

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
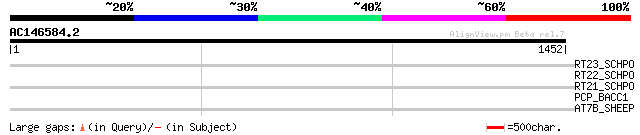
Query= AC146584.2 - phase: 0 /pseudo
(1452 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 37 0.51
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 37 0.51
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 37 0.51
PCP_BACC1 (Q735N6) Pyrrolidone-carboxylate peptidase (EC 3.4.19.... 33 4.4
AT7B_SHEEP (Q9XT50) Copper-transporting ATPase 2 (EC 3.6.3.4) (C... 32 9.7
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 36.6 bits (83), Expect = 0.51
Identities = 16/42 (38%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Query: 1330 PKFIGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 1370
P F GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1216 PSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 36.6 bits (83), Expect = 0.51
Identities = 16/42 (38%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Query: 1330 PKFIGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 1370
P F GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1216 PSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 36.6 bits (83), Expect = 0.51
Identities = 16/42 (38%), Positives = 25/42 (59%), Gaps = 1/42 (2%)
Query: 1330 PKFIGPYQILERVGTVAYRVGLPPHLSNL-HNVFHVSQLRKY 1370
P F GP+ +L++ G Y + LP + ++ + FHVS L KY
Sbjct: 1216 PSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKY 1257
>PCP_BACC1 (Q735N6) Pyrrolidone-carboxylate peptidase (EC 3.4.19.3)
(5-oxoprolyl-peptidase) (Pyroglutamyl-peptidase I)
(PGP-I) (Pyrase)
Length = 215
Score = 33.5 bits (75), Expect = 4.4
Identities = 29/117 (24%), Positives = 51/117 (42%), Gaps = 20/117 (17%)
Query: 1311 FESHSYDRCRTCFEV-KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFH--VSQL 1367
F+ + +EV K + K IG Y+I+ + + VFH +S L
Sbjct: 9 FDPFGGENINPAWEVAKNLHEKTIGEYKIISK---------------QVPTVFHKSISVL 53
Query: 1368 RKYVPD--PSHVIQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGAT 1422
++Y+ + P +I R ++T+E + + IDD ++ G + V VV G T
Sbjct: 54 KEYIEELAPEFIICIGQAGGRPDITIERVAINIDDARIADNEGNQPVDVPVVEEGPT 110
>AT7B_SHEEP (Q9XT50) Copper-transporting ATPase 2 (EC 3.6.3.4) (Copper
pump 2) (Wilson disease-associated protein homolog)
Length = 1505
Score = 32.3 bits (72), Expect = 9.7
Identities = 11/42 (26%), Positives = 25/42 (59%)
Query: 1329 DPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKY 1370
DP+ IGP I++ + + +R L + N H++ H +++++
Sbjct: 650 DPEIIGPRDIVKLIEEIGFRASLAQRIPNAHHLDHKVEIKQW 691
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.365 0.164 0.591
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 127,779,670
Number of Sequences: 164201
Number of extensions: 4318310
Number of successful extensions: 21503
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 21498
Number of HSP's gapped (non-prelim): 6
length of query: 1452
length of database: 59,974,054
effective HSP length: 123
effective length of query: 1329
effective length of database: 39,777,331
effective search space: 52864072899
effective search space used: 52864072899
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 72 (32.3 bits)
Medicago: description of AC146584.2


