

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.4 - phase: 0
(165 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
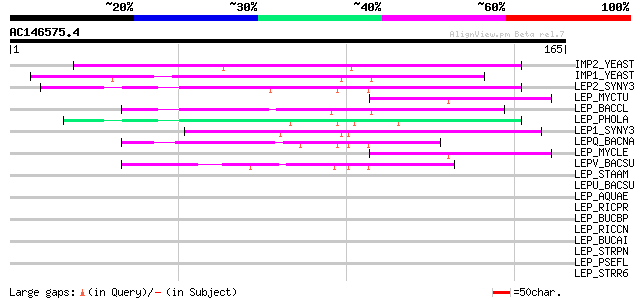
Score E
Sequences producing significant alignments: (bits) Value
IMP2_YEAST (P46972) Mitochondrial inner membrane protease subuni... 99 6e-21
IMP1_YEAST (P28627) Mitochondrial inner membrane protease subuni... 69 4e-12
LEP2_SYNY3 (P73157) Probable signal peptidase I-2 (EC 3.4.21.89)... 58 1e-08
LEP_MYCTU (Q10789) Probable signal peptidase I (EC 3.4.21.89) (S... 50 3e-06
LEP_BACCL (P41027) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 50 3e-06
LEP_PHOLA (Q51876) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 49 5e-06
LEP1_SYNY3 (P72660) Probable signal peptidase I-1 (EC 3.4.21.89)... 45 6e-05
LEPQ_BACNA (Q57350) Signal peptidase I P (EC 3.4.21.89) (SPase I... 44 2e-04
LEP_MYCLE (O33021) Probable signal peptidase I (EC 3.4.21.89) (S... 42 5e-04
LEPV_BACSU (O07560) Signal peptidase I V (EC 3.4.21.89) (SPase I... 42 5e-04
LEP_STAAM (P72365) Signal peptidase IB (EC 3.4.21.89) (SPase IB)... 41 0.001
LEPU_BACSU (P42959) Signal peptidase I U (EC 3.4.21.89) (SPase I... 38 0.009
LEP_AQUAE (O67088) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 37 0.020
LEP_RICPR (Q9ZE32) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 37 0.027
LEP_BUCBP (Q89AM6) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 36 0.035
LEP_RICCN (Q92JB1) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 34 0.13
LEP_BUCAI (P57347) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 34 0.13
LEP_STRPN (O07344) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 32 0.50
LEP_PSEFL (P26844) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 32 0.65
LEP_STRR6 (P59662) Signal peptidase I (EC 3.4.21.89) (SPase I) (... 32 0.85
>IMP2_YEAST (P46972) Mitochondrial inner membrane protease subunit 2
(EC 3.4.99.-)
Length = 177
Score = 98.6 bits (244), Expect = 6e-21
Identities = 47/136 (34%), Positives = 78/136 (56%), Gaps = 3/136 (2%)
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCL-DKFKFSHGDIVIFSSP 78
+ T+++ + V+G SM PT NP+T + D+V + K + + S DI++F +P
Sbjct: 23 VLLTINNNVVHIAQVKGTSMQPTLNPQTETLATDWVLLWKFGVKNPSNLSRDDIILFKAP 82
Query: 79 SNFKETHIKRIIALPGEWFVNR--HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGL 136
+N ++ + KR+ LP + + + + + +P GH WVEGDN S DS ++GP+ GL
Sbjct: 83 TNPRKVYCKRVKGLPFDTIDTKFPYPKPQVNLPRGHIWVEGDNYFHSIDSNTFGPISSGL 142
Query: 137 VRGRVTHVVWPPQRIG 152
V G+ +VWPP R G
Sbjct: 143 VIGKAITIVWPPSRWG 158
>IMP1_YEAST (P28627) Mitochondrial inner membrane protease subunit 1
(EC 3.4.99.-)
Length = 189
Score = 69.3 bits (168), Expect = 4e-12
Identities = 48/152 (31%), Positives = 70/152 (45%), Gaps = 22/152 (14%)
Query: 7 VWNVTKKLATIGLITFTVSDRYA-TVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKF 65
+W+ T A L + YA RG SM PT S T+DYV V K +
Sbjct: 7 IWSKTFSYAIRSLCFLHIIHMYAYEFTETRGESMLPTL-----SATNDYVHVLKNFQNGR 61
Query: 66 KFSHGDIVIFSSPSNFKETHIKRIIALPGEWF------VNRHNQDVL----------KVP 109
GD ++ P++ KR+ +PG+ + + DVL KVP
Sbjct: 62 GIKMGDCIVALKPTDPNHRICKRVTGMPGDLVLVDPSTIVNYVGDVLVDEERFGTYIKVP 121
Query: 110 EGHCWVEGDNAASSTDSKSYGPVPLGLVRGRV 141
EGH WV GDN + S DS++Y +P+GL+ G++
Sbjct: 122 EGHVWVTGDNLSHSLDSRTYNALPMGLIMGKI 153
>LEP2_SYNY3 (P73157) Probable signal peptidase I-2 (EC 3.4.21.89)
(SPase I-2) (Leader peptidase I-2)
Length = 218
Score = 57.8 bits (138), Expect = 1e-08
Identities = 51/169 (30%), Positives = 71/169 (41%), Gaps = 37/169 (21%)
Query: 10 VTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSH 69
VT + IG+ TF RY + +SM PT +D + +EK+
Sbjct: 29 VTAVILAIGIRTFVAEARY-----IPSSSMEPTLQ------INDRLIIEKISYRLRDPER 77
Query: 70 GDIVIFS-----SPSNFKETHIKRIIALPGEW--------FVNRHNQDV----------- 105
G+IV+F+ NF + IKRII LPG+ +VN D
Sbjct: 78 GEIVVFNPTDALKAKNFHDAFIKRIIGLPGDEVRVSQGNVYVNGKMLDENYIAAPPAYEY 137
Query: 106 --LKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
+KVP+ V GDN +S DS +G VP + GR WP R+G
Sbjct: 138 GPVKVPDDQYLVLGDNRNNSYDSHYWGFVPREKLLGRAFVRFWPVPRVG 186
>LEP_MYCTU (Q10789) Probable signal peptidase I (EC 3.4.21.89)
(SPase I) (Leader peptidase I)
Length = 294
Score = 49.7 bits (117), Expect = 3e-06
Identities = 24/65 (36%), Positives = 34/65 (51%), Gaps = 11/65 (16%)
Query: 108 VPEGHCWVEGDNAASSTDSKSY-----------GPVPLGLVRGRVTHVVWPPQRIGAVKN 156
VP G WV GDN S DS+++ G VP+ V G+ +VWPP R G V++
Sbjct: 228 VPPGRVWVMGDNRTHSADSRAHCPLLCTDDPLPGTVPVANVIGKARLIVWPPSRWGVVRS 287
Query: 157 TTPER 161
P++
Sbjct: 288 VNPQQ 292
>LEP_BACCL (P41027) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 182
Score = 49.7 bits (117), Expect = 3e-06
Identities = 42/149 (28%), Positives = 62/149 (41%), Gaps = 43/149 (28%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALP 93
V G SM PT + + + V KL D DI++F + N KE ++KR+I LP
Sbjct: 34 VEGKSMMPTLE------SGNLLIVNKLSYDIGPIRRFDIIVFHA--NKKEDYVKRVIGLP 85
Query: 94 G------------------EWFVNRHNQDVL-----------------KVPEGHCWVEGD 118
G E ++ + Q +L +VP G +V GD
Sbjct: 86 GDRIAYKNDILYVNGKKVDEPYLRPYKQKLLDGRLTGDFTLEEVTGKTRVPPGCIFVLGD 145
Query: 119 NAASSTDSKSYGPVPLGLVRGRVTHVVWP 147
N SS DS+ +G V + + G+V WP
Sbjct: 146 NRLSSWDSRHFGFVKINQIVGKVDFRYWP 174
>LEP_PHOLA (Q51876) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 203
Score = 48.9 bits (115), Expect = 5e-06
Identities = 46/164 (28%), Positives = 65/164 (39%), Gaps = 39/164 (23%)
Query: 17 IGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFS 76
+G+ TF RY + SM PT +D + VEK+ GDI++F
Sbjct: 43 LGIRTFVAEARY-----IPSESMLPTLE------VNDRLIVEKISYHFNPPRRGDIIVFH 91
Query: 77 SPSNFK-------ETHIKRIIALPGEW--------FVNRH--NQDVLKVPEGHCW----- 114
K E IKR+I LPGE +N ++ ++ P + W
Sbjct: 92 PTEALKQQNPSLNEAFIKRVIGLPGETVQVTGGRVLINGQPLEENYIQSPPDYQWGPEKV 151
Query: 115 ------VEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
V GDN +S DS +G VP + GR WP R+G
Sbjct: 152 PADSFLVLGDNRNNSYDSHFWGYVPRQNIIGRAVVRFWPVNRLG 195
>LEP1_SYNY3 (P72660) Probable signal peptidase I-1 (EC 3.4.21.89)
(SPase I-1) (Leader peptidase I-1)
Length = 196
Score = 45.4 bits (106), Expect = 6e-05
Identities = 37/134 (27%), Positives = 58/134 (42%), Gaps = 28/134 (20%)
Query: 53 DYVFVEKLCLDKFKFSHGDIVIFSSPS-------NFKETHIKRIIALPGEWF-VN----- 99
D + VEK+ GDI++F P + + IKR+IALPG+ VN
Sbjct: 53 DRLVVEKVSYHFHPPQVGDIIVFHPPELLQVQGYDLGQAFIKRVIALPGQTVEVNNGIVY 112
Query: 100 ---------------RHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHV 144
++N ++VP+G +V GDN +S DS +G +P + G
Sbjct: 113 RDGQPLQEEYILEPPQYNLPAVRVPDGQVFVMGDNRNNSNDSHVWGFLPQQNIIGHALFR 172
Query: 145 VWPPQRIGAVKNTT 158
+P R G + + T
Sbjct: 173 FFPASRWGQLGSFT 186
>LEPQ_BACNA (Q57350) Signal peptidase I P (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 185
Score = 43.9 bits (102), Expect = 2e-04
Identities = 39/127 (30%), Positives = 56/127 (43%), Gaps = 40/127 (31%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETH-IKRIIAL 92
V+G SM PT F + +FV K F GDIV+ + K+TH +KR+I L
Sbjct: 39 VQGESMKPTL------FNSERLFVNKFVKYTGDFKRGDIVVLNGEE--KKTHYVKRLIGL 90
Query: 93 PGEW--------FVN---------------RHNQDV--------LKVPEGHCWVEGDNAA 121
PG+ FVN H+ D+ +KVP+ +V GDN
Sbjct: 91 PGDTIEMKNDNLFVNGKRFNEEYLKENKKDAHDSDLNLTGDFGPIKVPKDKYFVMGDNRQ 150
Query: 122 SSTDSKS 128
+S DS++
Sbjct: 151 NSMDSRN 157
>LEP_MYCLE (O33021) Probable signal peptidase I (EC 3.4.21.89)
(SPase I) (Leader peptidase I)
Length = 289
Score = 42.4 bits (98), Expect = 5e-04
Identities = 24/72 (33%), Positives = 32/72 (44%), Gaps = 18/72 (25%)
Query: 108 VPEGHCWVEGDNAASSTDSKSY------------------GPVPLGLVRGRVTHVVWPPQ 149
VP+G WV GDN S DS+ + G VP+ V G+ VVWPP
Sbjct: 216 VPQGRLWVMGDNRIHSADSRYHCNSTDVVNGLSCTGDPNSGTVPVSNVIGKARVVVWPPS 275
Query: 150 RIGAVKNTTPER 161
R G V + ++
Sbjct: 276 RWGGVGSVNSQQ 287
>LEPV_BACSU (O07560) Signal peptidase I V (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 168
Score = 42.4 bits (98), Expect = 5e-04
Identities = 40/135 (29%), Positives = 56/135 (40%), Gaps = 45/135 (33%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHG-DIVIFSSPSNFKETHIKRIIAL 92
V G SM+PTF + + +FK H DIV+F P + + IKR+I L
Sbjct: 30 VEGVSMNPTFQEGNELLVNKFSH-------RFKTIHRFDIVLFKGPDH--KVLIKRVIGL 80
Query: 93 PGE--------WFVN---------RHNQDV------------------LKVPEGHCWVEG 117
PGE +VN +H + V KVP+G +V G
Sbjct: 81 PGETIKYKDDQLYVNGKQVAEPFLKHLKSVSAGSHVTGDFSLKDVTGTSKVPKGKYFVVG 140
Query: 118 DNAASSTDSKSYGPV 132
DN S DS+ +GP+
Sbjct: 141 DNRIYSFDSRHFGPI 155
>LEP_STAAM (P72365) Signal peptidase IB (EC 3.4.21.89) (SPase IB)
(Leader peptidase IB)
Length = 191
Score = 41.2 bits (95), Expect = 0.001
Identities = 43/181 (23%), Positives = 71/181 (38%), Gaps = 49/181 (27%)
Query: 6 LVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKF 65
L W ++ +A +I F V T ++G SM PT + V V +
Sbjct: 6 LEWIIS--IAVAFVILFIVGKFIVTPYTIKGESMDPTLKD------GERVAVNIIGYKTG 57
Query: 66 KFSHGDIVIFSSPSNFKETHIKRIIALPGE--------WFVNRHNQDV------LK---- 107
G++V+F + N + ++KR+I +PG+ +VN QD LK
Sbjct: 58 GLEKGNVVVFHANKN--DDYVKRVIGVPGDKVEYKNDTLYVNGKKQDEPYLNYNLKHKQG 115
Query: 108 ---------------------VPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVW 146
+P+G V GDN S DS+++G + + G+V+ W
Sbjct: 116 DYITGTFQVKDLPNANPKSNVIPKGKYLVLGDNREVSKDSRAFGLIDEDQIVGKVSFRFW 175
Query: 147 P 147
P
Sbjct: 176 P 176
>LEPU_BACSU (P42959) Signal peptidase I U (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 187
Score = 38.1 bits (87), Expect = 0.009
Identities = 34/149 (22%), Positives = 61/149 (40%), Gaps = 37/149 (24%)
Query: 10 VTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSH 69
V + I + FT+ + + G+SM+PT + + V+K F
Sbjct: 18 VVLSIIMIAALIFTIRLVFYKPFLIEGSSMAPTLKDS------ERILVDKAVKWTGGFHR 71
Query: 70 GDIVIFSSPSNFKETHIKRIIALPG------------------EWFVNRHNQDV------ 105
GDI++ + + + +KR+I LPG E ++ + Q+V
Sbjct: 72 GDIIVIHDKKSGR-SFVKRLIGLPGDSIKMKNDQLYINDKKVEEPYLKEYKQEVKESGVT 130
Query: 106 ------LKVPEGHCWVEGDNAASSTDSKS 128
++VP G +V GDN +S DS++
Sbjct: 131 LTGDFEVEVPSGKYFVMGDNRLNSLDSRN 159
>LEP_AQUAE (O67088) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 256
Score = 37.0 bits (84), Expect = 0.020
Identities = 25/83 (30%), Positives = 36/83 (43%), Gaps = 6/83 (7%)
Query: 13 KLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDI 72
+L I L + + A + ASM PT D++ V KL + GD+
Sbjct: 7 ELFLIILAVLFIREYIAQAYTIPSASMEPTL------LVGDFILVNKLVYSLSEPMRGDM 60
Query: 73 VIFSSPSNFKETHIKRIIALPGE 95
++F P N IKRIIA G+
Sbjct: 61 IVFKYPKNPDIDFIKRIIARGGD 83
Score = 36.6 bits (83), Expect = 0.027
Identities = 39/140 (27%), Positives = 63/140 (44%), Gaps = 26/140 (18%)
Query: 32 VPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIA 91
V V G T+ + N D Y + EKL + G+++ S F+ T +K
Sbjct: 102 VAVNGKLYELTYEGEKNYSYDCYQYREKLYRED-----GEVIQHSVC--FRNTLLK---- 150
Query: 92 LPGEWF-------VNRHNQD----VLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGR 140
+PG + ++N+D VPEG+ +V GDN +S DS+ +G VP + G+
Sbjct: 151 VPGMVYNAISSDLCLKYNEDGFCVKFVVPEGYYFVMGDNRDNSQDSRFWGFVPRENIEGK 210
Query: 141 VTHVVWPPQRIGAVKNTTPE 160
+ + G V + TPE
Sbjct: 211 AFVIYYS----GKVPSLTPE 226
>LEP_RICPR (Q9ZE32) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 264
Score = 36.6 bits (83), Expect = 0.027
Identities = 24/75 (32%), Positives = 33/75 (44%), Gaps = 20/75 (26%)
Query: 41 PTFNPKTNSFTDDYVFVEKLC-------LDKFKFSH-------------GDIVIFSSPSN 80
PT + K +DY+F K L F F H GDIV+F P++
Sbjct: 40 PTGSMKATILENDYIFSTKYSYGYSNYSLSFFDFIHLFKGRVFAREPERGDIVVFRPPND 99
Query: 81 FKETHIKRIIALPGE 95
+IKR+I LPG+
Sbjct: 100 MSVRYIKRLIGLPGD 114
Score = 28.1 bits (61), Expect = 9.4
Identities = 16/33 (48%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query: 102 NQDVLKVPEGHCWVEGDNAASSTDSK-SYGPVP 133
N DV VPEG + GDN S DS+ + G VP
Sbjct: 178 NTDVFYVPEGQYFFLGDNRDRSNDSRVNLGFVP 210
>LEP_BUCBP (Q89AM6) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 310
Score = 36.2 bits (82), Expect = 0.035
Identities = 26/89 (29%), Positives = 39/89 (43%), Gaps = 18/89 (20%)
Query: 19 LITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLD-KFKFSH-------- 69
+I F + + SM PT P D++ V+K K FS+
Sbjct: 63 IIVFIIRTFICEPFQIPSESMMPTLLP------GDFILVKKFSYGIKNPFSNNVIVFINT 116
Query: 70 ---GDIVIFSSPSNFKETHIKRIIALPGE 95
GDIV+F P+N ++KRI+ LPG+
Sbjct: 117 PKRGDIVVFKHPNNNAINYVKRIVGLPGD 145
Score = 30.4 bits (67), Expect = 1.9
Identities = 19/63 (30%), Positives = 29/63 (45%), Gaps = 10/63 (15%)
Query: 82 KETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRV 141
K+ + K+ G W V +H VL GDN +S DS+ +G VP + G+V
Sbjct: 234 KDLYFKQFSQKQGTWIVPKHKYFVL----------GDNRDNSLDSRYWGFVPEKNLIGKV 283
Query: 142 THV 144
+
Sbjct: 284 VFI 286
>LEP_RICCN (Q92JB1) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 266
Score = 34.3 bits (77), Expect = 0.13
Identities = 21/75 (28%), Positives = 31/75 (41%), Gaps = 20/75 (26%)
Query: 41 PTFNPKTNSFTDDYVFVEKLCLDKFKFS--------------------HGDIVIFSSPSN 80
PT + K +DY+F K +S GDIV+F P++
Sbjct: 42 PTGSMKATILENDYIFSTKYSYGYSNYSLSFFDFIPLFKGRIFAREPDRGDIVVFRPPND 101
Query: 81 FKETHIKRIIALPGE 95
+IKR+I LPG+
Sbjct: 102 MSVRYIKRLIGLPGD 116
Score = 28.5 bits (62), Expect = 7.2
Identities = 16/33 (48%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query: 102 NQDVLKVPEGHCWVEGDNAASSTDSK-SYGPVP 133
N DV VPEG + GDN S DS+ + G VP
Sbjct: 180 NTDVFYVPEGQYFFLGDNRDQSNDSRVNLGFVP 212
>LEP_BUCAI (P57347) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 314
Score = 34.3 bits (77), Expect = 0.13
Identities = 22/74 (29%), Positives = 33/74 (43%), Gaps = 18/74 (24%)
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEK------------LCLDKFKFSHGDIVIFSSPSNF 81
+ SM PT D++ VEK + + K + GDI +F P++
Sbjct: 84 IPSGSMMPTL------LVGDFILVEKFSYGIKEPITHKILIRTKKPNRGDIAVFQHPTDH 137
Query: 82 KETHIKRIIALPGE 95
+IKRII LPG+
Sbjct: 138 NINYIKRIIGLPGD 151
Score = 28.1 bits (61), Expect = 9.4
Identities = 18/62 (29%), Positives = 30/62 (48%), Gaps = 10/62 (16%)
Query: 72 IVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGP 131
I++ +S N KE + ++ W V P+G ++ GDN +S DS+ +G
Sbjct: 228 ILLLNSIKNTKENYFQQKNMPKLTWIV----------PKGEYFMMGDNRDNSLDSRYWGF 277
Query: 132 VP 133
VP
Sbjct: 278 VP 279
>LEP_STRPN (O07344) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 204
Score = 32.3 bits (72), Expect = 0.50
Identities = 18/55 (32%), Positives = 25/55 (44%)
Query: 98 VNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
VN + VPEG + GD+ S+DS+ G + G +WP RIG
Sbjct: 148 VNYNTNFSFTVPEGEYLLLGDDRLVSSDSRHVGTFKAKDITGEAKFRLWPITRIG 202
>LEP_PSEFL (P26844) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 284
Score = 32.0 bits (71), Expect = 0.65
Identities = 15/48 (31%), Positives = 28/48 (58%), Gaps = 2/48 (4%)
Query: 57 VEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQD 104
++K ++ GD+++F PS+ +IKR++ LPG+ V R+ D
Sbjct: 115 IDKKVIEVGDPQRGDVMVFRYPSDPNVNYIKRVVGLPGD--VVRYTSD 160
>LEP_STRR6 (P59662) Signal peptidase I (EC 3.4.21.89) (SPase I)
(Leader peptidase I)
Length = 204
Score = 31.6 bits (70), Expect = 0.85
Identities = 18/55 (32%), Positives = 24/55 (42%)
Query: 98 VNRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
VN + VPEG + GD+ S+DS+ G + G WP RIG
Sbjct: 148 VNYNTNFSFTVPEGEYLLLGDDRLVSSDSRHVGTFKAKDITGEAKFRFWPITRIG 202
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,385,272
Number of Sequences: 164201
Number of extensions: 904676
Number of successful extensions: 1670
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1623
Number of HSP's gapped (non-prelim): 47
length of query: 165
length of database: 59,974,054
effective HSP length: 102
effective length of query: 63
effective length of database: 43,225,552
effective search space: 2723209776
effective search space used: 2723209776
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146575.4


