

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.2 + phase: 0
(305 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
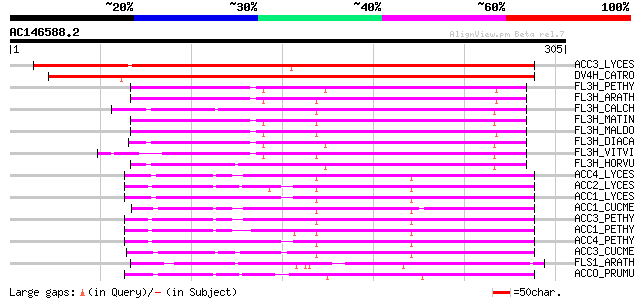
Score E
Sequences producing significant alignments: (bits) Value
ACC3_LYCES (P10967) 1-aminocyclopropane-1-carboxylate oxidase ho... 277 3e-74
DV4H_CATRO (O04847) Desacetoxyvindoline 4-hydroxylase (EC 1.14.1... 266 5e-71
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 138 2e-32
FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 134 2e-31
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 134 3e-31
FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 133 6e-31
FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 132 1e-30
FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 131 2e-30
FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 128 2e-29
FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 123 5e-28
ACC4_LYCES (P24157) 1-aminocyclopropane-1-carboxylate oxidase 4 ... 123 5e-28
ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 ... 123 6e-28
ACC1_LYCES (P05116) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 122 1e-27
ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 122 1e-27
ACC3_PETHY (Q08507) 1-aminocyclopropane-1-carboxylate oxidase 3 ... 120 5e-27
ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 120 5e-27
ACC4_PETHY (Q08508) 1-aminocyclopropane-1-carboxylate oxidase 4 ... 118 2e-26
ACC3_CUCME (P54847) 1-aminocyclopropane-1-carboxylate oxidase 3 ... 117 3e-26
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 117 5e-26
ACCO_PRUMU (Q9MB94) 1-aminocyclopropane-1-carboxylate oxidase (E... 117 5e-26
>ACC3_LYCES (P10967) 1-aminocyclopropane-1-carboxylate oxidase
homolog (Protein E8)
Length = 363
Score = 277 bits (708), Expect = 3e-74
Identities = 134/277 (48%), Positives = 188/277 (67%), Gaps = 4/277 (1%)
Query: 14 DSTYDRNDELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPIKNSMLNVPIIDLK 73
+ +YD+ ELK FDD+K GV+GL++ G+TK+P++F + K + + P+IDL+
Sbjct: 7 EESYDKMSELKAFDDTKAGVKGLVDSGITKVPQIFVLPPKDRAKKCETHFVF--PVIDLQ 64
Query: 74 DIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFY 133
I DP + E+++++R A ++WGFFQV+NHGIP VLD T+ G R+F EQD EV+KQ+Y
Sbjct: 65 GIDEDPIKHKEIVDKVRDASEKWGFFQVVNHGIPTSVLDRTLQGTRQFFEQDNEVKKQYY 124
Query: 134 NRDMKKKIVYLSTTSLYRDK--SANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVA 191
RD KK+VY S LY+ +A+WRD++ C+MAPNPP +E P + ++++SK V
Sbjct: 125 TRDTAKKVVYTSNLDLYKSSVPAASWRDTIFCYMAPNPPSLQEFPTPCGESLIDFSKDVK 184
Query: 192 TLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTI 251
LG T+ EL SE LGL SYLK+ +F +YYPPCP+PELTMG HTD F+TI
Sbjct: 185 KLGFTLLELLSEGLGLDRSYLKDYMDCFHLFCSCNYYPPCPQPELTMGTIQHTDIGFVTI 244
Query: 252 VLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
+LQ+ + GLQVL N W +V P G+LVVNIGD LQ+
Sbjct: 245 LLQDDMGGLQVLHQNHWVDVPPTPGSLVVNIGDFLQL 281
>DV4H_CATRO (O04847) Desacetoxyvindoline 4-hydroxylase (EC
1.14.11.20)
Length = 401
Score = 266 bits (680), Expect = 5e-71
Identities = 123/270 (45%), Positives = 187/270 (68%), Gaps = 3/270 (1%)
Query: 22 ELKVFDDSKLGVRGLMERGVTKIPRMFYSGEVNIIKNPI--KNSMLNVPIIDLKDIHIDP 79
ELK FD++K GV+G+++ G+TKIPR+F N+ + + S + +P+I+L + +
Sbjct: 44 ELKAFDETKAGVKGIVDTGITKIPRIFIDQPKNLDRISVCRGKSDIKIPVINLNGLSSNS 103
Query: 80 SRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPEVRKQFYNRD-MK 138
R E++ +I A +++GFFQ++NHGIP DV+D+ +DG+R+FHEQD ++++Q+Y+RD
Sbjct: 104 EIRREIVEKIGEASEKYGFFQIVNHGIPQDVMDKMVDGVRKFHEQDDQIKRQYYSRDRFN 163
Query: 139 KKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHEELPEVFRDIIMEYSKKVATLGSTIS 198
K +Y S L + NWRD++ C M N P +E P+V RDI+M+YS V LG +
Sbjct: 164 KNFLYSSNYVLIPGIACNWRDTMECIMNSNQPDPQEFPDVCRDILMKYSNYVRNLGLILF 223
Query: 199 ELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLA 258
EL SE LGL P++L+E + EG+ + GHYYP CP+PELT G S H+D F+TI++Q+Q+
Sbjct: 224 ELLSEALGLKPNHLEEMDCAEGLILLGHYYPACPQPELTFGTSKHSDSGFLTILMQDQIG 283
Query: 259 GLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
GLQ+L +NQW +V + GALV+NI DLLQ+
Sbjct: 284 GLQILLENQWIDVPFIPGALVINIADLLQL 313
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 138 bits (347), Expect = 2e-32
Identities = 76/226 (33%), Positives = 125/226 (54%), Gaps = 11/226 (4%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
+PII L+ I + +R E+ ++I AC++WG FQV++HG+ +V+ + + F P
Sbjct: 41 IPIISLEGIDDETGKRAEICDKIVKACEDWGVFQVVDHGVDAEVISQMTTFAKEFFALPP 100
Query: 127 EVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKH----EELPEVFR 180
E + +F DM KK ++ ++ L + +WR+ V F P + + PE +
Sbjct: 101 EEKLRF---DMSGGKKGGFIVSSHLQGEVVQDWREIVTYFSYPTRARDYSRWPDKPEGWI 157
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGA 240
+ +YS+K+ L + ++ SE +GL L + + ++YP CPEP+LT+G
Sbjct: 158 AVTQKYSEKLMELACKLLDVLSEAMGLEKEALTKACVDMDQKVVVNFYPKCPEPDLTLGL 217
Query: 241 STHTDPAFMTIVLQEQLAGLQVLRDN--QWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+Q+ GLQ +DN W V PV GA VVN+GD
Sbjct: 218 KRHTDPGTITLLLQDQVGGLQATKDNGKTWITVQPVEGAFVVNLGD 263
>FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Naringenin 3-dioxygenase) (Flavanone
3-hydroxylase) (FH3) (TRANSPARENT TESTA 6 protein)
Length = 358
Score = 134 bits (338), Expect = 2e-31
Identities = 79/226 (34%), Positives = 122/226 (53%), Gaps = 11/226 (4%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
+P+I L I +R E+ QI AC+ WG FQV++HG+ +++ + R F P
Sbjct: 38 IPVISLAGIDDVDGKRGEICRQIVEACENWGIFQVVDHGVDTNLVADMTRLARDFFALPP 97
Query: 127 EVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMAP----NPPKHEELPEVFR 180
E + +F DM KK ++ ++ L + +WR+ V F P + + + PE +
Sbjct: 98 EDKLRF---DMSGGKKGGFIVSSHLQGEAVQDWREIVTYFSYPVRNRDYSRWPDKPEGWV 154
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGA 240
+ EYS+++ +L + E+ SE +GL L I +YYP CP+P+LT+G
Sbjct: 155 KVTEEYSERLMSLACKLLEVLSEAMGLEKESLTNACVDMDQKIVVNYYPKCPQPDLTLGL 214
Query: 241 STHTDPAFMTIVLQEQLAGLQVLRDN--QWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+Q+ GLQ RDN W V PV GA VVN+GD
Sbjct: 215 KRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQPVEGAFVVNLGD 260
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 134 bits (337), Expect = 3e-31
Identities = 79/234 (33%), Positives = 123/234 (51%), Gaps = 9/234 (3%)
Query: 57 KNPIKNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETID 116
K P +P+I L I D RR E+ ++I AC++WG FQV++HG+ +L + +
Sbjct: 28 KVPYNTFSNEIPVISLAGI--DGCRRAEICDEIVKACEDWGIFQVVDHGVDTKLLSD-MT 84
Query: 117 GIRRFHEQDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAP----NPPKH 172
G+ R P K ++ KK ++ ++ L + +WR+ V F P + +
Sbjct: 85 GLARDFFHLPTQEKLRFDMTGGKKGGFIVSSHLQGEAVQDWREIVTYFSYPIKARDYSRW 144
Query: 173 EELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCP 232
+ P +R + EYSK + L + E+ SE +GL L + + +YYP CP
Sbjct: 145 PDKPNEWRAVTEEYSKVLMGLACKLLEVLSEAMGLEKEALTKACVDMDQKVVVNYYPKCP 204
Query: 233 EPELTMGASTHTDPAFMTIVLQEQLAGLQVLRD--NQWFNVAPVHGALVVNIGD 284
+P+LT+G HTDP +T++LQ+Q+ GLQ RD W V PV GA VVN+GD
Sbjct: 205 QPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGESWITVKPVEGAFVVNLGD 258
>FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 357
Score = 133 bits (334), Expect = 6e-31
Identities = 78/226 (34%), Positives = 120/226 (52%), Gaps = 11/226 (4%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
+P+I L I +R E+ +I AC+ WG FQV++HG+ ++ + R F P
Sbjct: 37 IPVISLAGIDDVDGKRGEICREIVEACENWGIFQVVDHGVDTSLVADMTRLARDFFALPP 96
Query: 127 EVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMAP----NPPKHEELPEVFR 180
E + +F DM KK ++ ++ L + +WR+ V F P + + + P+ +
Sbjct: 97 EEKLRF---DMSGGKKGGFIVSSHLQGEAVQDWREIVTYFSYPVRNRDYSRWPDKPQGWA 153
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGA 240
+ EYS+K+ L + E+ SE +GL L I +YYP CP+P+LT+G
Sbjct: 154 KVTEEYSEKLMGLACKLLEVLSEAMGLEKESLTNACVDMDQKIVVNYYPKCPQPDLTLGL 213
Query: 241 STHTDPAFMTIVLQEQLAGLQVLRD--NQWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+Q+ GLQ RD N W V PV GA VVN+GD
Sbjct: 214 KRHTDPGTITLLLQDQVGGLQATRDDGNTWITVQPVEGAFVVNLGD 259
>FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 132 bits (331), Expect = 1e-30
Identities = 75/226 (33%), Positives = 122/226 (53%), Gaps = 11/226 (4%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
+PII L I RR E+ +I AC++WG FQ+++HG+ +++ E R F
Sbjct: 39 IPIISLAGIDEVEGRRGEICKKIVAACEDWGIFQIVDHGVDAELISEMTGLAREFFALPS 98
Query: 127 EVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMAP----NPPKHEELPEVFR 180
E + +F DM KK ++ ++ L + +WR+ V F P + + + PE +R
Sbjct: 99 EEKLRF---DMSGGKKGGFIVSSHLQGEAVQDWREIVTYFSYPIRHRDYSRWPDKPEAWR 155
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGA 240
++ +YS ++ L + + SE +GL L + + ++YP CP+P+LT+G
Sbjct: 156 EVTKKYSDELMGLACKLLGVLSEAMGLDTEALTKACVDMDQKVVVNFYPKCPQPDLTLGL 215
Query: 241 STHTDPAFMTIVLQEQLAGLQVLRDN--QWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+Q+ GLQ RD+ W V PV GA VVN+GD
Sbjct: 216 KRHTDPGTITLLLQDQVGGLQATRDDGKTWITVQPVEGAFVVNLGD 261
>FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 365
Score = 131 bits (330), Expect = 2e-30
Identities = 80/227 (35%), Positives = 122/227 (53%), Gaps = 13/227 (5%)
Query: 66 NVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQD 125
++P+I L I D +R E+ +I AC++WG FQV++HG+ D++ + R F
Sbjct: 40 DIPVISLAGI--DGEKRGEICRKIVEACEDWGIFQVVDHGVGDDLIADMTRLAREFFALP 97
Query: 126 PEVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKH----EELPEVF 179
E + +F DM KK ++ ++ L + +WR+ V F P + + PE +
Sbjct: 98 AEEKLRF---DMSGGKKGGFIVSSHLQGEVVQDWREIVTYFSYPTNSRDYTRWPDKPEGW 154
Query: 180 RDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMG 239
+ EYS K+ TL T+ + SE +GL L + I +YYP CP+P+LT+G
Sbjct: 155 IKVTEEYSNKLMTLACTLLGVLSEAMGLELEALTKACVDMDQKIVVNYYPKCPQPDLTLG 214
Query: 240 ASTHTDPAFMTIVLQEQLAGLQVLRD--NQWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+Q+ GLQ RD W V PV GA VVN+GD
Sbjct: 215 LKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQPVPGAFVVNLGD 261
>FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 128 bits (322), Expect = 2e-29
Identities = 80/244 (32%), Positives = 125/244 (50%), Gaps = 24/244 (9%)
Query: 49 YSGEVNIIKNPIKNSMLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPI 108
+S E+ +I + K SM ++D E+ +I AC++WG FQV+NHG+
Sbjct: 34 FSNEIPVI-SLTKESMKLAAVVD------------EICRKIVEACEDWGIFQVVNHGVDS 80
Query: 109 DVLDETIDGIRRFHEQDPEVRKQFYNRDMK--KKIVYLSTTSLYRDKSANWRDSVGCFMA 166
+++ E R F PE +F DM KK ++ ++ L + +WR+ V F
Sbjct: 81 NLISEMTRLAREFFALPPEENVRF---DMSGGKKGGFIVSSHLQGEAVQDWREIVTYFSY 137
Query: 167 P----NPPKHEELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIF 222
P + + + PE +R + EYS+K+ L + E+ SE + L L
Sbjct: 138 PLRTRDYSRWPDKPEGWRSVTQEYSEKLMGLACKLLEVLSEAMDLDKDALTNACVDMDQK 197
Query: 223 IQGHYYPPCPEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRD--NQWFNVAPVHGALVV 280
+ ++YP CP+P+LT+G HTDP +T++LQ+Q+ GLQ RD W V PV GA VV
Sbjct: 198 VVVNFYPQCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQPVEGAFVV 257
Query: 281 NIGD 284
N+GD
Sbjct: 258 NLGD 261
>FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 377
Score = 123 bits (309), Expect = 5e-28
Identities = 72/224 (32%), Positives = 118/224 (52%), Gaps = 9/224 (4%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
VP+I L I D +RR ++ +++ AC++WG FQVI+HG+ D++ + R F P
Sbjct: 43 VPLISLHGI--DGARRAQIRDRVAAACEDWGIFQVIDHGVDADLIADMTRLAREFFAL-P 99
Query: 127 EVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKH----EELPEVFRDI 182
K Y+ KK ++ ++ L + +WR+ V F P + E P + +
Sbjct: 100 AEDKLRYDMSGGKKGGFIVSSHLQGEAVQDWREIVTYFSYPVKARDYGRWPEKPAGWCAV 159
Query: 183 IMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQGHYYPPCPEPELTMGAST 242
+ YS+++ L + + SE +GL L + + ++YP CP+P+LT+G
Sbjct: 160 VERYSERLMGLSCNLMGVLSEAMGLETEALAKACVDMDQKVVVNFYPRCPQPDLTLGLKR 219
Query: 243 HTDPAFMTIVLQEQLAGLQVLRD--NQWFNVAPVHGALVVNIGD 284
HTDP +T++LQ+ + GLQ RD W V P+ GA VVN+GD
Sbjct: 220 HTDPGTITLLLQDLVGGLQATRDGGKNWITVQPISGAFVVNLGD 263
>ACC4_LYCES (P24157) 1-aminocyclopropane-1-carboxylate oxidase 4 (EC
1.14.17.4) (ACC oxidase 4) (Ethylene-forming enzyme)
(EFE) (Protein pHTOM5)
Length = 316
Score = 123 bits (309), Expect = 5e-28
Identities = 71/232 (30%), Positives = 129/232 (55%), Gaps = 16/232 (6%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L++++ D R + + I+ AC+ WGFF+++NHGIP +V+D T++ + + H
Sbjct: 1 MENFPIINLENLNGD--ERAKTMEMIKDACENWGFFELVNHGIPHEVMD-TVEKLTKGH- 56
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRD-KSANWRDSVGCFMAP--NPPKHEELPEVFR 180
K+ + K+ + ++ + +W + P N + +L E +R
Sbjct: 57 -----YKKCMEQRFKELVASKGLEAVQAEVTDLDWESTFFLRHLPTSNISQVPDLDEEYR 111
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELT 237
+++ +++K++ L + +L E LGL YLK Y + YPPCP+P+L
Sbjct: 112 EVMRDFAKRLEKLAEELLDLLCENLGLEKGYLKNAFYGSKGPNFGTKVSNYPPCPKPDLI 171
Query: 238 MGASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
G HTD + ++ Q ++++GLQ+L+D QW +V P+ ++VVN+GD L+V
Sbjct: 172 KGLRAHTDAGGIILLFQDDKVSGLQLLKDEQWIDVPPMRHSIVVNLGDQLEV 223
>ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 (EC
1.14.17.4) (ACC oxidase 2) (Ethylene-forming enzyme)
(EFE) (Protein GTOMA)
Length = 316
Score = 123 bits (308), Expect = 6e-28
Identities = 74/233 (31%), Positives = 132/233 (55%), Gaps = 18/233 (7%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L+ ++ + RV + +I AC+ WGFF+++NHGIP +V+D T++ + + H
Sbjct: 1 MENFPIINLEKLN--GAERVATMEKINDACENWGFFELVNHGIPHEVMD-TVEKLTKGHY 57
Query: 124 QDPEVRKQFYNRDMKKKI--VYLSTTSLYRDKSANWRDSVGCFMAP--NPPKHEELPEVF 179
+ + ++F KK + V + T + +W + P N + +L +V+
Sbjct: 58 KKC-MEQRFKELVAKKGLEGVEVEVTDM------DWESTFFLRHLPSSNISQLPDLDDVY 110
Query: 180 RDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPEL 236
R+++ ++ K++ L + +L E LGL SYLK Y + YPPCP+P+L
Sbjct: 111 REVMRDFRKRLEKLAEELLDLLCENLGLEKSYLKNTFYGSKGPNFGTKVSNYPPCPKPDL 170
Query: 237 TMGASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
G HTD + ++ Q ++++GLQ+L+D +W +V P+ ++VVN+GD L+V
Sbjct: 171 IKGLRAHTDAGGIILLFQDDKVSGLQLLKDGRWIDVPPMRHSIVVNLGDQLEV 223
>ACC1_LYCES (P05116) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE) (Protein pTOM 13)
Length = 315
Score = 122 bits (306), Expect = 1e-27
Identities = 72/231 (31%), Positives = 123/231 (53%), Gaps = 14/231 (6%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L+ ++ D R + I+ AC+ WGFF+++NHGIP +V+D + ++
Sbjct: 1 MENFPIINLEKLNGD--ERANTMEMIKDACENWGFFELVNHGIPHEVMDTVEKMTKGHYK 58
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAP--NPPKHEELPEVFRD 181
+ E R + + V T L +W + P N + +L E +R+
Sbjct: 59 KCMEQRFKELVASKGLEAVQAEVTDL------DWESTFFLRHLPTSNISQVPDLDEEYRE 112
Query: 182 IIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELTM 238
++ +++K++ L + +L E LGL YLK Y + YPPCP+P+L
Sbjct: 113 VMRDFAKRLEKLAEELLDLLCENLGLEKGYLKNAFYGSKGPNFGTKVSNYPPCPKPDLIK 172
Query: 239 GASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
G HTD + ++ Q ++++GLQ+L+D QW +V P+ ++VVN+GD L+V
Sbjct: 173 GLRAHTDAGGIILLFQDDKVSGLQLLKDEQWIDVPPMRHSIVVNLGDQLEV 223
>ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE) (PMEL1)
Length = 318
Score = 122 bits (305), Expect = 1e-27
Identities = 74/230 (32%), Positives = 125/230 (54%), Gaps = 20/230 (8%)
Query: 68 PIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDPE 127
PII+L++I+ D R +++ QI AC+ WGFF+++NHGIP + LD ++ + R H
Sbjct: 5 PIINLENINDDG--RAKILEQIEDACQNWGFFELVNHGIPHEFLD-MVEKMTRDH----- 56
Query: 128 VRKQFYNRDMKKKIVYLSTTSLYRD-KSANWRDSVGCFMAP--NPPKHEELPEVFRDIIM 184
K+ K+ ++ + + +W + P N + +L E ++ I+
Sbjct: 57 -YKKCMEERFKETVLSKGLEAAQAEVNDMDWESTFFLRHLPESNISQMSDLDEEYKKIMK 115
Query: 185 EYSKKVATLGSTISELFSETLGLHPSYLKERNYFE-----GIFIQGHYYPPCPEPELTMG 239
E++KK+ L + +L E LGL YLK+ Y G + YPPCP+P+L G
Sbjct: 116 EFAKKLENLAEELLDLLCENLGLEKGYLKKAFYGSKGPTFGTKVSN--YPPCPKPDLIKG 173
Query: 240 ASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
HTD + ++ Q ++++GLQ+L+D W +V P+ A+VVN+GD L+V
Sbjct: 174 LRAHTDAGGIILLFQDDKVSGLQLLKDGNWIDVPPMRHAIVVNLGDQLEV 223
>ACC3_PETHY (Q08507) 1-aminocyclopropane-1-carboxylate oxidase 3 (EC
1.14.17.4) (ACC oxidase 3) (Ethylene-forming enzyme)
(EFE)
Length = 320
Score = 120 bits (300), Expect = 5e-27
Identities = 70/232 (30%), Positives = 128/232 (55%), Gaps = 16/232 (6%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L+ ++ S R + I+ AC+ WGFF+++NHGIP +V+D T++ + + H
Sbjct: 1 MENFPIINLEKLN--GSERDATMEMIKDACENWGFFELVNHGIPHEVMD-TVEKLTKGH- 56
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRD-KSANWRDSVGCFMAP--NPPKHEELPEVFR 180
K+ + K+ + ++ + +W + P N + +L + +R
Sbjct: 57 -----YKKCMEQRFKELVASKGLEAVQAEVTDLDWESTFFLRHLPVSNISEVPDLDDEYR 111
Query: 181 DIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELT 237
+++ +++K++ L + +L E LGL YLK+ Y + YPPCP+P+L
Sbjct: 112 EVMRDFAKRLEKLAEELLDLLCENLGLEKGYLKKAFYGSKGPNFGTKVSNYPPCPKPDLI 171
Query: 238 MGASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
G HTD + ++ Q ++++GLQ+L+D QW +V P+ ++VVN+GD L+V
Sbjct: 172 KGLRAHTDAGGIILLFQDDKVSGLQLLKDGQWIDVPPMRHSIVVNLGDQLEV 223
>ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 319
Score = 120 bits (300), Expect = 5e-27
Identities = 72/236 (30%), Positives = 124/236 (52%), Gaps = 24/236 (10%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII L ++ R + I+ AC+ WGFF+++NHGIP +V+D T++ + + H
Sbjct: 1 MENFPIISLDKVN--GVERAATMEMIKDACENWGFFELVNHGIPREVMD-TVEKMTKGH- 56
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSA-----NWRDSVGCFMAP--NPPKHEELP 176
Y + M+++ L + A +W + P N + +L
Sbjct: 57 ---------YKKCMEQRFKELVASKALEGVQAEVTDMDWESTFFLKHLPISNISEVPDLD 107
Query: 177 EVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPE 233
E +R+++ +++K++ L + +L E LGL YLK Y + YPPCP+
Sbjct: 108 EEYREVMRDFAKRLEKLAEELLDLLCENLGLEKGYLKNAFYGSKGPNFGTKVSNYPPCPK 167
Query: 234 PELTMGASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
P+L G HTD + ++ Q ++++GLQ+L+D QW +V P+ ++VVN+GD L+V
Sbjct: 168 PDLIKGLRAHTDAGGIILLFQDDKVSGLQLLKDGQWIDVPPMRHSIVVNLGDQLEV 223
>ACC4_PETHY (Q08508) 1-aminocyclopropane-1-carboxylate oxidase 4 (EC
1.14.17.4) (ACC oxidase 4) (Ethylene-forming enzyme)
(EFE)
Length = 319
Score = 118 bits (295), Expect = 2e-26
Identities = 69/231 (29%), Positives = 125/231 (53%), Gaps = 14/231 (6%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L+++ + R + I+ AC+ WGFF+++NHGIP +V+D + ++
Sbjct: 1 MENFPIINLENLC--GAERDATMEMIKDACENWGFFELVNHGIPHEVMDTVEKFTKGHYK 58
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAP--NPPKHEELPEVFRD 181
+ E R + + V T L +W + P N + +L + +R+
Sbjct: 59 KCMEQRFKELVASKGLEAVQAEVTDL------DWESTFFLRHLPVSNISEVPDLDDEYRE 112
Query: 182 IIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELTM 238
++ +++K++ L + +L E LGL YLK+ Y + YPPCP+P+L
Sbjct: 113 VMRDFAKRLEKLAEELLDLLCENLGLEKGYLKKAFYGSKGPNFGTKVSNYPPCPKPDLIK 172
Query: 239 GASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
G HTD + ++ Q ++++GLQ+L+D+QW +V P+ ++V+N+GD L+V
Sbjct: 173 GLRAHTDAGGIILLFQDDKVSGLQLLKDDQWIDVPPMRHSIVINLGDQLEV 223
>ACC3_CUCME (P54847) 1-aminocyclopropane-1-carboxylate oxidase 3 (EC
1.14.17.4) (ACC oxidase 3) (Ethylene-forming enzyme)
(EFE)
Length = 320
Score = 117 bits (294), Expect = 3e-26
Identities = 67/230 (29%), Positives = 129/230 (55%), Gaps = 14/230 (6%)
Query: 65 LNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQ 124
++ P+I++ +++ + RV V+NQI AC+ WGFF+++NHGI +++D+ ++ + + H +
Sbjct: 3 MDFPVINMNNLNGES--RVSVLNQINDACENWGFFELVNHGISHELMDK-VEKLTKEHYR 59
Query: 125 DPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAP--NPPKHEELPEVFRDI 182
+ +Q + + K + T + +W + P N + +L E ++ +
Sbjct: 60 --KCMEQRFKEMVASKGLDSVETEI---NDTDWESTFFLRHLPVSNMSEIGDLDEEYKKV 114
Query: 183 IMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFE---GIFIQGHYYPPCPEPELTMG 239
+ E++ ++ L + +L E LGL YLK+ Y + YPPCP+PEL G
Sbjct: 115 MKEFADELEKLAEEVLDLLCENLGLEKGYLKKVFYGSKGPNFGTKVSNYPPCPKPELIKG 174
Query: 240 ASTHTDPAFMTIVLQ-EQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
HTD + ++ Q ++++GL VL+D +W +V P+H ++V+N+GD L+V
Sbjct: 175 LRAHTDAGGLILLFQDDKVSGLHVLKDGKWVDVPPMHHSIVINLGDQLEV 224
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 117 bits (292), Expect = 5e-26
Identities = 78/243 (32%), Positives = 121/243 (49%), Gaps = 29/243 (11%)
Query: 67 VPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHEQDP 126
+P++DL D + RR V A +EWG FQV+NHGIP +++ D R+F E P
Sbjct: 43 IPVVDLSDPDEESVRRAVV-----KASEEWGLFQVVNHGIPTELIRRLQDVGRKFFEL-P 96
Query: 127 EVRKQFYNRDMKKKIVYLSTTSLYRDKSAN--WRDSV-------GC----FMAPNPPKHE 173
K+ + K + T L +D W D + C F NPP++
Sbjct: 97 SSEKESVAKPEDSKDIEGYGTKLQKDPEGKKAWVDHLFHRIWPPSCVNYRFWPKNPPEYR 156
Query: 174 ELPEVFRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKER--NYFEGIFIQGHYYPPC 231
E+ E EY+ V L T+ + S+ LGL LKE ++ +YYPPC
Sbjct: 157 EVNE-------EYAVHVKKLSETLLGILSDGLGLKRDALKEGLGGEMAEYMMKINYYPPC 209
Query: 232 PEPELTMGASTHTDPAFMTIVLQEQLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQVNLF 291
P P+L +G HTD + +T+++ ++ GLQV +D+ WF+ + A++V+IGD + + L
Sbjct: 210 PRPDLALGVPAHTDLSGITLLVPNEVPGLQVFKDDHWFDAEYIPSAVIVHIGDQI-LRLS 268
Query: 292 KGR 294
GR
Sbjct: 269 NGR 271
>ACCO_PRUMU (Q9MB94) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 319
Score = 117 bits (292), Expect = 5e-26
Identities = 73/234 (31%), Positives = 126/234 (53%), Gaps = 20/234 (8%)
Query: 64 MLNVPIIDLKDIHIDPSRRVEVINQIRTACKEWGFFQVINHGIPIDVLDETIDGIRRFHE 123
M N PII+L+ ++ + R + +I+ AC+ WGFF++++HGIP + LD T++ + + H
Sbjct: 1 MENFPIINLEGLNGEG--RKATMEKIKDACENWGFFELVSHGIPTEFLD-TVERLTKEHY 57
Query: 124 QDPEVRKQFYNRDMKKKIVYLSTTSLYRDKSANWRDSVGCFMAPNPPKHE-----ELPEV 178
+ + ++F K + + T N D F + PK +L +
Sbjct: 58 RQC-LEQRFKELVASKGLEAVKT-------EVNDMDWESTFYLRHLPKSNISEVPDLEDQ 109
Query: 179 FRDIIMEYSKKVATLGSTISELFSETLGLHPSYLKERNYFEGIFIQG---HYYPPCPEPE 235
+R+++ E++ K+ L + +L E LGL YLK+ Y G YPPCP PE
Sbjct: 110 YRNVMKEFALKLEKLAEQLLDLLCENLGLEKGYLKKAFYGTNGPTFGTKVSNYPPCPNPE 169
Query: 236 LTMGASTHTDPAFMTIVLQE-QLAGLQVLRDNQWFNVAPVHGALVVNIGDLLQV 288
L G HTD + ++ Q+ +++GLQ+L+D QW +V P+ ++V+N+GD L+V
Sbjct: 170 LIKGLRAHTDAGGLILLFQDDKVSGLQLLKDGQWIDVPPMRHSIVINLGDQLEV 223
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,996,481
Number of Sequences: 164201
Number of extensions: 1645962
Number of successful extensions: 4216
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 4000
Number of HSP's gapped (non-prelim): 88
length of query: 305
length of database: 59,974,054
effective HSP length: 110
effective length of query: 195
effective length of database: 41,911,944
effective search space: 8172829080
effective search space used: 8172829080
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146588.2


