDSSDDSSGDGDSDSNGDGDSESEDKD 529
Query: 271 ESDGSPHRSSDDSENASENGDVS--GTESADGEECSREEHEEDGDHDNKVESEGEAEGMA 328
ESD S H +S DSE+ S++ D S ++S+D + S D + ++ +
Sbjct: 530 ESDSSDHDNSSDSESKSDSSDSSDDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNSSS 589
Query: 329 DANDVEGDGASLPYSE 344
D++D G S S+
Sbjct: 590 DSSDSSGSSDSSDSSD 605
Score = 45.1 bits (105), Expect = 6e-04
Identities = 61/328 (18%), Positives = 126/328 (37%), Gaps = 16/328 (4%)
Query: 23 DTEDAVKAKKDSAKTGTA-SIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKE 81
D+ D+ + S + ++ S DSS ++ S + S+SC + +
Sbjct: 608 DSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSSSDSSDSSSCSDSSDSSDSSDSS 667
Query: 82 DDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVE 141
D SD S + + +S+ + ++S D + D S + + S+ +
Sbjct: 668 DSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSSDSSGSSDSSDSSASSD 727
Query: 142 QSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADST----RPDTSTNGAIIEDTK 197
S+ + +++S ++ S + SS +DS+ D+S + + +
Sbjct: 728 SSSSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDSSDSSDSSDSSDSSD 787
Query: 198 AHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRRE 257
+ S S+ + S N D+ ++ D + S + +
Sbjct: 788 SSDSSNSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 847
Query: 258 EQICGVAGGEND-----DESDGSPHRSSDDSENASENGDV----SGTESADGEECSREEH 308
+ +D D SD S S DS N+S++ D S ++S+DG+ S +
Sbjct: 848 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSDSKDSSSDSSDGDSKSGNGN 907
Query: 309 EEDGDHDNKVESEGEAEGMADANDVEGD 336
D + D+ +S+ ++EG +D+N D
Sbjct: 908 -SDSNSDSNSDSDSDSEG-SDSNHSTSD 933
Score = 41.6 bits (96), Expect = 0.006
Identities = 51/310 (16%), Positives = 119/310 (37%), Gaps = 18/310 (5%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ DS+ + +S + DSS ++ S + S+ + + D
Sbjct: 547 DSSDSSDDSSDSSDSSDSSDSS-DSSDSSDSSDSSDSSDSNSSSDSSDSSGSSDSSDSSD 605
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
C SD S + S + + ++S + S + S + ++ S SS
Sbjct: 606 TCDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSSSDSSDSSSCSDSSDSSDSSD 665
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ ++ +++S ++ S + SS+ +DS+ S++ + D+
Sbjct: 666 SSDSSDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSSDSSGSSDSSDSSAS 725
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
+S S + + S + D + + + ++ E+ ++S +
Sbjct: 726 SDS----SSSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDS-------- 773
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
++ D SD S SSD S++++ + ++S+D + S D +
Sbjct: 774 ---SDSSDSSDSSD--SSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSS 828
Query: 323 EAEGMADAND 332
++ +D++D
Sbjct: 829 DSSDSSDSSD 838
Score = 39.7 bits (91), Expect = 0.024
Identities = 58/292 (19%), Positives = 99/292 (33%), Gaps = 22/292 (7%)
Query: 45 GDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQG 104
G P V NT E+ N + + +GV E+ RS ++
Sbjct: 93 GKDFPTQPILVNEQGNTAEEHNDIETYGHDGVHARGENSTANGIRSQV-------GIVEN 145
Query: 105 NVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGY 164
+S+ + + K + + N + ++ E +I N++ A + G
Sbjct: 146 AEEAESSVHGQAGQNTKSGGASDVSQNGDATLVQ--ENEPPEASIKNSTNHEAGIHGSGV 203
Query: 165 VYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREE-----GELS 219
E P EG S T +I ED G+ E E+ GE +
Sbjct: 204 ATHE---TTPQREGLGSENQGTEVTPSIGEDAGLDDTDGSPSGNGVEEDEDTGSGDGEGA 260
Query: 220 PNGDFEEDNFAVYANAGLE----AVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGS 275
GD E + G H+G+ +++ ++ +E G+N E +G
Sbjct: 261 EAGDGRESHDGTKGQGGQSHGGNTDHRGQSSVSTEDDDSKEQEGFPNGHNGDNSSEENGV 320
Query: 276 PHRSSDDSENASENGDVSGTESADGEECSREEHEEDG-DHDNKVESEGEAEG 326
S + E T A+G S+ E G D +E+EG +G
Sbjct: 321 EEGDSTQATQDKEKLSPKDTRDAEGGIISQSEACPSGKSQDQGIETEGPNKG 372
Score = 38.5 bits (88), Expect = 0.054
Identities = 53/310 (17%), Positives = 118/310 (37%), Gaps = 40/310 (12%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D+ D+ + S+ ++S DSS ++ S ++ + S+S ++G D
Sbjct: 665 DSSDSSDSSDSSSSDSSSSSNSSDSSDSSDSS--SSSDSSDSSDSSDSSDSSGSSD-SSD 721
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD S + SS+ + ++S + S + +S + +++ S SS
Sbjct: 722 SSASSDSSSSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDSSDSSDSSD 781
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ ++ +++S + SS+ +DS+ S+N + D+
Sbjct: 782 SSDSSDSSDSSNSS-----------------DSSDSSDSSDSSDSSNSSDSSDSSDSSDS 824
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
+S + + S + D + + + ++ ++ ++S
Sbjct: 825 SDS----SDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD------------ 868
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
++ D SD S S DS++ + D S +S G + D + D+ +SEG
Sbjct: 869 --SSDSSDSSDSSNSSDSSDSDSKDSSSDSSDGDSKSGN--GNSDSNSDSNSDSDSDSEG 924
Query: 323 EAEGMADAND 332
+ ++D
Sbjct: 925 SDSNHSTSDD 934
Score = 35.8 bits (81), Expect = 0.35
Identities = 46/310 (14%), Positives = 110/310 (34%), Gaps = 23/310 (7%)
Query: 23 DTEDAVKAKKDSAKTGTASIAKGDSSPGVGATVMSPKNTFEQSNSCKEWQTNGVGGVKED 82
D ++K DS+ + S DSS ++ S + S+ + ++
Sbjct: 537 DNSSDSESKSDSSDSSDDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNSSSDSSDSSG 596
Query: 83 DCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQDHSIEQLVNANVSMSSRVEQ 142
SD S + S + + ++S + S + S + +++ S SS
Sbjct: 597 SSDSSDSSDTCDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSSSSDSSDSSSCSD 656
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAHRCH 202
S+ ++ +++S SS+ +DS+ D+S++ + + +
Sbjct: 657 SSDSSDSSDSSD--------------------SSDSSDSSSSDSSSSSNSSDSSDSSDSS 696
Query: 203 KESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICG 262
S S+ + S D+ A ++ + D ++S + +
Sbjct: 697 SSSDSSDSSDSSDSSDSSGSSDSSDSSA---SSDSSSSSDSSDSSSSSDSSDSSDSSDSS 753
Query: 263 VAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEG 322
+ +D + SSD S+++ + ++S+D S D + +
Sbjct: 754 DSSESSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSS 813
Query: 323 EAEGMADAND 332
++ +D++D
Sbjct: 814 DSSDSSDSSD 823
Score = 35.0 bits (79), Expect = 0.60
Identities = 47/269 (17%), Positives = 97/269 (35%), Gaps = 24/269 (8%)
Query: 63 EQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVNKQ 122
++ NS K+ +++ G E S+ + +GSS+ + H + DE + +
Sbjct: 399 DKRNSPKQGESDKPQGTAEKSAAHSNLGHSR---IGSSS-NSDGHDSYEFDDESMQGDDP 454
Query: 123 DHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADST 182
S E S+ ++S+ + N SG S + D S G D +
Sbjct: 455 KSSDE---------SNGSDESDTNSESANESGSRGDASYTSDESSDDDNDSDSHAGEDDS 505
Query: 183 RPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHK 242
D+S +G +S G SE E+ + S + D + + + + ++
Sbjct: 506 SDDSSGDG-----------DSDSNGDGDSESEDKDESDSSDHDNSSDSESKSDSSDSSDD 554
Query: 243 GKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEE 302
D + S + + + +D S SSD S ++ + +S+D +
Sbjct: 555 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNSSSDSSDSSGSSDSSDSSDTCDSSDSSD 614
Query: 303 CSREEHEEDGDHDNKVESEGEAEGMADAN 331
S D + ++ +D++
Sbjct: 615 SSDSSDSSDSSDSSDSSDSSDSSDSSDSS 643
Score = 33.1 bits (74), Expect = 2.3
Identities = 23/85 (27%), Positives = 35/85 (41%), Gaps = 17/85 (20%)
Query: 270 DESDGSPHRSSDDSENASENGDVSGTESADGEEC-------SREEHEEDGDH-------- 314
D++DGSP + + + + +GD G E+ DG E + H + DH
Sbjct: 235 DDTDGSPSGNGVEEDEDTGSGDGEGAEAGDGRESHDGTKGQGGQSHGGNTDHRGQSSVST 294
Query: 315 --DNKVESEGEAEGMADANDVEGDG 337
D+ E EG G N E +G
Sbjct: 295 EDDDSKEQEGFPNGHNGDNSSEENG 319
>UBF1_RAT (P25977) Nucleolar transcription factor 1 (Upstream
binding factor 1) (UBF-1)
Length = 764
Score = 48.1 bits (113), Expect = 7e-05
Identities = 24/72 (33%), Positives = 40/72 (55%), Gaps = 6/72 (8%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEG 326
E DDE+ S D SE++SE+ ES DG+E ++ +ED D D+ + + E+EG
Sbjct: 694 EEDDENGDSSEDGGDSSESSSED------ESEDGDENEDDDDDEDDDEDDDEDEDNESEG 747
Query: 327 MADANDVEGDGA 338
+ ++ GD +
Sbjct: 748 SSSSSSSSGDSS 759
Score = 42.0 bits (97), Expect = 0.005
Identities = 26/76 (34%), Positives = 37/76 (48%), Gaps = 6/76 (7%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDH---DNKVESEGE 323
E+DDE D ++ E ENGD S + D E S E+ EDGD D+ E + E
Sbjct: 679 EDDDEEDDDD--DDEEEEEDDENGD-SSEDGGDSSESSSEDESEDGDENEDDDDDEDDDE 735
Query: 324 AEGMADANDVEGDGAS 339
+ + N+ EG +S
Sbjct: 736 DDDEDEDNESEGSSSS 751
>UBF2_XENLA (P25980) Nucleolar transcription factor 2 (Upstream
binding factor-2) (UBF-2) (xUBF-2)
Length = 701
Score = 47.4 bits (111), Expect = 1e-04
Identities = 23/76 (30%), Positives = 41/76 (53%), Gaps = 4/76 (5%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGT----ESADGEECSREEHEEDGDHDNKVESEG 322
E+D++ + +D E++SE+GD S + +S +GEE EE EED D DN+ + +
Sbjct: 621 EDDEDEEDDDDDDDEDKEDSSEDGDSSDSSSDEDSEEGEENEDEEDEEDDDEDNEEDDDD 680
Query: 323 EAEGMADANDVEGDGA 338
G + ++ D +
Sbjct: 681 NESGSSSSSSSSADSS 696
Score = 35.0 bits (79), Expect = 0.60
Identities = 22/99 (22%), Positives = 40/99 (40%)
Query: 241 HKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADG 300
+K K Q ++ + I + + DDE D DD + E+ G S
Sbjct: 591 NKRKSTTKIQAPSSKSKLVIQSKSDDDEDDEDDEDEEDDDDDDDEDKEDSSEDGDSSDSS 650
Query: 301 EECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
+ EE EE+ D +++ + + + E D N+ +S
Sbjct: 651 SDEDSEEGEENEDEEDEEDDDEDNEEDDDDNESGSSSSS 689
>DSPP_RAT (Q62598) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 687
Score = 47.4 bits (111), Expect = 1e-04
Identities = 58/290 (20%), Positives = 106/290 (36%), Gaps = 39/290 (13%)
Query: 60 NTFEQSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRV 119
+T + + +E NG GG D S E G+ T Q N +++ + +S+
Sbjct: 292 STEDDDSKEQEGSPNGRGG----DNTSSSEETGIEEGDGTQTTQDNQNLSPTEGGIISQA 347
Query: 120 NKQDHSIEQLVNANVSMSSRVEQSNGRTNINNASG-LAATLSRPGYVYREGGLDLPSSEG 178
Q N + + + +++I SG L+ + G+ G++L
Sbjct: 348 EACPSGQSQ----NQGLETEGSSTGNKSSITKESGKLSGSKDSNGH----HGMELDKRNS 399
Query: 179 ADSTRPDTSTNGAIIEDTKAHRCHKE------SVGH----FKSEREEGELSPNGDFEEDN 228
D A DT + H S GH F E +G+ PN E +
Sbjct: 400 PKQGESDKPQGAAEKSDTHNNMGHSRIGSSSNSDGHDSYDFDDESMQGD-DPNSSDESNG 458
Query: 229 FAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASE 288
D N+ +N + +D+ SD H DDS + +
Sbjct: 459 S-----------DGSDDANSESAIENGNHGDASYTSDESSDNGSDSDSHAGEDDSSDDTS 507
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGA 338
+ D S + D E ++ ++ +HDN + ++E +D++D + D +
Sbjct: 508 DTDDSDSNGDDDSESKDKDESDNSNHDN----DSDSESKSDSSDSDSDSS 553
Score = 43.5 bits (101), Expect = 0.002
Identities = 59/278 (21%), Positives = 105/278 (37%), Gaps = 34/278 (12%)
Query: 81 EDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEV---SRVNKQDHS---IEQLVNANV 134
E D ++ VP G++ Q +HIN + D+ S + +Q HS E+ N +
Sbjct: 27 ERDIVEKSADVPFLAHPGTAA-QNELHINNATNDDSPKGSELGRQVHSNGGYERDRNGSE 85
Query: 135 SMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGL--------------DLPSSEGAD 180
S++ + S + + NA G +A Y G+ + +E A+
Sbjct: 86 SIAVGGKSSPTQPILANAQGNSAKEREDVETYGHDGIHAGGENSTANGIRGQVGIAENAE 145
Query: 181 STRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAV 240
+ ++ +G +DTK S + +E E G N V + A
Sbjct: 146 EAK-ESKVHGQPHQDTKTGLASDTSQNGDATLVQENEPQVAGSKNSTNHEVGTHGSGVAA 204
Query: 241 HKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADG 300
+ + +N+ E + G D ++GSP + + + + +GD G ++ DG
Sbjct: 205 QETTPQREGEGSENQGAEVTPSIGEGAGLDNTEGSPSGNGIEEDEDTGSGDGVGADAGDG 264
Query: 301 EECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGA 338
E HD EG++ G ND G G+
Sbjct: 265 RE----------SHDGTEGHEGQSSG--GNNDNRGQGS 290
Score = 40.8 bits (94), Expect = 0.011
Identities = 35/180 (19%), Positives = 70/180 (38%), Gaps = 10/180 (5%)
Query: 168 EGGLDLPSSEGADSTRPDTS------TNGAIIEDTKAHRCHKESVGHFKSEREEGELSPN 221
+ G D S G D + DTS +NG +D+++ + + ++ + S +
Sbjct: 488 DNGSDSDSHAGEDDSSDDTSDTDDSDSNGD--DDSESKDKDESDNSNHDNDSDSESKSDS 545
Query: 222 GDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGEND--DESDGSPHRS 279
D + D+ ++ + D + S + + + +D D SD S S
Sbjct: 546 SDSDSDSSDSSDSSDSSDSSETSDSSDSSDTSDSSDSSDSSDSSNSSDTSDSSDSSDGDS 605
Query: 280 SDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
SD + S++ D + S+D + + D+ +S+ +D N G+G S
Sbjct: 606 SDGDSSDSDSSDSDSSNSSDSDSSDSSDSSSSDSSDSDSDSKDSTSDSSDDNSKSGNGNS 665
Score = 40.0 bits (92), Expect = 0.019
Identities = 42/231 (18%), Positives = 87/231 (37%), Gaps = 36/231 (15%)
Query: 143 SNGRTNINNASGLAATLSRPGYVYREGGL-----DLPS----SEGADSTRPDTSTNGAII 193
S+ T I G T EGG+ PS ++G ++ T +I
Sbjct: 315 SSEETGIEEGDGTQTTQDNQNLSPTEGGIISQAEACPSGQSQNQGLETEGSSTGNKSSIT 374
Query: 194 EDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAV-----YANAGLEAVHKGKDCNT 248
+++ K+S GH E ++ G+ ++ A + N G + G N+
Sbjct: 375 KESGKLSGSKDSNGHHGMELDKRNSPKQGESDKPQGAAEKSDTHNNMGHSRI--GSSSNS 432
Query: 249 SQHYQNRREEQICGVAGGENDDESDGSP--------------------HRSSDDSENASE 288
H +++ + DES+GS + S + S+N S+
Sbjct: 433 DGHDSYDFDDESMQGDDPNSSDESNGSDGSDDANSESAIENGNHGDASYTSDESSDNGSD 492
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGAS 339
+ +G + + + ++ + +GD D++ + + E++ ND + + S
Sbjct: 493 SDSHAGEDDSSDDTSDTDDSDSNGDDDSESKDKDESDNSNHDNDSDSESKS 543
Score = 37.0 bits (84), Expect = 0.16
Identities = 49/219 (22%), Positives = 87/219 (39%), Gaps = 26/219 (11%)
Query: 131 NANVSMSSRVEQSNGRTNINNASGLAATLSRPG--YVYREGGLDLPSSEGADSTRPDTST 188
NA + S+V +G+ + + +GLA+ S+ G + +E + S+ + + T
Sbjct: 143 NAEEAKESKV---HGQPHQDTKTGLASDTSQNGDATLVQENEPQVAGSKNSTNHEVGTHG 199
Query: 189 NGAIIEDTKAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNT 248
+G ++T R E G SE + E++P+ + AGL+ N
Sbjct: 200 SGVAAQETTPQR---EGEG---SENQGAEVTPS---------IGEGAGLDNTEGSPSGNG 244
Query: 249 SQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASEN---GDVSGTESADGEECSR 305
+ ++ G G+ + DG+ S ++N G VS TE D +E
Sbjct: 245 IEEDEDTGSGDGVGADAGDGRESHDGTEGHEGQSSGGNNDNRGQGSVS-TEDDDSKEQEG 303
Query: 306 EEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSE 344
+ GD+ + E G EG D D +L +E
Sbjct: 304 SPNGRGGDNTSSSEETGIEEG--DGTQTTQDNQNLSPTE 340
Score = 36.2 bits (82), Expect = 0.27
Identities = 58/296 (19%), Positives = 107/296 (35%), Gaps = 39/296 (13%)
Query: 64 QSNSCKEWQTNGVGGVKEDDCLKSDRSVPKTETLGSSTLQGNVHINASIPDEVSRVN--K 121
Q+ +C Q+ G E + S+ K S L G+ N E+ + N K
Sbjct: 346 QAEACPSGQSQNQGLETEGSSTGNKSSITKE----SGKLSGSKDSNGHHGMELDKRNSPK 401
Query: 122 QDHSIEQLVNANVSMSSRVEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADS 181
Q S + E+S+ N+ + S + ++ + G+ + D S +G D
Sbjct: 402 QGESDKP--------QGAAEKSDTHNNMGH-SRIGSSSNSDGHDSYD--FDDESMQGDDP 450
Query: 182 TRPDTSTNGAIIEDTKAHRCHKESVGHFKSEREEGELSPNG-------------DFEEDN 228
D S NG+ D E+ H + E S NG D D
Sbjct: 451 NSSDES-NGSDGSDDANSESAIENGNHGDASYTSDESSDNGSDSDSHAGEDDSSDDTSDT 509
Query: 229 FAVYANAGLEAVHKGKDCNTSQHYQNRREEQICGVAGGENDDESDGSPHRSSDDSENASE 288
+N ++ K KD + + ++ N + E+ +S S SSD S+++
Sbjct: 510 DDSDSNGDDDSESKDKDESDNSNHDNDSDS--------ESKSDSSDSDSDSSDSSDSSDS 561
Query: 289 NGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSE 344
+ ++S+D + S D + ++ +D + +GD + S+
Sbjct: 562 SDSSETSDSSDSSDTSDSSDSSDSSDSSNSSDTSDSSDSSDGDSSDGDSSDSDSSD 617
Score = 35.4 bits (80), Expect = 0.46
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 2/69 (2%)
Query: 264 AGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGE 323
+ + D SD S SSD S++ S++ D + S D + + D D D+ +SEG
Sbjct: 621 SNSSDSDSSDSSDSSSSDSSDSDSDSKDSTSDSSDDNSKSGNGNSDSDSDSDS--DSEGS 678
Query: 324 AEGMADAND 332
+ ++D
Sbjct: 679 DSNHSTSDD 687
>UBF1_MOUSE (P25976) Nucleolar transcription factor 1 (Upstream
binding factor 1) (UBF-1)
Length = 765
Score = 47.0 bits (110), Expect = 2e-04
Identities = 24/73 (32%), Positives = 39/73 (52%), Gaps = 2/73 (2%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEG 326
E DDE+ S D SE++SE+ G E+ D ++ +E ++D D DN ESEG +
Sbjct: 695 EEDDENGDSSEDGGDSSESSSEDESEDGDENDDDDDDEDDEDDDDEDEDN--ESEGSSSS 752
Query: 327 MADANDVEGDGAS 339
+ + D G++
Sbjct: 753 SSSSGDSSDSGSN 765
Score = 43.1 bits (100), Expect = 0.002
Identities = 22/83 (26%), Positives = 39/83 (46%), Gaps = 11/83 (13%)
Query: 267 ENDDESDGSPHRSSDDSENASENGD-----------VSGTESADGEECSREEHEEDGDHD 315
E DD+ + ++ E ENGD S ES DG+E ++ +ED + D
Sbjct: 678 EEDDDEEEEDDEEEEEEEEDDENGDSSEDGGDSSESSSEDESEDGDENDDDDDDEDDEDD 737
Query: 316 NKVESEGEAEGMADANDVEGDGA 338
+ + + E+EG + ++ GD +
Sbjct: 738 DDEDEDNESEGSSSSSSSSGDSS 760
Score = 40.4 bits (93), Expect = 0.014
Identities = 23/77 (29%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADG---EECSREEHEEDGD-HDNKVESEG 322
+++ E D D+ E E D +G S DG E S E+ EDGD +D+ + E
Sbjct: 674 KSESEEDDDEEEEDDEEEEEEEEDDENGDSSEDGGDSSESSSEDESEDGDENDDDDDDED 733
Query: 323 EAEGMADANDVEGDGAS 339
+ + + D E +G+S
Sbjct: 734 DEDDDDEDEDNESEGSS 750
>SG1_MOUSE (P16014) Secretogranin I precursor (SgI) (Chromogranin B)
(CgB)
Length = 677
Score = 47.0 bits (110), Expect = 2e-04
Identities = 55/199 (27%), Positives = 82/199 (40%), Gaps = 32/199 (16%)
Query: 140 VEQSNGRTNINNASGLAATLSRPGYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDTKAH 199
VE S G+T + N + EGG S EG D +N ++ K +
Sbjct: 108 VEDSQGQTKVGNEK------------WTEGGGH--SREGVDDQESLRPSNQQASKEAKIY 153
Query: 200 RCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRREEQ 259
+E VG + E+EEG++ P G+ ED AG E H + N+R E
Sbjct: 154 HS-EERVGK-EREKEEGKIYPMGEHRED-------AGEEKKHIEDSGEKPNTFSNKRSE- 203
Query: 260 ICGVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEED-GDHD-NK 317
A + DES S + E + + + S ES GEE R+E ++ D D ++
Sbjct: 204 ----ASAKKKDESVARADAHSMELEEKTHSREQSSQES--GEETRRQEKPQELTDQDQSQ 257
Query: 318 VESEGEAEGMADANDVEGD 336
ES+ EG EG+
Sbjct: 258 EESQEGEEGEEGEEGEEGE 276
>UBF1_XENLA (P25979) Nucleolar transcription factor 1 (Upstream
binding factor-1) (UBF-1) (xUBF-1)
Length = 677
Score = 45.4 bits (106), Expect = 4e-04
Identities = 27/79 (34%), Positives = 42/79 (52%), Gaps = 9/79 (11%)
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGT----ESADGEECSREEHEED-----GDHDNK 317
++DDE + +D E++SE+GD S + +S DGEE EE EED D DN+
Sbjct: 599 DDDDEDEDDDDDEDEDKEDSSEDGDSSESSSDEDSEDGEENEDEEGEEDEDEEEDDDDNE 658
Query: 318 VESEGEAEGMADANDVEGD 336
S + AD++D + +
Sbjct: 659 SGSSSSSSSSADSSDSDSN 677
Score = 40.0 bits (92), Expect = 0.019
Identities = 21/72 (29%), Positives = 35/72 (48%), Gaps = 1/72 (1%)
Query: 268 NDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAEGM 327
+DD+ D DD ++ E+ + S +E D E S +E EDG+ + E E + +
Sbjct: 593 DDDDEDDDDDEDEDDDDDEDEDKEDS-SEDGDSSESSSDEDSEDGEENEDEEGEEDEDEE 651
Query: 328 ADANDVEGDGAS 339
D +D E +S
Sbjct: 652 EDDDDNESGSSS 663
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 77,190,676
Number of Sequences: 164201
Number of extensions: 3435895
Number of successful extensions: 12482
Number of sequences better than 10.0: 325
Number of HSP's better than 10.0 without gapping: 122
Number of HSP's successfully gapped in prelim test: 213
Number of HSP's that attempted gapping in prelim test: 9673
Number of HSP's gapped (non-prelim): 1412
length of query: 637
length of database: 59,974,054
effective HSP length: 117
effective length of query: 520
effective length of database: 40,762,537
effective search space: 21196519240
effective search space used: 21196519240
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146588.5


