

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146570.9 + phase: 0
(475 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
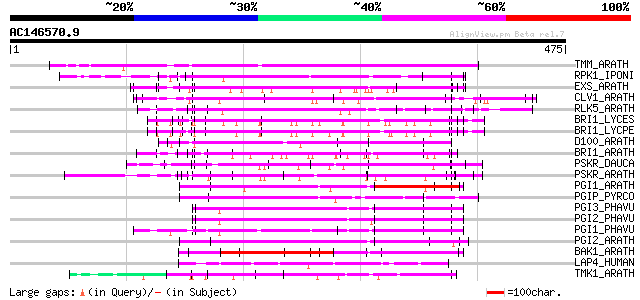
Score E
Sequences producing significant alignments: (bits) Value
TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM) 182 1e-45
RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC 2... 146 1e-34
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 127 8e-29
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 119 1e-26
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 118 3e-26
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 117 5e-26
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 117 8e-26
D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100 ... 113 1e-24
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 109 2e-23
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 101 4e-21
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 97 9e-20
PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor (Pol... 96 2e-19
PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor (Polyg... 96 2e-19
PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor (Pol... 91 5e-18
PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor (Pol... 91 6e-18
PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor (Pol... 88 5e-17
PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor (Pol... 82 4e-15
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 81 7e-15
LAP4_HUMAN (Q14160) LAP4 protein (Scribble homolog protein) (hSc... 80 1e-14
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 79 2e-14
>TMM_ARATH (Q9SSD1) TOO MANY MOUTHS protein precursor (TMM)
Length = 496
Score = 182 bits (463), Expect = 1e-45
Identities = 124/372 (33%), Positives = 198/372 (52%), Gaps = 17/372 (4%)
Query: 35 EKAEQEALYSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSC-DIFNGLWYVTVINIGPIHE 93
E EQ+A+Y ++ GN W + PD C G+ C + +++V ++ G + +
Sbjct: 55 EPDEQDAVYDIMRA-TGNDWAAA--IPDVCRGRW-HGIECMPDQDNVYHVVSLSFGALSD 110
Query: 94 NSL--PCANEKLEFKPELFQLKHLKAISFFNCF-QSPNKLPVSIPTGNWEKLAESLESIE 150
++ C ++ L +LKHLKA+ F+ C ++P ++P + +L SL+++
Sbjct: 111 DTAFPTCDPQRSYVSESLTRLKHLKALFFYRCLGRAPQRIPAFLG-----RLGSSLQTLV 165
Query: 151 FRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
R N G +G IP G L NL+ L L +N L G+IP L+ L LSGN +G+IP
Sbjct: 166 LREN-GFLGPIPDELGNLTNLKVLDLHKNHLNGSIPLSFNRFSGLRSLDLSGNRLTGSIP 224
Query: 211 DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLM 270
L L +LDL++N L+G +P TL S++K+DLS N + G + L L L+
Sbjct: 225 GFV--LPALSVLDLNQNLLTGPVPPTLTSCGSLIKIDLSRNRVTGPIPESINRLNQLVLL 282
Query: 271 DLRNNRLCCGLVLSLQEMNSLEEMVLSNNP-LGGDIRTLKWENLQNLVILELSNMELIGE 329
DL NRL SLQ +NSL+ ++L N I ++ L+NL+IL LSN + G
Sbjct: 283 DLSYNRLSGPFPSSLQGLNSLQALMLKGNTKFSTTIPENAFKGLKNLMILVLSNTNIQGS 342
Query: 330 IPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLG 389
IP+SL++L LR L L NN+TG + + + L+ L L+ N+L G + F + ++
Sbjct: 343 IPKSLTRLNSLRVLHLEGNNLTGEIPLEFRDVKHLSELRLNDNSLTGPVPFERDTVWRMR 402
Query: 390 RRFGAWSNPKLC 401
R+ ++N LC
Sbjct: 403 RKLRLYNNAGLC 414
>RPK1_IPONI (P93194) Receptor-like protein kinase precursor (EC
2.7.1.37)
Length = 1109
Score = 146 bits (368), Expect = 1e-34
Identities = 115/348 (33%), Positives = 166/348 (47%), Gaps = 29/348 (8%)
Query: 43 YSTIQGFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEK 102
+++I + SWN SD PC S GV CD +V +N+ +
Sbjct: 38 WTSIPSDITQSWNASD--STPC---SWLGVECDRRQ---FVDTLNLSSYGISG------- 82
Query: 103 LEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGN 160
EF PE+ LKHLK + S N SIP+ GN LE I+ SN GN
Sbjct: 83 -EFGPEISHLKHLKKVVL-----SGNGFFGSIPSQLGN----CSLLEHIDLSSN-SFTGN 131
Query: 161 IPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLL 220
IP T G L+NL++L L N L G P+ + ++ L+ + +GN +G+IP G +S+L
Sbjct: 132 IPDTLGALQNLRNLSLFFNSLIGPFPESLLSIPHLETVYFTGNGLNGSIPSNIGNMSELT 191
Query: 221 ILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCG 280
L L N SG +P +LG + ++ +L L+ N L G L NL+NL +D+RNN L
Sbjct: 192 TLWLDDNQFSGPVPSSLGNITTLQELYLNDNNLVGTLPVTLNNLENLVYLDVRNNSLVGA 251
Query: 281 LVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKL 340
+ L ++ + LSNN G + N +L + L G IP QL KL
Sbjct: 252 IPLDFVSCKQIDTISLSNNQFTGGLPP-GLGNCTSLREFGAFSCALSGPIPSCFGQLTKL 310
Query: 341 RFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
L L+ N+ +G + P+L S+ L L N L+GEI G +L
Sbjct: 311 DTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGMLSQL 358
Score = 120 bits (302), Expect = 6e-27
Identities = 87/245 (35%), Positives = 118/245 (47%), Gaps = 26/245 (10%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGL-----------------------TGNIPQEIGNLV 193
L G++PS G L+ L+L EN L TG IP +GNL
Sbjct: 464 LEGSVPSDLGGCSTLERLILEENNLRGGLPDFVEKQNLLFFDLSGNNFTGPIPPSLGNLK 523
Query: 194 KLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFL 253
+ + LS N SG+IP G L L L+LS N L G LP L + +LD SHN L
Sbjct: 524 NVTAIYLSSNQLSGSIPPELGSLVKLEHLNLSHNILKGILPSELSNCHKLSELDASHNLL 583
Query: 254 EGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENL 313
G + + G+L LT + L N G+ SL + N L + L N L GDI + L
Sbjct: 584 NGSIPSTLGSLTELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGDIPPV--GAL 641
Query: 314 QNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNN 373
Q L L LS+ +L G++P L +LK L L +S NN++G L L T+ SL + +S N
Sbjct: 642 QALRSLNLSSNKLNGQLPIDLGKLKMLEELDVSHNNLSGTLR-VLSTIQSLTFINISHNL 700
Query: 374 LKGEI 378
G +
Sbjct: 701 FSGPV 705
Score = 118 bits (296), Expect = 3e-26
Identities = 82/247 (33%), Positives = 119/247 (47%), Gaps = 2/247 (0%)
Query: 144 ESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGN 203
E+L ++ R+N L+G IP F K + ++ L N TG +P +GN L+
Sbjct: 236 ENLVYLDVRNN-SLVGAIPLDFVSCKQIDTISLSNNQFTGGLPPGLGNCTSLREFGAFSC 294
Query: 204 NFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGN 263
SG IP FG L+ L L L+ N SG +P LG+ S++ L L N LEG++ E G
Sbjct: 295 ALSGPIPSCFGQLTKLDTLYLAGNHFSGRIPPELGKCKSMIDLQLQQNQLEGEIPGELGM 354
Query: 264 LKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSN 323
L L + L N L + LS+ ++ SL+ + L N L G++ + L+ LV L L
Sbjct: 355 LSQLQYLHLYTNNLSGEVPLSIWKIQSLQSLQLYQNNLSGEL-PVDMTELKQLVSLALYE 413
Query: 324 MELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
G IP+ L L L L+ N TG++ P L + L L L N L+G + G
Sbjct: 414 NHFTGVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQKKLKRLLLGYNYLEGSVPSDLG 473
Query: 384 FFGKLGR 390
L R
Sbjct: 474 GCSTLER 480
Score = 115 bits (287), Expect = 3e-25
Identities = 88/261 (33%), Positives = 134/261 (50%), Gaps = 5/261 (1%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
N+L IP G L++ L+ + +N L G +P + +++LQSL L +N L+G +P
Sbjct: 342 NQLEGEIP-GELGMLSQ-LQYLHLYTN-NLSGEVPLSIWKIQSLQSLQLYQNNLSGELPV 398
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
++ L +L L L N+F+G IP G S L +LDL+RN +G +P L + +L
Sbjct: 399 DMTELKQLVSLALYENHFTGVIPQDLGANSSLEVLDLTRNMFTGHIPPNLCSQKKLKRLL 458
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
L +N+LEG + ++ G L + L N L GL +++ N L + NN G +
Sbjct: 459 LGYNYLEGSVPSDLGGCSTLERLILEENNLRGGLPDFVEKQNLLFFDLSGNNFTGPIPPS 518
Query: 308 LKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNAL 367
L NL+N+ + LS+ +L G IP L L KL L LS N + G L +L L+ L
Sbjct: 519 L--GNLKNVTAIYLSSNQLSGSIPPELGSLVKLEHLNLSHNILKGILPSELSNCHKLSEL 576
Query: 368 YLSGNNLKGEIQFSKGFFGKL 388
S N L G I + G +L
Sbjct: 577 DASHNLLNGSIPSTLGSLTEL 597
Score = 102 bits (254), Expect = 2e-21
Identities = 70/203 (34%), Positives = 107/203 (52%), Gaps = 8/203 (3%)
Query: 151 FRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
+ S+ L G+IP G L L+ L L N L G +P E+ N KL L S N +G+IP
Sbjct: 529 YLSSNQLSGSIPPELGSLVKLEHLNLSHNILKGILPSELSNCHKLSELDASHNLLNGSIP 588
Query: 211 DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLM 270
G L++L L L NS SG +P +L + +L L L N L G + G L+ L +
Sbjct: 589 STLGSLTELTKLSLGENSFSGGIPTSLFQSNKLLNLQLGGNLLAGD-IPPVGALQALRSL 647
Query: 271 DLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEI 330
+L +N+L L + L ++ LEE+ +S+N L G +R L +Q+L + +S+ G +
Sbjct: 648 NLSSNKLNGQLPIDLGKLKMLEELDVSHNNLSGTLRVL--STIQSLTFINISHNLFSGPV 705
Query: 331 PESLSQLKKLRFLGLSDNNITGN 353
P SL+ +FL S + +GN
Sbjct: 706 PPSLT-----KFLNSSPTSFSGN 723
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells
protein) (EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 127 bits (318), Expect = 8e-29
Identities = 88/237 (37%), Positives = 123/237 (51%), Gaps = 13/237 (5%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG--- 215
G IP G +L +L L N L G IP +I L +L+ LVLS NN SG+IP
Sbjct: 510 GKIPVELGDCTSLTTLDLGSNNLQGQIPDKITALAQLQCLVLSYNNLSGSIPSKPSAYFH 569
Query: 216 ---LSDLLIL------DLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKN 266
+ DL L DLS N LSG +P LG + ++++ LS+N L G++ L N
Sbjct: 570 QIEMPDLSFLQHHGIFDLSYNRLSGPIPEELGECLVLVEISLSNNHLSGEIPASLSRLTN 629
Query: 267 LTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMEL 326
LT++DL N L + + L+ + L+NN L G I + L +LV L L+ +L
Sbjct: 630 LTILDLSGNALTGSIPKEMGNSLKLQGLNLANNQLNGHIPE-SFGLLGSLVKLNLTKNKL 688
Query: 327 IGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
G +P SL LK+L + LS NN++G LS +L T+ L LY+ N GEI G
Sbjct: 689 DGPVPASLGNLKELTHMDLSFNNLSGELSSELSTMEKLVGLYIEQNKFTGEIPSELG 745
Score = 125 bits (314), Expect = 2e-28
Identities = 96/273 (35%), Positives = 132/273 (48%), Gaps = 17/273 (6%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQ 187
N++ SIP W+ L +++ SN G IP + NL N L G +P
Sbjct: 411 NQINGSIPEDLWKL---PLMALDLDSN-NFTGEIPKSLWKSTNLMEFTASYNRLEGYLPA 466
Query: 188 EIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD 247
EIGN LKRLVLS N +G IP G L+ L +L+L+ N G +PV LG S+ LD
Sbjct: 467 EIGNAASLKRLVLSDNQLTGEIPREIGKLTSLSVLNLNANMFQGKIPVELGDCTSLTTLD 526
Query: 248 LSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGL---------VLSLQEMNSLEE---MV 295
L N L+G++ ++ L L + L N L + + + +++ L+
Sbjct: 527 LGSNNLQGQIPDKITALAQLQCLVLSYNNLSGSIPSKPSAYFHQIEMPDLSFLQHHGIFD 586
Query: 296 LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLS 355
LS N L G I E L LV + LSN L GEIP SLS+L L L LS N +TG++
Sbjct: 587 LSYNRLSGPIPEELGECLV-LVEISLSNNHLSGEIPASLSRLTNLTILDLSGNALTGSIP 645
Query: 356 PKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
++ L L L+ N L G I S G G L
Sbjct: 646 KEMGNSLKLQGLNLANNQLNGHIPESFGLLGSL 678
Score = 120 bits (302), Expect = 6e-27
Identities = 80/207 (38%), Positives = 116/207 (55%), Gaps = 6/207 (2%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFR-SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGN 184
S N+L IP E+L E L +E SN L G IP++ L NL L L N LTG+
Sbjct: 588 SYNRLSGPIP----EELGECLVLVEISLSNNHLSGEIPASLSRLTNLTILDLSGNALTGS 643
Query: 185 IPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVL 244
IP+E+GN +KL+ L L+ N +G+IP+ FG L L+ L+L++N L G +P +LG L +
Sbjct: 644 IPKEMGNSLKLQGLNLANNQLNGHIPESFGLLGSLVKLNLTKNKLDGPVPASLGNLKELT 703
Query: 245 KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGD 304
+DLS N L G+L +E ++ L + + N+ + L + LE + +S N L G+
Sbjct: 704 HMDLSFNNLSGELSSELSTMEKLVGLYIEQNKFTGEIPSELGNLTQLEYLDVSENLLSGE 763
Query: 305 IRTLKWENLQNLVILELSNMELIGEIP 331
I T K L NL L L+ L GE+P
Sbjct: 764 IPT-KICGLPNLEFLNLAKNNLRGEVP 789
Score = 120 bits (300), Expect = 1e-26
Identities = 103/311 (33%), Positives = 149/311 (47%), Gaps = 45/311 (14%)
Query: 104 EFKPELFQLKHLKAISF-FNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIP 162
E E+ +L L ++ N FQ K+PV + SL +++ SN L G IP
Sbjct: 487 EIPREIGKLTSLSVLNLNANMFQG--KIPVELGD------CTSLTTLDLGSN-NLQGQIP 537
Query: 163 STFGVLKNLQSLVLLENGLTGNIPQ---------EIGNLVKLKR---LVLSGNNFSGNIP 210
L LQ LVL N L+G+IP E+ +L L+ LS N SG IP
Sbjct: 538 DKITALAQLQCLVLSYNNLSGSIPSKPSAYFHQIEMPDLSFLQHHGIFDLSYNRLSGPIP 597
Query: 211 DIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLM 270
+ G L+ + LS N LSG +P +L RL ++ LDLS N L G + E GN L +
Sbjct: 598 EELGECLVLVEISLSNNHLSGEIPASLSRLTNLTILDLSGNALTGSIPKEMGNSLKLQGL 657
Query: 271 DLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI------------RTLKWENLQNLVI 318
+L NN+L + S + SL ++ L+ N L G + L + NL +
Sbjct: 658 NLANNQLNGHIPESFGLLGSLVKLNLTKNKLDGPVPASLGNLKELTHMDLSFNNLSGELS 717
Query: 319 LELSNMELI-----------GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNAL 367
ELS ME + GEIP L L +L +L +S+N ++G + K+ LP+L L
Sbjct: 718 SELSTMEKLVGLYIEQNKFTGEIPSELGNLTQLEYLDVSENLLSGEIPTKICGLPNLEFL 777
Query: 368 YLSGNNLKGEI 378
L+ NNL+GE+
Sbjct: 778 NLAKNNLRGEV 788
Score = 113 bits (283), Expect = 9e-25
Identities = 98/298 (32%), Positives = 131/298 (43%), Gaps = 42/298 (14%)
Query: 128 NKLPVSIPTGNWEKLAESLESIEFRSNPG-LIGNIPSTFGVLKNLQSLVLLENGLTGNIP 186
N IP W+ S +EF ++ L G +P+ G +L+ LVL +N LTG IP
Sbjct: 434 NNFTGEIPKSLWK----STNLMEFTASYNRLEGYLPAEIGNAASLKRLVLSDNQLTGEIP 489
Query: 187 QEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKL 246
+EIG L L L L+ N F G IP G + L LDL N+L G +P + L + L
Sbjct: 490 REIGKLTSLSVLNLNANMFQGKIPVELGDCTSLTTLDLGSNNLQGQIPDKITALAQLQCL 549
Query: 247 DLSHNFLEGKL------------LNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEM 294
LS+N L G + + + L++ + DL NRL + L E L E+
Sbjct: 550 VLSYNNLSGSIPSKPSAYFHQIEMPDLSFLQHHGIFDLSYNRLSGPIPEELGECLVLVEI 609
Query: 295 VLSNNPLGGDIRTLKWENLQNLVILELS------------------------NMELIGEI 330
LSNN L G+I L NL IL+LS N +L G I
Sbjct: 610 SLSNNHLSGEIPA-SLSRLTNLTILDLSGNALTGSIPKEMGNSLKLQGLNLANNQLNGHI 668
Query: 331 PESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
PES L L L L+ N + G + L L L + LS NNL GE+ KL
Sbjct: 669 PESFGLLGSLVKLNLTKNKLDGPVPASLGNLKELTHMDLSFNNLSGELSSELSTMEKL 726
Score = 112 bits (279), Expect = 3e-24
Identities = 95/301 (31%), Positives = 143/301 (46%), Gaps = 41/301 (13%)
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFR--SNPGLIGNIPST 164
PE++ LKHL+ + S N L TG +L L + + S+ G++P +
Sbjct: 107 PEIWNLKHLQTLDL-----SGNSL-----TGLLPRLLSELPQLLYLDLSDNHFSGSLPPS 156
Query: 165 FGV-LKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILD 223
F + L L SL + N L+G IP EIG L L L + N+FSG IP G +S L
Sbjct: 157 FFISLPALSSLDVSNNSLSGEIPPEIGKLSNLSNLYMGLNSFSGQIPSEIGNISLLKNFA 216
Query: 224 LSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVL 283
+G LP + +L + KLDLS+N L+ + FG L NL++++L + L +
Sbjct: 217 APSCFFNGPLPKEISKLKHLAKLDLSYNPLKCSIPKSFGELHNLSILNLVSAELIGLIPP 276
Query: 284 SLQEMNSLEEMVLSNNPLGG-------DIRTL------------------KWENLQNLVI 318
L SL+ ++LS N L G +I L KW+ L +L+
Sbjct: 277 ELGNCKSLKSLMLSFNSLSGPLPLELSEIPLLTFSAERNQLSGSLPSWMGKWKVLDSLL- 335
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L+N GEIP + L+ L L+ N ++G++ +L SL A+ LSGN L G I
Sbjct: 336 --LANNRFSGEIPHEIEDCPMLKHLSLASNLLSGSIPRELCGSGSLEAIDLSGNLLSGTI 393
Query: 379 Q 379
+
Sbjct: 394 E 394
Score = 110 bits (275), Expect = 8e-24
Identities = 81/227 (35%), Positives = 115/227 (49%), Gaps = 10/227 (4%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP LKNL+ L L N +G IP EI NL L+ L LSGN+ +G +P + L
Sbjct: 77 LRGQIPKEISSLKNLRELCLAGNQFSGKIPPEIWNLKHLQTLDLSGNSLTGLLPRLLSEL 136
Query: 217 SDLLILDLSRNSLSGTLPVTLG-RLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
LL LDLS N SG+LP + L ++ LD+S+N L G++ E G L NL+ + + N
Sbjct: 137 PQLLYLDLSDNHFSGSLPPSFFISLPALSSLDVSNNSLSGEIPPEIGKLSNLSNLYMGLN 196
Query: 276 RLCCGLVLSLQEMNSLEEMV----LSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
+ + ++ L+ N PL +I LK +L L+LS L IP
Sbjct: 197 SFSGQIPSEIGNISLLKNFAAPSCFFNGPLPKEISKLK-----HLAKLDLSYNPLKCSIP 251
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+S +L L L L + G + P+L SL +L LS N+L G +
Sbjct: 252 KSFGELHNLSILNLVSAELIGLIPPELGNCKSLKSLMLSFNSLSGPL 298
Score = 110 bits (275), Expect = 8e-24
Identities = 95/274 (34%), Positives = 132/274 (47%), Gaps = 15/274 (5%)
Query: 107 PELFQLKHLKAISF-FNCFQSPNKLPVS-IPTGNWEKLAESLESIEFRSNPGLIGNIPST 164
PEL K LK++ FN P L +S IP L S E + L G++PS
Sbjct: 276 PELGNCKSLKSLMLSFNSLSGPLPLELSEIPL-----LTFSAERNQ------LSGSLPSW 324
Query: 165 FGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDL 224
G K L SL+L N +G IP EI + LK L L+ N SG+IP G L +DL
Sbjct: 325 MGKWKVLDSLLLANNRFSGEIPHEIEDCPMLKHLSLASNLLSGSIPRELCGSGSLEAIDL 384
Query: 225 SRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLS 284
S N LSGT+ S+ +L L++N + G + + L L +DL +N + S
Sbjct: 385 SGNLLSGTIEEVFDGCSSLGELLLTNNQINGSIPEDLWKLP-LMALDLDSNNFTGEIPKS 443
Query: 285 LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLG 344
L + +L E S N L G + + N +L L LS+ +L GEIP + +L L L
Sbjct: 444 LWKSTNLMEFTASYNRLEGYL-PAEIGNAASLKRLVLSDNQLTGEIPREIGKLTSLSVLN 502
Query: 345 LSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L+ N G + +L SL L L NNL+G+I
Sbjct: 503 LNANMFQGKIPVELGDCTSLTTLDLGSNNLQGQI 536
Score = 100 bits (250), Expect = 6e-21
Identities = 84/245 (34%), Positives = 120/245 (48%), Gaps = 4/245 (1%)
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
L ++ NP L +IP +FG L NL L L+ L G IP E+GN LK L+LS N+
Sbjct: 236 LAKLDLSYNP-LKCSIPKSFGELHNLSILNLVSAELIGLIPPELGNCKSLKSLMLSFNSL 294
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLK 265
SG +P + LL RN LSG+LP +G+ + L L++N G++ +E +
Sbjct: 295 SGPLPLELSEIP-LLTFSAERNQLSGSLPSWMGKWKVLDSLLLANNRFSGEIPHEIEDCP 353
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
L + L +N L + L SLE + LS N L G I + ++ +L L L+N +
Sbjct: 354 MLKHLSLASNLLSGSIPRELCGSGSLEAIDLSGNLLSGTIEEV-FDGCSSLGELLLTNNQ 412
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFF 385
+ G IPE L +L L L L NN TG + L +L S N L+G + G
Sbjct: 413 INGSIPEDLWKLP-LMALDLDSNNFTGEIPKSLWKSTNLMEFTASYNRLEGYLPAEIGNA 471
Query: 386 GKLGR 390
L R
Sbjct: 472 ASLKR 476
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 119 bits (299), Expect = 1e-26
Identities = 96/295 (32%), Positives = 141/295 (47%), Gaps = 16/295 (5%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGN-NFSGNIP-DIFG 214
L G I G+L +L +L L N TG +P E+ +L LK L +S N N +G P +I
Sbjct: 82 LFGTISPEIGMLTHLVNLTLAANNFTGELPLEMKSLTSLKVLNISNNGNLTGTFPGEILK 141
Query: 215 GLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRN 274
+ DL +LD N+ +G LP + L + L NF G++ +G++++L + L
Sbjct: 142 AMVDLEVLDTYNNNFNGKLPPEMSELKKLKYLSFGGNFFSGEIPESYGDIQSLEYLGLNG 201
Query: 275 NRLCCGLVLSLQEMNSLEEMVLS--NNPLGGDIRTLKWENLQNLVILELSNMELIGEIPE 332
L L + +L EM + N+ GG ++ L L IL++++ L GEIP
Sbjct: 202 AGLSGKSPAFLSRLKNLREMYIGYYNSYTGG--VPPEFGGLTKLEILDMASCTLTGEIPT 259
Query: 333 SLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRF 392
SLS LK L L L NN+TG++ P+L L SL +L LS N L GEI S F LG
Sbjct: 260 SLSNLKHLHTLFLHINNLTGHIPPELSGLVSLKSLDLSINQLTGEIPQS---FINLG--- 313
Query: 393 GAWSNPKLCYPFELMSTNNVPYGVKPCHQEEIHLVKSNAKTEVINGDINHNSNFI 447
N L F +P + + E+ V N T + ++ N N I
Sbjct: 314 ----NITLINLFRNNLYGQIPEAIGELPKLEVFEVWENNFTLQLPANLGRNGNLI 364
Score = 111 bits (277), Expect = 5e-24
Identities = 89/275 (32%), Positives = 137/275 (49%), Gaps = 14/275 (5%)
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFR--SNPGLIGNIPST 164
PE+ +LK LK +SF F S ++P S ++S+E+ + GL G P+
Sbjct: 162 PEMSELKKLKYLSFGGNFFS-GEIPESYG---------DIQSLEYLGLNGAGLSGKSPAF 211
Query: 165 FGVLKNLQSLVL-LENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILD 223
LKNL+ + + N TG +P E G L KL+ L ++ +G IP L L L
Sbjct: 212 LSRLKNLREMYIGYYNSYTGGVPPEFGGLTKLEILDMASCTLTGEIPTSLSNLKHLHTLF 271
Query: 224 LSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVL 283
L N+L+G +P L L+S+ LDLS N L G++ F NL N+TL++L N L +
Sbjct: 272 LHINNLTGHIPPELSGLVSLKSLDLSINQLTGEIPQSFINLGNITLINLFRNNLYGQIPE 331
Query: 284 SLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFL 343
++ E+ LE + N + N NL+ L++S+ L G IP+ L + +KL L
Sbjct: 332 AIGELPKLEVFEVWENNFTLQLPANLGRN-GNLIKLDVSDNHLTGLIPKDLCRGEKLEML 390
Query: 344 GLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
LS+N G + +L SL + + N L G +
Sbjct: 391 ILSNNFFFGPIPEELGKCKSLTKIRIVKNLLNGTV 425
Score = 110 bits (275), Expect = 8e-24
Identities = 91/305 (29%), Positives = 141/305 (45%), Gaps = 12/305 (3%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP++ LK+L +L L N LTG+IP E+ LV LK L LS N +G IP F L
Sbjct: 253 LTGEIPTSLSNLKHLHTLFLHINNLTGHIPPELSGLVSLKSLDLSINQLTGEIPQSFINL 312
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNR 276
++ +++L RN+L G +P +G L + ++ N +L G NL +D+ +N
Sbjct: 313 GNITLINLFRNNLYGQIPEAIGELPKLEVFEVWENNFTLQLPANLGRNGNLIKLDVSDNH 372
Query: 277 LCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
L + L LE ++LSNN G I + ++L + + L G +P L
Sbjct: 373 LTGLIPKDLCRGEKLEMLILSNNFFFGPIPE-ELGKCKSLTKIRIVKNLLNGTVPAGLFN 431
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWS 396
L + + L+DN +G L P + L+ +YLS N GEI + G F L F +
Sbjct: 432 LPLVTIIELTDNFFSGEL-PVTMSGDVLDQIYLSNNWFSGEIPPAIGNFPNLQTLFLDRN 490
Query: 397 NPKLCYPFELM----------STNNVPYGVKPCHQEEIHLVKSNAKTEVINGDINHNSNF 446
+ P E+ S NN+ G+ L+ + ING+I N
Sbjct: 491 RFRGNIPREIFELKHLSRINTSANNITGGIPDSISRCSTLISVDLSRNRINGEIPKGINN 550
Query: 447 ITSMG 451
+ ++G
Sbjct: 551 VKNLG 555
Score = 107 bits (267), Expect = 7e-23
Identities = 103/349 (29%), Positives = 162/349 (45%), Gaps = 28/349 (8%)
Query: 107 PELFQLKHLKAISFF-NCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTF 165
PE+ L HL ++ N F +LP+ + K SL+ + +N L G P
Sbjct: 88 PEIGMLTHLVNLTLAANNFTG--ELPLEM------KSLTSLKVLNISNNGNLTGTFPGE- 138
Query: 166 GVLKNLQSLVLLE---NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLIL 222
+LK + L +L+ N G +P E+ L KLK L GN FSG IP+ +G + L L
Sbjct: 139 -ILKAMVDLEVLDTYNNNFNGKLPPEMSELKKLKYLSFGGNFFSGEIPESYGDIQSLEYL 197
Query: 223 DLSRNSLSGTLPVTLGRLISVLKLDLS-HNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGL 281
L+ LSG P L RL ++ ++ + +N G + EFG L L ++D+ + L +
Sbjct: 198 GLNGAGLSGKSPAFLSRLKNLREMYIGYYNSYTGGVPPEFGGLTKLEILDMASCTLTGEI 257
Query: 282 VLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLR 341
SL + L + L N L G I + L +L L+LS +L GEIP+S L +
Sbjct: 258 PTSLSNLKHLHTLFLHINNLTGHIPP-ELSGLVSLKSLDLSINQLTGEIPQSFINLGNIT 316
Query: 342 FLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSN---- 397
+ L NN+ G + + LP L + NN ++ + G G L + + ++
Sbjct: 317 LINLFRNNLYGQIPEAIGELPKLEVFEVWENNFTLQLPANLGRNGNLIKLDVSDNHLTGL 376
Query: 398 -PK-LCYPFE---LMSTNNVPYGVKPCHQEEIHLVKSNAKTEVINGDIN 441
PK LC + L+ +NN +G P EE+ KS K ++ +N
Sbjct: 377 IPKDLCRGEKLEMLILSNNFFFGPIP---EELGKCKSLTKIRIVKNLLN 422
Score = 99.4 bits (246), Expect = 2e-20
Identities = 91/313 (29%), Positives = 138/313 (44%), Gaps = 31/313 (9%)
Query: 114 HLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQS 173
+L I+ N F+ N L IP E LE E N + +P+ G NL
Sbjct: 311 NLGNITLINLFR--NNLYGQIPEAIGE--LPKLEVFEVWENNFTL-QLPANLGRNGNLIK 365
Query: 174 LVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTL 233
L + +N LTG IP+++ KL+ L+LS N F G IP+ G L + + +N L+GT+
Sbjct: 366 LDVSDNHLTGLIPKDLCRGEKLEMLILSNNFFFGPIPEELGKCKSLTKIRIVKNLLNGTV 425
Query: 234 PVTLGRLISVLKLDLSHNFLEGKL--------LNEF---------------GNLKNLTLM 270
P L L V ++L+ NF G+L L++ GN NL +
Sbjct: 426 PAGLFNLPLVTIIELTDNFFSGELPVTMSGDVLDQIYLSNNWFSGEIPPAIGNFPNLQTL 485
Query: 271 DLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEI 330
L NR + + E+ L + S N + G I L+ ++LS + GEI
Sbjct: 486 FLDRNRFRGNIPREIFELKHLSRINTSANNITGGIPD-SISRCSTLISVDLSRNRINGEI 544
Query: 331 PESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
P+ ++ +K L L +S N +TG++ + + SL L LS N+L G + F
Sbjct: 545 PKGINNVKNLGTLNISGNQLTGSIPTGIGNMTSLTTLDLSFNDLSGRVPLGGQFLVFNET 604
Query: 391 RFGAWSNPKLCYP 403
F N LC P
Sbjct: 605 SFA--GNTYLCLP 615
Score = 88.2 bits (217), Expect = 4e-17
Identities = 61/169 (36%), Positives = 88/169 (51%), Gaps = 9/169 (5%)
Query: 109 LFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVL 168
LF L + I + F S +LPV++ + L+ I + SN G IP G
Sbjct: 429 LFNLPLVTIIELTDNFFS-GELPVTMS-------GDVLDQI-YLSNNWFSGEIPPAIGNF 479
Query: 169 KNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNS 228
NLQ+L L N GNIP+EI L L R+ S NN +G IPD S L+ +DLSRN
Sbjct: 480 PNLQTLFLDRNRFRGNIPREIFELKHLSRINTSANNITGGIPDSISRCSTLISVDLSRNR 539
Query: 229 LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
++G +P + + ++ L++S N L G + GN+ +LT +DL N L
Sbjct: 540 INGEIPKGINNVKNLGTLNISGNQLTGSIPTGIGNMTSLTTLDLSFNDL 588
Score = 58.9 bits (141), Expect = 3e-08
Identities = 44/157 (28%), Positives = 74/157 (47%), Gaps = 3/157 (1%)
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
++ L++S L GT+ +G L ++ L L+ N G+L E +L +L ++++ NN
Sbjct: 72 VISLNVSFTPLFGTISPEIGMLTHLVNLTLAANNFTGELPLEMKSLTSLKVLNISNNGNL 131
Query: 279 CGLVLS--LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQ 336
G L+ M LE + NN G + + L+ L L GEIPES
Sbjct: 132 TGTFPGEILKAMVDLEVLDTYNNNFNGKLPP-EMSELKKLKYLSFGGNFFSGEIPESYGD 190
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNN 373
++ L +LGL+ ++G L L +L +Y+ N
Sbjct: 191 IQSLEYLGLNGAGLSGKSPAFLSRLKNLREMYIGYYN 227
Score = 35.4 bits (80), Expect = 0.32
Identities = 23/62 (37%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Query: 316 LVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN-NL 374
++ L +S L G I + L L L L+ NN TG L ++++L SL L +S N NL
Sbjct: 72 VISLNVSFTPLFGTISPEIGMLTHLVNLTLAANNFTGELPLEMKSLTSLKVLNISNNGNL 131
Query: 375 KG 376
G
Sbjct: 132 TG 133
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 118 bits (296), Expect = 3e-26
Identities = 80/222 (36%), Positives = 118/222 (53%), Gaps = 6/222 (2%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G IP+ L+ L+L++N +G I +G L R+ LS N SG IP F GL
Sbjct: 369 GEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRLSNNKLSGQIPHGFWGLPR 428
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L +L+LS NS +G++P T+ ++ L +S N G + NE G+L + + N
Sbjct: 429 LSLLELSDNSFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNEIGSLNGIIEISGAENDFS 488
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDI-RTLK-WENLQNLVILELSNMELIGEIPESLSQ 336
+ SL ++ L + LS N L G+I R L+ W+NL L L+N L GEIP+ +
Sbjct: 489 GEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGWKNLNE---LNLANNHLSGEIPKEVGI 545
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L L +L LS N +G + +L+ L LN L LS N+L G+I
Sbjct: 546 LPVLNYLDLSSNQFSGEIPLELQNL-KLNVLNLSYNHLSGKI 586
Score = 115 bits (287), Expect = 3e-25
Identities = 104/322 (32%), Positives = 150/322 (46%), Gaps = 27/322 (8%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S NKL IP N L +LES+ N L G +P + K L L L N LTG +
Sbjct: 292 SMNKLTGKIPD-NLNLL--NLESLNLFENM-LEGPLPESITRSKTLSELKLFNNRLTGVL 347
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P ++G L+ + LS N FSG IP G L L L NS SG + LG+ S+ +
Sbjct: 348 PSQLGANSPLQYVDLSYNRFSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTR 407
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+ LS+N L G++ + F L L+L++L +N + ++ +L + +S N G I
Sbjct: 408 VRLSNNKLSGQIPHGFWGLPRLSLLELSDNSFTGSIPKTIIGAKNLSNLRISKNRFSGSI 467
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ +L ++ + + + GEIPESL +LK+L L LS N ++G + +L +LN
Sbjct: 468 PN-EIGSLNGIIEISGAENDFSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGWKNLN 526
Query: 366 ALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPFELMSTNNVPYGVKPCHQEEIH 425
L L+ N+L GEI G P L Y +L S EI
Sbjct: 527 ELNLANNHLSGEIPKEVGIL------------PVLNY-LDLSSNQ---------FSGEIP 564
Query: 426 LVKSNAKTEVINGDINHNSNFI 447
L N K V+N NH S I
Sbjct: 565 LELQNLKLNVLNLSYNHLSGKI 586
Score = 108 bits (270), Expect = 3e-23
Identities = 95/301 (31%), Positives = 146/301 (47%), Gaps = 35/301 (11%)
Query: 110 FQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLK 169
F L +LK + S N L +IP+ E LES+ N L G IP++ G +
Sbjct: 136 FNLPNLKFLEI-----SGNNLSDTIPSSFGE--FRKLESLNLAGN-FLSGTIPASLGNVT 187
Query: 170 NLQSLVLLENGLT-GNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNS 228
L+ L L N + IP ++GNL +L+ L L+G N G IP L+ L+ LDL+ N
Sbjct: 188 TLKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLTFNQ 247
Query: 229 LSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGL-----VL 283
L+G++P + +L +V +++L +N G+L GN+ L D N+L + +L
Sbjct: 248 LTGSIPSWITQLKTVEQIELFNNSFSGELPESMGNMTTLKRFDASMNKLTGKIPDNLNLL 307
Query: 284 SLQEMN------------------SLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
+L+ +N +L E+ L NN L G + + N L ++LS
Sbjct: 308 NLESLNLFENMLEGPLPESITRSKTLSELKLFNNRLTGVLPSQLGAN-SPLQYVDLSYNR 366
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFF 385
GEIP ++ KL +L L DN+ +G +S L SL + LS N L G+I GF+
Sbjct: 367 FSGEIPANVCGEGKLEYLILIDNSFSGEISNNLGKCKSLTRVRLSNNKLSGQI--PHGFW 424
Query: 386 G 386
G
Sbjct: 425 G 425
Score = 102 bits (254), Expect = 2e-21
Identities = 68/198 (34%), Positives = 106/198 (53%), Gaps = 2/198 (1%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G I + G K+L + L N L+G IP L +L L LS N+F+G+IP G +
Sbjct: 393 GEISNNLGKCKSLTRVRLSNNKLSGQIPHGFWGLPRLSLLELSDNSFTGSIPKTIIGAKN 452
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L L +S+N SG++P +G L ++++ + N G++ LK L+ +DL N+L
Sbjct: 453 LSNLRISKNRFSGSIPNEIGSLNGIIEISGAENDFSGEIPESLVKLKQLSRLDLSKNQLS 512
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK 338
+ L+ +L E+ L+NN L G+I + L L L+LS+ + GEIP L L
Sbjct: 513 GEIPRELRGWKNLNELNLANNHLSGEI-PKEVGILPVLNYLDLSSNQFSGEIPLELQNL- 570
Query: 339 KLRFLGLSDNNITGNLSP 356
KL L LS N+++G + P
Sbjct: 571 KLNVLNLSYNHLSGKIPP 588
Score = 99.8 bits (247), Expect = 1e-20
Identities = 81/250 (32%), Positives = 121/250 (48%), Gaps = 31/250 (12%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENG-------------------------LTGNIPQEIG- 190
L+G PS L +L SL L N L G+IP+ +
Sbjct: 77 LVGPFPSILCHLPSLHSLSLYNNSINGSLSADDFDTCHNLISLDLSENLLVGSIPKSLPF 136
Query: 191 NLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSH 250
NL LK L +SGNN S IP FG L L+L+ N LSGT+P +LG + ++ +L L++
Sbjct: 137 NLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAGNFLSGTIPASLGNVTTLKELKLAY 196
Query: 251 N-FLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLK 309
N F ++ ++ GNL L ++ L L + SL + SL + L+ N L G I +
Sbjct: 197 NLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLTFNQLTGSIPS-- 254
Query: 310 W-ENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALY 368
W L+ + +EL N GE+PES+ + L+ S N +TG + L L +L +L
Sbjct: 255 WITQLKTVEQIELFNNSFSGELPESMGNMTTLKRFDASMNKLTGKIPDNLNLL-NLESLN 313
Query: 369 LSGNNLKGEI 378
L N L+G +
Sbjct: 314 LFENMLEGPL 323
Score = 97.1 bits (240), Expect = 9e-20
Identities = 66/179 (36%), Positives = 97/179 (53%), Gaps = 2/179 (1%)
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
SN L G IP F L L L L +N TG+IP+ I L L +S N FSG+IP+
Sbjct: 411 SNNKLSGQIPHGFWGLPRLSLLELSDNSFTGSIPKTIIGAKNLSNLRISKNRFSGSIPNE 470
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDL 272
G L+ ++ + + N SG +P +L +L + +LDLS N L G++ E KNL ++L
Sbjct: 471 IGSLNGIIEISGAENDFSGEIPESLVKLKQLSRLDLSKNQLSGEIPRELRGWKNLNELNL 530
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
NN L + + + L + LS+N G+I L+ +NL+ L +L LS L G+IP
Sbjct: 531 ANNHLSGEIPKEVGILPVLNYLDLSSNQFSGEI-PLELQNLK-LNVLNLSYNHLSGKIP 587
Score = 90.9 bits (224), Expect = 6e-18
Identities = 73/230 (31%), Positives = 116/230 (49%), Gaps = 3/230 (1%)
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP-DIFGGLSDLLILDLSRNS 228
N+ S+ L L G P + +L L L L N+ +G++ D F +L+ LDLS N
Sbjct: 66 NVVSVDLSSFMLVGPFPSILCHLPSLHSLSLYNNSINGSLSADDFDTCHNLISLDLSENL 125
Query: 229 LSGTLPVTLGRLISVLK-LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQE 287
L G++P +L + LK L++S N L + + FG + L ++L N L + SL
Sbjct: 126 LVGSIPKSLPFNLPNLKFLEISGNNLSDTIPSSFGEFRKLESLNLAGNFLSGTIPASLGN 185
Query: 288 MNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSD 347
+ +L+E+ L+ N + NL L +L L+ L+G IP SLS+L L L L+
Sbjct: 186 VTTLKELKLAYNLFSPSQIPSQLGNLTELQVLWLAGCNLVGPIPPSLSRLTSLVNLDLTF 245
Query: 348 NNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSN 397
N +TG++ + L ++ + L N+ GE+ S G L +RF A N
Sbjct: 246 NQLTGSIPSWITQLKTVEQIELFNNSFSGELPESMGNMTTL-KRFDASMN 294
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor
(EC 2.7.1.37) (tBRI1) (Altered brassinolide sensitivity
1) (Systemin receptor SR160)
Length = 1207
Score = 117 bits (294), Expect = 5e-26
Identities = 85/244 (34%), Positives = 125/244 (50%), Gaps = 7/244 (2%)
Query: 140 EKLAE--SLESIEFRSNPGLIGNIP-STFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLK 196
E L E SLE ++ N G +P T L N++++VL N G +P NL+KL+
Sbjct: 346 ESLGECSSLELVDISYN-NFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLPDSFSNLLKLE 404
Query: 197 RLVLSGNNFSGNIPDIF--GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLE 254
L +S NN +G IP +++L +L L N G +P +L ++ LDLS N+L
Sbjct: 405 TLDMSSNNLTGVIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLT 464
Query: 255 GKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQ 314
G + + G+L L + L N+L + L + +LE ++L N L G I N
Sbjct: 465 GSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPA-SLSNCT 523
Query: 315 NLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNL 374
L + LSN +L GEIP SL +L L L L +N+I+GN+ +L SL L L+ N L
Sbjct: 524 KLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFL 583
Query: 375 KGEI 378
G I
Sbjct: 584 NGSI 587
Score = 108 bits (270), Expect = 3e-23
Identities = 71/230 (30%), Positives = 120/230 (51%), Gaps = 8/230 (3%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQE-IGNLVKLKRLVLSGNNFSGNIPDIFGGLS 217
G +P + G +L+ + + N +G +P + + L +K +VLS N F G +PD F L
Sbjct: 342 GMVPESLGECSSLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLPDSFSNLL 401
Query: 218 DLLILDLSRNSLSGTLPVTLGR--LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
L LD+S N+L+G +P + + + ++ L L +N +G + + N L +DL N
Sbjct: 402 KLETLDMSSNNLTGVIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFN 461
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDI--RTLKWENLQNLVILELSNMELIGEIPES 333
L + SL ++ L++++L N L G+I + + L+NL+ L +L G IP S
Sbjct: 462 YLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLI---LDFNDLTGPIPAS 518
Query: 334 LSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
LS KL ++ LS+N ++G + L L +L L L N++ G I G
Sbjct: 519 LSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELG 568
Score = 107 bits (268), Expect = 5e-23
Identities = 91/262 (34%), Positives = 124/262 (46%), Gaps = 36/262 (13%)
Query: 156 GLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVK-LKRLVLSGNNFSGNIPDIFG 214
GL+ +PS ++LQ L L N G P ++ +L K + L LS NNFSG +P+ G
Sbjct: 295 GLVPKLPS-----ESLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLG 349
Query: 215 GLSDLLILDLSRNSLSGTLPV-TLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLR 273
S L ++D+S N+ SG LPV TL +L ++ + LS N G L + F NL L +D+
Sbjct: 350 ECSSLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNKFVGGLPDSFSNLLKLETLDMS 409
Query: 274 NNRLCCGLVLS---LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEI 330
+N L G++ S MN+L+ + L NN G I N LV L+LS L G I
Sbjct: 410 SNNLT-GVIPSGICKDPMNNLKVLYLQNNLFKGPIPD-SLSNCSQLVSLDLSFNYLTGSI 467
Query: 331 PESLSQLKKLRFLGL------------------------SDNNITGNLSPKLETLPSLNA 366
P SL L KL+ L L N++TG + L LN
Sbjct: 468 PSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLNW 527
Query: 367 LYLSGNNLKGEIQFSKGFFGKL 388
+ LS N L GEI S G L
Sbjct: 528 ISLSNNQLSGEIPASLGRLSNL 549
Score = 107 bits (268), Expect = 5e-23
Identities = 109/342 (31%), Positives = 144/342 (41%), Gaps = 64/342 (18%)
Query: 119 SFFNCFQ------SPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKN 170
S NC Q S N L SIP+ G+ KL + + + L G IP L+
Sbjct: 446 SLSNCSQLVSLDLSFNYLTGSIPSSLGSLSKLKDLILWLN-----QLSGEIPQELMYLQA 500
Query: 171 LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLS 230
L++L+L N LTG IP + N KL + LS N SG IP G LS+L IL L NS+S
Sbjct: 501 LENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSIS 560
Query: 231 GTLPVTLGRLISVLKLDLSHNFLEGKL--------------------------------- 257
G +P LG S++ LDL+ NFL G +
Sbjct: 561 GNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSKECH 620
Query: 258 ----LNEFGNLKNLTLMDLRNNRLCCGLVL--------SLQEMNSLEEMVLSNNPLGGDI 305
L EFG ++ L D + R C + S+ + LS N L G I
Sbjct: 621 GAGNLLEFGGIRQEQL-DRISTRHPCNFTRVYRGITQPTFNHNGSMIFLDLSYNKLEGSI 679
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ + L IL L + +L G IP+ L LK + L LS N G + L +L L
Sbjct: 680 PK-ELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLG 738
Query: 366 ALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLC-YPFEL 406
+ LS NNL G I S F RF +N LC YP +
Sbjct: 739 EIDLSNNNLSGMIPESAPFDTFPDYRF---ANNSLCGYPLPI 777
Score = 85.9 bits (211), Expect = 2e-16
Identities = 82/296 (27%), Positives = 124/296 (41%), Gaps = 44/296 (14%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S N L IP+G + +L+ + ++N G IP + L SL L N LTG+I
Sbjct: 409 SSNNLTGVIPSGICKDPMNNLKVLYLQNNL-FKGPIPDSLSNCSQLVSLDLSFNYLTGSI 467
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P +G+L KLK L+L N SG IP L L L L N L+G +P +L +
Sbjct: 468 PSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLNW 527
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+ LS+N L G++ G L NL ++ L NN + + L SL + L+ N L G I
Sbjct: 528 ISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNGSI 587
Query: 306 RTLKWENLQNLVILELSN--------------------MELIGEIPESLSQLK------- 338
++ N+ + L+ +E G E L ++
Sbjct: 588 PPPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRHPCNF 647
Query: 339 ----------------KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ FL LS N + G++ +L + L+ L L N+L G I
Sbjct: 648 TRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMI 703
Score = 81.3 bits (199), Expect = 5e-15
Identities = 74/235 (31%), Positives = 107/235 (45%), Gaps = 10/235 (4%)
Query: 149 IEFRSNPG--LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFS 206
+EF S G L G+IP KNL L L N + P + L+ L LS N F
Sbjct: 214 LEFFSLKGNKLAGSIPELD--FKNLSYLDLSANNFSTVFPS-FKDCSNLQHLDLSSNKFY 270
Query: 207 GNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNL-K 265
G+I L L+L+ N G +P S+ L L N +G N+ +L K
Sbjct: 271 GDIGSSLSSCGKLSFLNLTNNQFVGLVPKLPSE--SLQYLYLRGNDFQGVYPNQLADLCK 328
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
+ +DL N + SL E +SLE + +S N G + L N+ + LS +
Sbjct: 329 TVVELDLSYNNFSGMVPESLGECSSLELVDISYNNFSGKLPVDTLSKLSNIKTMVLSFNK 388
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKL--ETLPSLNALYLSGNNLKGEI 378
+G +P+S S L KL L +S NN+TG + + + + +L LYL N KG I
Sbjct: 389 FVGGLPDSFSNLLKLETLDMSSNNLTGVIPSGICKDPMNNLKVLYLQNNLFKGPI 443
Score = 73.6 bits (179), Expect = 1e-12
Identities = 76/217 (35%), Positives = 108/217 (49%), Gaps = 14/217 (6%)
Query: 168 LKNLQSLVLLENGLTGNIPQEIGNL--VKLKRLVLSGNNFSGNIPDI--FGGLSDLLILD 223
L NL+SLVL L+G++ + V L + L+ N SG I DI FG S+L L+
Sbjct: 107 LSNLESLVLKNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLN 166
Query: 224 LSRNSLSGTLPVTL-GRLISVLKLDLSHNFLEGKLLNEFGN---LKNLTLMDLRNNRLCC 279
LS+N L L S+ LDLS+N + G L + + L L+ N+L
Sbjct: 167 LSKNFLDPPGKEMLKAATFSLQVLDLSYNNISGFNLFPWVSSMGFVELEFFSLKGNKL-A 225
Query: 280 GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKK 339
G + L + +L + LS N + K + NL L+LS+ + G+I SLS K
Sbjct: 226 GSIPEL-DFKNLSYLDLSANNFSTVFPSFK--DCSNLQHLDLSSNKFYGDIGSSLSSCGK 282
Query: 340 LRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKG 376
L FL L++N G L PKL + SL LYL GN+ +G
Sbjct: 283 LSFLNLTNNQFVG-LVPKLPS-ESLQYLYLRGNDFQG 317
Score = 70.5 bits (171), Expect = 9e-12
Identities = 75/263 (28%), Positives = 117/263 (43%), Gaps = 37/263 (14%)
Query: 145 SLESIEFRSNP--GLIGNIPSTFGVLKNLQSLVLLENGL---------TGNIPQEIGNL- 192
+L+SI+ N G I +I S+FGV NL+SL L +N L ++ +L
Sbjct: 135 TLDSIDLAENTISGPISDI-SSFGVCSNLKSLNLSKNFLDPPGKEMLKAATFSLQVLDLS 193
Query: 193 ------------------VKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLP 234
V+L+ L GN +G+IP++ +L LDLS N+ S P
Sbjct: 194 YNNISGFNLFPWVSSMGFVELEFFSLKGNKLAGSIPEL--DFKNLSYLDLSANNFSTVFP 251
Query: 235 VTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEM 294
+ ++ LDLS N G + + + L+ ++L NN+ GLV L SL+ +
Sbjct: 252 -SFKDCSNLQHLDLSSNKFYGDIGSSLSSCGKLSFLNLTNNQF-VGLVPKLPS-ESLQYL 308
Query: 295 VLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNL 354
L N G + + +V L+LS G +PESL + L + +S NN +G L
Sbjct: 309 YLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLGECSSLELVDISYNNFSGKL 368
Query: 355 S-PKLETLPSLNALYLSGNNLKG 376
L L ++ + LS N G
Sbjct: 369 PVDTLSKLSNIKTMVLSFNKFVG 391
Score = 51.2 bits (121), Expect = 6e-06
Identities = 57/181 (31%), Positives = 80/181 (43%), Gaps = 12/181 (6%)
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVL--KLDLSHNFLEGKL--LNEFGNLKNLTLMD 271
LS+L L L +LSG+L V +DL+ N + G + ++ FG NL ++
Sbjct: 107 LSNLESLVLKNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLN 166
Query: 272 LRNNRLCC-GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILE---LSNMELI 327
L N L G + SL+ + LS N + G W + V LE L +L
Sbjct: 167 LSKNFLDPPGKEMLKAATFSLQVLDLSYNNISG-FNLFPWVSSMGFVELEFFSLKGNKLA 225
Query: 328 GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGK 387
G IPE K L +L LS NN + + P + +L L LS N G+I S GK
Sbjct: 226 GSIPEL--DFKNLSYLDLSANNFS-TVFPSFKDCSNLQHLDLSSNKFYGDIGSSLSSCGK 282
Query: 388 L 388
L
Sbjct: 283 L 283
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 117 bits (292), Expect = 8e-26
Identities = 86/244 (35%), Positives = 125/244 (50%), Gaps = 7/244 (2%)
Query: 140 EKLAE--SLESIEFRSNPGLIGNIP-STFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLK 196
E L E SLE ++ SN G +P T L N++++VL N G +P NL KL+
Sbjct: 346 ESLGECSSLELVDI-SNNNFSGKLPVDTLLKLSNIKTMVLSFNKFVGGLPDSFSNLPKLE 404
Query: 197 RLVLSGNNFSGNIPDIF--GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLE 254
L +S NN +G IP +++L +L L N G +P +L ++ LDLS N+L
Sbjct: 405 TLDMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFNYLT 464
Query: 255 GKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQ 314
G + + G+L L + L N+L + L + +LE ++L N L G I N
Sbjct: 465 GSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPA-SLSNCT 523
Query: 315 NLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNL 374
L + LSN +L GEIP SL +L L L L +N+I+GN+ +L SL L L+ N L
Sbjct: 524 KLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFL 583
Query: 375 KGEI 378
G I
Sbjct: 584 NGSI 587
Score = 108 bits (270), Expect = 3e-23
Identities = 110/342 (32%), Positives = 144/342 (41%), Gaps = 64/342 (18%)
Query: 119 SFFNCFQ------SPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKN 170
S NC Q S N L SIP+ G+ KL + + + L G IP L+
Sbjct: 446 SLSNCSQLVSLDLSFNYLTGSIPSSLGSLSKLKDLILWLN-----QLSGEIPQELMYLQA 500
Query: 171 LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLS 230
L++L+L N LTG IP + N KL + LS N SG IP G LS+L IL L NS+S
Sbjct: 501 LENLILDFNDLTGPIPASLSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSIS 560
Query: 231 GTLPVTLGRLISVLKLDLSHNFLEGKL--------------------------------- 257
G +P LG S++ LDL+ NFL G +
Sbjct: 561 GNIPAELGNCQSLIWLDLNTNFLNGSIPPPLFKQSGNIAVALLTGKRYVYIKNDGSKECH 620
Query: 258 ----LNEFGNLKNLTLMDLRNNRLCCGLVL--------SLQEMNSLEEMVLSNNPLGGDI 305
L EFG ++ L D + R C + S+ + LS N L G I
Sbjct: 621 GAGNLLEFGGIRQEQL-DRISTRHPCNFTRVYRGITQPTFNHNGSMIFLDLSYNKLEGSI 679
Query: 306 RTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLN 365
+ + L IL L + +L G IP+ L LK + L LS N G + L +L L
Sbjct: 680 PK-ELGAMYYLSILNLGHNDLSGMIPQQLGGLKNVAILDLSYNRFNGTIPNSLTSLTLLG 738
Query: 366 ALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLC-YPFEL 406
+ LS NNL G I S F RF +N LC YP L
Sbjct: 739 EIDLSNNNLSGMIPESAPFDTFPDYRF---ANNSLCGYPLPL 777
Score = 107 bits (268), Expect = 5e-23
Identities = 71/230 (30%), Positives = 120/230 (51%), Gaps = 8/230 (3%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQE-IGNLVKLKRLVLSGNNFSGNIPDIFGGLS 217
G +P + G +L+ + + N +G +P + + L +K +VLS N F G +PD F L
Sbjct: 342 GMVPESLGECSSLELVDISNNNFSGKLPVDTLLKLSNIKTMVLSFNKFVGGLPDSFSNLP 401
Query: 218 DLLILDLSRNSLSGTLPVTLGR--LISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
L LD+S N+L+G +P + + + ++ L L +N +G + + N L +DL N
Sbjct: 402 KLETLDMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPIPDSLSNCSQLVSLDLSFN 461
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDI--RTLKWENLQNLVILELSNMELIGEIPES 333
L + SL ++ L++++L N L G+I + + L+NL+ L +L G IP S
Sbjct: 462 YLTGSIPSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLI---LDFNDLTGPIPAS 518
Query: 334 LSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKG 383
LS KL ++ LS+N ++G + L L +L L L N++ G I G
Sbjct: 519 LSNCTKLNWISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELG 568
Score = 107 bits (268), Expect = 5e-23
Identities = 91/262 (34%), Positives = 124/262 (46%), Gaps = 36/262 (13%)
Query: 156 GLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVK-LKRLVLSGNNFSGNIPDIFG 214
GL+ +PS ++LQ L L N G P ++ +L K + L LS NNFSG +P+ G
Sbjct: 295 GLVPKLPS-----ESLQYLYLRGNDFQGVYPNQLADLCKTVVELDLSYNNFSGMVPESLG 349
Query: 215 GLSDLLILDLSRNSLSGTLPV-TLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLR 273
S L ++D+S N+ SG LPV TL +L ++ + LS N G L + F NL L +D+
Sbjct: 350 ECSSLELVDISNNNFSGKLPVDTLLKLSNIKTMVLSFNKFVGGLPDSFSNLPKLETLDMS 409
Query: 274 NNRLCCGLVLS---LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEI 330
+N L G++ S MN+L+ + L NN G I N LV L+LS L G I
Sbjct: 410 SNNLT-GIIPSGICKDPMNNLKVLYLQNNLFKGPIPD-SLSNCSQLVSLDLSFNYLTGSI 467
Query: 331 PESLSQLKKLRFLGL------------------------SDNNITGNLSPKLETLPSLNA 366
P SL L KL+ L L N++TG + L LN
Sbjct: 468 PSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLNW 527
Query: 367 LYLSGNNLKGEIQFSKGFFGKL 388
+ LS N L GEI S G L
Sbjct: 528 ISLSNNQLSGEIPASLGRLSNL 549
Score = 85.9 bits (211), Expect = 2e-16
Identities = 82/296 (27%), Positives = 124/296 (41%), Gaps = 44/296 (14%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S N L IP+G + +L+ + ++N G IP + L SL L N LTG+I
Sbjct: 409 SSNNLTGIIPSGICKDPMNNLKVLYLQNNL-FKGPIPDSLSNCSQLVSLDLSFNYLTGSI 467
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P +G+L KLK L+L N SG IP L L L L N L+G +P +L +
Sbjct: 468 PSSLGSLSKLKDLILWLNQLSGEIPQELMYLQALENLILDFNDLTGPIPASLSNCTKLNW 527
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+ LS+N L G++ G L NL ++ L NN + + L SL + L+ N L G I
Sbjct: 528 ISLSNNQLSGEIPASLGRLSNLAILKLGNNSISGNIPAELGNCQSLIWLDLNTNFLNGSI 587
Query: 306 RTLKWENLQNLVILELSN--------------------MELIGEIPESLSQLK------- 338
++ N+ + L+ +E G E L ++
Sbjct: 588 PPPLFKQSGNIAVALLTGKRYVYIKNDGSKECHGAGNLLEFGGIRQEQLDRISTRHPCNF 647
Query: 339 ----------------KLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ FL LS N + G++ +L + L+ L L N+L G I
Sbjct: 648 TRVYRGITQPTFNHNGSMIFLDLSYNKLEGSIPKELGAMYYLSILNLGHNDLSGMI 703
Score = 83.6 bits (205), Expect = 1e-15
Identities = 75/235 (31%), Positives = 108/235 (45%), Gaps = 10/235 (4%)
Query: 149 IEFRSNPG--LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFS 206
+EF S G L G+IP KNL L L N + P + L+ L LS N F
Sbjct: 214 LEFFSIKGNKLAGSIPELD--FKNLSYLDLSANNFSTVFPS-FKDCSNLQHLDLSSNKFY 270
Query: 207 GNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNL-K 265
G+I L L+L+ N G +P S+ L L N +G N+ +L K
Sbjct: 271 GDIGSSLSSCGKLSFLNLTNNQFVGLVPKLPSE--SLQYLYLRGNDFQGVYPNQLADLCK 328
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
+ +DL N + SL E +SLE + +SNN G + L N+ + LS +
Sbjct: 329 TVVELDLSYNNFSGMVPESLGECSSLELVDISNNNFSGKLPVDTLLKLSNIKTMVLSFNK 388
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKL--ETLPSLNALYLSGNNLKGEI 378
+G +P+S S L KL L +S NN+TG + + + + +L LYL N KG I
Sbjct: 389 FVGGLPDSFSNLPKLETLDMSSNNLTGIIPSGICKDPMNNLKVLYLQNNLFKGPI 443
Score = 75.5 bits (184), Expect = 3e-13
Identities = 77/219 (35%), Positives = 112/219 (50%), Gaps = 18/219 (8%)
Query: 168 LKNLQSLVLLENGLTGNIPQEIGNL--VKLKRLVLSGNNFSGNIPDI--FGGLSDLLILD 223
L NL+SLVL L+G++ + V L + L+ N SG I DI FG S+L L+
Sbjct: 107 LSNLESLVLKNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLN 166
Query: 224 LSRNSLSGTLPVTL-GRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDL-----RNNRL 277
LS+N L L G S+ LDLS+N + G N F + ++ ++L + N+L
Sbjct: 167 LSKNFLDPPGKEMLKGATFSLQVLDLSYNNISG--FNLFPWVSSMGFVELEFFSIKGNKL 224
Query: 278 CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQL 337
G + L + +L + LS N + K + NL L+LS+ + G+I SLS
Sbjct: 225 -AGSIPEL-DFKNLSYLDLSANNFSTVFPSFK--DCSNLQHLDLSSNKFYGDIGSSLSSC 280
Query: 338 KKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKG 376
KL FL L++N G L PKL + SL LYL GN+ +G
Sbjct: 281 GKLSFLNLTNNQFVG-LVPKLPS-ESLQYLYLRGNDFQG 317
Score = 50.1 bits (118), Expect = 1e-05
Identities = 56/181 (30%), Positives = 81/181 (43%), Gaps = 12/181 (6%)
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVL--KLDLSHNFLEGKL--LNEFGNLKNLTLMD 271
LS+L L L +LSG+L V +DL+ N + G + ++ FG NL ++
Sbjct: 107 LSNLESLVLKNANLSGSLTSAAKSQCGVTLDSIDLAENTISGPISDISSFGVCSNLKSLN 166
Query: 272 LRNNRLCC-GLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME---LI 327
L N L G + SL+ + LS N + G W + V LE +++ L
Sbjct: 167 LSKNFLDPPGKEMLKGATFSLQVLDLSYNNISG-FNLFPWVSSMGFVELEFFSIKGNKLA 225
Query: 328 GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGK 387
G IPE K L +L LS NN + + P + +L L LS N G+I S GK
Sbjct: 226 GSIPEL--DFKNLSYLDLSANNFS-TVFPSFKDCSNLQHLDLSSNKFYGDIGSSLSSCGK 282
Query: 388 L 388
L
Sbjct: 283 L 283
>D100_ARATH (Q00874) DNA-damage-repair/toleration protein DRT100
precursor
Length = 372
Score = 113 bits (282), Expect = 1e-24
Identities = 79/221 (35%), Positives = 117/221 (52%), Gaps = 3/221 (1%)
Query: 136 TGNWEKLAESLESIEFRSNPG--LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLV 193
TG SL S+ G + G IP+ G L L L L EN ++G IP + +L+
Sbjct: 124 TGEIPPCITSLASLRILDLAGNKITGEIPAEIGKLSKLAVLNLAENQMSGEIPASLTSLI 183
Query: 194 KLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFL 253
+LK L L+ N +G IP FG L L + L RN L+G++P ++ + + LDLS N +
Sbjct: 184 ELKHLELTENGITGVIPADFGSLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHI 243
Query: 254 EGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENL 313
EG + GN+K L+L++L N L + SL + L+ LS N L G I + + +
Sbjct: 244 EGPIPEWMGNMKVLSLLNLDCNSLTGPIPGSLLSNSGLDVANLSRNALEGTIPDV-FGSK 302
Query: 314 QNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNL 354
LV L+LS+ L G IP+SLS K + L +S N + G +
Sbjct: 303 TYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRI 343
Score = 107 bits (267), Expect = 7e-23
Identities = 80/222 (36%), Positives = 116/222 (52%), Gaps = 4/222 (1%)
Query: 159 GNIPSTFGVLKNLQSLVLLE-NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLS 217
G+I L L SLVL + G+TG IP I +L L+ L L+GN +G IP G LS
Sbjct: 100 GSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGNKITGEIPAEIGKLS 159
Query: 218 DLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
L +L+L+ N +SG +P +L LI + L+L+ N + G + +FG+LK L+ + L N L
Sbjct: 160 KLAVLNLAENQMSGEIPASLTSLIELKHLELTENGITGVIPADFGSLKMLSRVLLGRNEL 219
Query: 278 CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKW-ENLQNLVILELSNMELIGEIPESLSQ 336
+ S+ M L ++ LS N + G I +W N++ L +L L L G IP SL
Sbjct: 220 TGSIPESISGMERLADLDLSKNHIEGPIP--EWMGNMKVLSLLNLDCNSLTGPIPGSLLS 277
Query: 337 LKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
L LS N + G + + L +L LS N+L G I
Sbjct: 278 NSGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRI 319
Score = 89.7 bits (221), Expect = 1e-17
Identities = 63/165 (38%), Positives = 91/165 (54%), Gaps = 7/165 (4%)
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
L+ +E N G+ G IP+ FG LK L ++L N LTG+IP+ I + +L L LS N+
Sbjct: 185 LKHLELTEN-GITGVIPADFGSLKMLSRVLLGRNELTGSIPESISGMERLADLDLSKNHI 243
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLD---LSHNFLEGKLLNEFG 262
G IP+ G + L +L+L NSL+G +P G L+S LD LS N LEG + + FG
Sbjct: 244 EGPIPEWMGNMKVLSLLNLDCNSLTGPIP---GSLLSNSGLDVANLSRNALEGTIPDVFG 300
Query: 263 NLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
+ L +DL +N L + SL + + +S+N L G I T
Sbjct: 301 SKTYLVSLDLSHNSLSGRIPDSLSSAKFVGHLDISHNKLCGRIPT 345
Score = 71.6 bits (174), Expect = 4e-12
Identities = 58/155 (37%), Positives = 78/155 (49%), Gaps = 8/155 (5%)
Query: 128 NKLPVSIPTG--NWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
N+L SIP E+LA+ ++ N + G IP G +K L L L N LTG I
Sbjct: 217 NELTGSIPESISGMERLAD----LDLSKNH-IEGPIPEWMGNMKVLSLLNLDCNSLTGPI 271
Query: 186 PQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLK 245
P + + L LS N G IPD+FG + L+ LDLS NSLSG +P +L V
Sbjct: 272 PGSLLSNSGLDVANLSRNALEGTIPDVFGSKTYLVSLDLSHNSLSGRIPDSLSSAKFVGH 331
Query: 246 LDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCG 280
LD+SHN L G++ F +L +N+ CG
Sbjct: 332 LDISHNKLCGRIPTGF-PFDHLEATSFSDNQCLCG 365
Score = 42.4 bits (98), Expect = 0.003
Identities = 22/56 (39%), Positives = 31/56 (55%)
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFS 381
+ GEIP ++ L LR L L+ N ITG + ++ L L L L+ N + GEI S
Sbjct: 123 ITGEIPPCITSLASLRILDLAGNKITGEIPAEIGKLSKLAVLNLAENQMSGEIPAS 178
Score = 37.0 bits (84), Expect = 0.11
Identities = 24/62 (38%), Positives = 32/62 (50%), Gaps = 1/62 (1%)
Query: 328 GEIPESLSQLKKLRFLGLSD-NNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFG 386
G I ++ L L L L+D ITG + P + +L SL L L+GN + GEI G
Sbjct: 100 GSIDPAVCDLTALTSLVLADWKGITGEIPPCITSLASLRILDLAGNKITGEIPAEIGKLS 159
Query: 387 KL 388
KL
Sbjct: 160 KL 161
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 109 bits (272), Expect = 2e-23
Identities = 91/231 (39%), Positives = 119/231 (51%), Gaps = 8/231 (3%)
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEI-GNLVKLKRLVLSGNNFSGNIPD 211
S+ +G IP LK+LQ L L EN TG IP + G L L LSGN+F G +P
Sbjct: 277 SSNQFVGPIPPL--PLKSLQYLSLAENKFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPP 334
Query: 212 IFGGLSDLLILDLSRNSLSGTLPV-TLGRLISVLKLDLSHNFLEGKLLNEFGNLK-NLTL 269
FG S L L LS N+ SG LP+ TL ++ + LDLS N G+L NL +L
Sbjct: 335 FFGSCSLLESLALSSNNFSGELPMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASLLT 394
Query: 270 MDLRNNRLCCGLVLSLQE--MNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELI 327
+DL +N ++ +L + N+L+E+ L NN G I N LV L LS L
Sbjct: 395 LDLSSNNFSGPILPNLCQNPKNTLQELYLQNNGFTGKIPPTL-SNCSELVSLHLSFNYLS 453
Query: 328 GEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
G IP SL L KLR L L N + G + +L + +L L L N+L GEI
Sbjct: 454 GTIPSSLGSLSKLRDLKLWLNMLEGEIPQELMYVKTLETLILDFNDLTGEI 504
Score = 98.2 bits (243), Expect = 4e-20
Identities = 76/243 (31%), Positives = 114/243 (46%), Gaps = 21/243 (8%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IP +K L++L+L N LTG IP + N L + LS N +G IP G L
Sbjct: 476 LEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNCTNLNWISLSNNRLTGEIPKWIGRL 535
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKL------------LNEFGNL 264
+L IL LS NS SG +P LG S++ LDL+ N G + N
Sbjct: 536 ENLAILKLSNNSFSGNIPAELGDCRSLIWLDLNTNLFNGTIPAAMFKQSGKIAANFIAGK 595
Query: 265 KNLTLMDLRNNRLC--CGLVLSLQEMNSLEEMVLSN-NPLG------GDIRTLKWENLQN 315
+ + + + + C G +L Q + S + LS NP G + ++N +
Sbjct: 596 RYVYIKNDGMKKECHGAGNLLEFQGIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGS 655
Query: 316 LVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLK 375
++ L++S L G IP+ + + L L L N+I+G++ ++ L LN L LS N L
Sbjct: 656 MMFLDMSYNMLSGYIPKEIGSMPYLFILNLGHNDISGSIPDEVGDLRGLNILDLSSNKLD 715
Query: 376 GEI 378
G I
Sbjct: 716 GRI 718
Score = 95.9 bits (237), Expect = 2e-19
Identities = 92/268 (34%), Positives = 121/268 (44%), Gaps = 29/268 (10%)
Query: 144 ESLESIEFRSNPGLIGNIPSTF-GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSG 202
+SL+ + N G IP G L L L N G +P G+ L+ L LS
Sbjct: 291 KSLQYLSLAENK-FTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSCSLLESLALSS 349
Query: 203 NNFSGNIP-DIFGGLSDLLILDLSRNSLSGTLPVTLGRL-ISVLKLDLSHNFLEGKLL-N 259
NNFSG +P D + L +LDLS N SG LP +L L S+L LDLS N G +L N
Sbjct: 350 NNFSGELPMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASLLTLDLSSNNFSGPILPN 409
Query: 260 EFGNLKN-LTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI----------RTL 308
N KN L + L+NN + +L + L + LS N L G I R L
Sbjct: 410 LCQNPKNTLQELYLQNNGFTGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDL 469
Query: 309 K-WENL------------QNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLS 355
K W N+ + L L L +L GEIP LS L ++ LS+N +TG +
Sbjct: 470 KLWLNMLEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNCTNLNWISLSNNRLTGEIP 529
Query: 356 PKLETLPSLNALYLSGNNLKGEIQFSKG 383
+ L +L L LS N+ G I G
Sbjct: 530 KWIGRLENLAILKLSNNSFSGNIPAELG 557
Score = 86.7 bits (213), Expect = 1e-16
Identities = 82/249 (32%), Positives = 111/249 (43%), Gaps = 36/249 (14%)
Query: 109 LFQLKHLKAISF-FNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGV 167
L +++ LK + FN F +LP S+ L+ SL +++ SN +P+
Sbjct: 361 LLKMRGLKVLDLSFNEFSG--ELPESLTN-----LSASLLTLDLSSNNFSGPILPNLCQN 413
Query: 168 LKN-LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSR 226
KN LQ L L NG TG IP + N +L L LS N SG IP G LS L L L
Sbjct: 414 PKNTLQELYLQNNGFTGKIPPTLSNCSELVSLHLSFNYLSGTIPSSLGSLSKLRDLKLWL 473
Query: 227 NSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQ 286
N L G +P L + ++ L L N L G++ + N NL + L NNRL
Sbjct: 474 NMLEGEIPQELMYVKTLETLILDFNDLTGEIPSGLSNCTNLNWISLSNNRLT-------- 525
Query: 287 EMNSLEEMVLSNNPLGGDIRTLKW-ENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
G+I KW L+NL IL+LSN G IP L + L +L L
Sbjct: 526 ----------------GEIP--KWIGRLENLAILKLSNNSFSGNIPAELGDCRSLIWLDL 567
Query: 346 SDNNITGNL 354
+ N G +
Sbjct: 568 NTNLFNGTI 576
Score = 82.4 bits (202), Expect = 2e-15
Identities = 67/213 (31%), Positives = 103/213 (47%), Gaps = 7/213 (3%)
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
NL+ L + N + IP +G+ L+ L +SGN SG+ ++L +L++S N
Sbjct: 223 NLEFLDVSSNNFSTGIPF-LGDCSALQHLDISGNKLSGDFSRAISTCTELKLLNISSNQF 281
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEF-GNLKNLTLMDLRNNRLCCGLVLSLQEM 288
G +P L S+ L L+ N G++ + G LT +DL N +
Sbjct: 282 VGPIPPL--PLKSLQYLSLAENKFTGEIPDFLSGACDTLTGLDLSGNHFYGAVPPFFGSC 339
Query: 289 NSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK-KLRFLGLSD 347
+ LE + LS+N G++ ++ L +L+LS E GE+PESL+ L L L LS
Sbjct: 340 SLLESLALSSNNFSGELPMDTLLKMRGLKVLDLSFNEFSGELPESLTNLSASLLTLDLSS 399
Query: 348 NNITGNLSPKLETLP--SLNALYLSGNNLKGEI 378
NN +G + P L P +L LYL N G+I
Sbjct: 400 NNFSGPILPNLCQNPKNTLQELYLQNNGFTGKI 432
Score = 72.8 bits (177), Expect = 2e-12
Identities = 75/268 (27%), Positives = 113/268 (41%), Gaps = 51/268 (19%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L G IPS NL + L N LTG IP+ IG L L L LS N+FSGNIP G
Sbjct: 500 LTGEIPSGLSNCTNLNWISLSNNRLTGEIPKWIGRLENLAILKLSNNSFSGNIPAELGDC 559
Query: 217 SDLLILD--------------------LSRNSLSGTLPV--------------------- 235
L+ LD ++ N ++G V
Sbjct: 560 RSLIWLDLNTNLFNGTIPAAMFKQSGKIAANFIAGKRYVYIKNDGMKKECHGAGNLLEFQ 619
Query: 236 -----TLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNS 290
L RL + +++ G F N ++ +D+ N L + + M
Sbjct: 620 GIRSEQLNRLSTRNPCNITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGSMPY 679
Query: 291 LEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNI 350
L + L +N + G I + +L+ L IL+LS+ +L G IP+++S L L + LS+NN+
Sbjct: 680 LFILNLGHNDISGSIPD-EVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDLSNNNL 738
Query: 351 TGNLSP--KLETLPSLNALYLSGNNLKG 376
+G + + ET P A +L+ L G
Sbjct: 739 SGPIPEMGQFETFPP--AKFLNNPGLCG 764
Score = 71.2 bits (173), Expect = 5e-12
Identities = 47/125 (37%), Positives = 65/125 (51%), Gaps = 1/125 (0%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+ TF ++ L + N L+G IP+EIG++ L L L N+ SG+IPD G L
Sbjct: 644 GHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGSMPYLFILNLGHNDISGSIPDEVGDLRG 703
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L ILDLS N L G +P + L + ++DLS+N L G + E G + NN
Sbjct: 704 LNILDLSSNKLDGRIPQAMSALTMLTEIDLSNNNLSGP-IPEMGQFETFPPAKFLNNPGL 762
Query: 279 CGLVL 283
CG L
Sbjct: 763 CGYPL 767
Score = 68.9 bits (167), Expect = 3e-11
Identities = 60/226 (26%), Positives = 94/226 (41%), Gaps = 49/226 (21%)
Query: 126 SPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNI 185
S N+L IP W E+L ++ SN GNIP+ G ++L L L N G I
Sbjct: 520 SNNRLTGEIP--KWIGRLENLAILKL-SNNSFSGNIPAELGDCRSLIWLDLNTNLFNGTI 576
Query: 186 PQEI--------GNLVKLKRLVLSGNN--------------------------------- 204
P + N + KR V N+
Sbjct: 577 PAAMFKQSGKIAANFIAGKRYVYIKNDGMKKECHGAGNLLEFQGIRSEQLNRLSTRNPCN 636
Query: 205 -----FSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLN 259
+ G+ F ++ LD+S N LSG +P +G + + L+L HN + G + +
Sbjct: 637 ITSRVYGGHTSPTFDNNGSMMFLDMSYNMLSGYIPKEIGSMPYLFILNLGHNDISGSIPD 696
Query: 260 EFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
E G+L+ L ++DL +N+L + ++ + L E+ LSNN L G I
Sbjct: 697 EVGDLRGLNILDLSSNKLDGRIPQAMSALTMLTEIDLSNNNLSGPI 742
Score = 64.3 bits (155), Expect = 6e-10
Identities = 61/196 (31%), Positives = 96/196 (48%), Gaps = 15/196 (7%)
Query: 191 NLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSG--TLPVTLGRLISVLKLDL 248
+L L+ L LS ++ +G++ F + L LDLSRNSLSG T +LG + L++
Sbjct: 97 SLTGLESLFLSNSHINGSVSG-FKCSASLTSLDLSRNSLSGPVTTLTSLGSCSGLKFLNV 155
Query: 249 SHNFLE--GKLLNEFGNLKNLTLMDLRNNRL----CCGLVLSLQEMNSLEEMVLSNNPLG 302
S N L+ GK+ L +L ++DL N + G VLS L+ + +S N +
Sbjct: 156 SSNTLDFPGKVSGGL-KLNSLEVLDLSANSISGANVVGWVLS-DGCGELKHLAISGNKIS 213
Query: 303 GDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLP 362
GD+ + NL+ L++S+ IP L L+ L +S N ++G+ S + T
Sbjct: 214 GDVDVSRCVNLE---FLDVSSNNFSTGIP-FLGDCSALQHLDISGNKLSGDFSRAISTCT 269
Query: 363 SLNALYLSGNNLKGEI 378
L L +S N G I
Sbjct: 270 ELKLLNISSNQFVGPI 285
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 101 bits (252), Expect = 4e-21
Identities = 92/322 (28%), Positives = 136/322 (41%), Gaps = 53/322 (16%)
Query: 133 SIPTGNWEKLAESLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNL 192
SIP G S+E + SN L G+IP L NL L L N L+G + ++G L
Sbjct: 197 SIPVGIGN--CSSVEYLGLASN-NLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKL 253
Query: 193 VKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNF 252
L RL +S N FSG IPD+F L+ L N +G +P +L S+ L L +N
Sbjct: 254 SNLGRLDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSLSNSRSISLLSLRNNT 313
Query: 253 LEGKLLNEFGNLKNLTLMDL-------------------------------------RNN 275
L G++ + NLT +DL +N
Sbjct: 314 LSGQIYLNCSAMTNLTSLDLASNSFSGSIPSNLPNCLRLKTINFAKIKFIAQIPESFKNF 373
Query: 276 RLCCGLVLS-------------LQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELS 322
+ L S LQ +L+ +VL+ N ++ ++ +NL +L ++
Sbjct: 374 QSLTSLSFSNSSIQNISSALEILQHCQNLKTLVLTLNFQKEELPSVPSLQFKNLKVLIIA 433
Query: 323 NMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
+ +L G +P+ LS L+ L LS N ++G + P L +L SL L LS N GEI S
Sbjct: 434 SCQLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNTFIGEIPHSL 493
Query: 383 GFFGKLGRRFGAWSNPKLCYPF 404
L + A P +PF
Sbjct: 494 TSLQSLVSKENAVEEPSPDFPF 515
Score = 100 bits (250), Expect = 6e-21
Identities = 70/223 (31%), Positives = 109/223 (48%), Gaps = 5/223 (2%)
Query: 159 GNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSD 218
G+IP G +++ L L N L+G+IPQE+ L L L L N SG + G LS+
Sbjct: 196 GSIPVGIGNCSSVEYLGLASNNLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLSN 255
Query: 219 LLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLC 278
L LD+S N SG +P L + N G++ N ++++L+ LRNN L
Sbjct: 256 LGRLDISSNKFSGKIPDVFLELNKLWYFSAQSNLFNGEMPRSLSNSRSISLLSLRNNTLS 315
Query: 279 CGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLK 338
+ L+ M +L + L++N G I + N L + + ++ I +IPES +
Sbjct: 316 GQIYLNCSAMTNLTSLDLASNSFSGSIPS-NLPNCLRLKTINFAKIKFIAQIPESFKNFQ 374
Query: 339 KLRFLGLSDNNITGNLSPKLETL---PSLNALYLSGNNLKGEI 378
L L S+++I N+S LE L +L L L+ N K E+
Sbjct: 375 SLTSLSFSNSSIQ-NISSALEILQHCQNLKTLVLTLNFQKEEL 416
Score = 85.5 bits (210), Expect = 3e-16
Identities = 68/245 (27%), Positives = 104/245 (41%), Gaps = 61/245 (24%)
Query: 169 KNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNS 228
KNL+ L++ L G +PQ + N L+ L LS N SG IP G L+ L LDLS N+
Sbjct: 425 KNLKVLIIASCQLRGTVPQWLSNSPSLQLLDLSWNQLSGTIPPWLGSLNSLFYLDLSNNT 484
Query: 229 LSGTLPVTLGRLISVLK------------------------------------LDLSHNF 252
G +P +L L S++ +DLS+N
Sbjct: 485 FIGEIPHSLTSLQSLVSKENAVEEPSPDFPFFKKKNTNAGGLQYNQPSSFPPMIDLSYNS 544
Query: 253 LEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWEN 312
L G + EFG+L+ L +++L+NN L + +L M SLE
Sbjct: 545 LNGSIWPEFGDLRQLHVLNLKNNNLSGNIPANLSGMTSLE-------------------- 584
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
+L+LS+ L G IP SL +L L ++ N ++G + ++ N+ +
Sbjct: 585 -----VLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKLSGPIPTGVQFQTFPNSSFEGNQ 639
Query: 373 NLKGE 377
L GE
Sbjct: 640 GLCGE 644
Score = 84.0 bits (206), Expect = 8e-16
Identities = 54/169 (31%), Positives = 84/169 (48%), Gaps = 1/169 (0%)
Query: 222 LDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGL 281
L+L R LSG L ++ +L + L+L+HN L G + NL NL ++DL +N GL
Sbjct: 91 LELGRRKLSGKLSESVAKLDQLKVLNLTHNSLSGSIAASLLNLSNLEVLDLSSNDFS-GL 149
Query: 282 VLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLR 341
SL + SL + + N G I NL + ++L+ G IP + +
Sbjct: 150 FPSLINLPSLRVLNVYENSFHGLIPASLCNNLPRIREIDLAMNYFDGSIPVGIGNCSSVE 209
Query: 342 FLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGR 390
+LGL+ NN++G++ +L L +L+ L L N L G + G LGR
Sbjct: 210 YLGLASNNLSGSIPQELFQLSNLSVLALQNNRLSGALSSKLGKLSNLGR 258
Score = 64.3 bits (155), Expect = 6e-10
Identities = 40/102 (39%), Positives = 58/102 (56%), Gaps = 5/102 (4%)
Query: 156 GLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG 215
GL N PS+F + +L N L G+I E G+L +L L L NN SGNIP G
Sbjct: 525 GLQYNQPSSFPPMIDLSY-----NSLNGSIWPEFGDLRQLHVLNLKNNNLSGNIPANLSG 579
Query: 216 LSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKL 257
++ L +LDLS N+LSG +P +L +L + +++N L G +
Sbjct: 580 MTSLEVLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKLSGPI 621
Score = 63.2 bits (152), Expect = 1e-09
Identities = 66/222 (29%), Positives = 100/222 (44%), Gaps = 27/222 (12%)
Query: 101 EKLEFKPELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGN 160
E+L P L Q K+LK + +C +L ++P W + SL+ ++ N L G
Sbjct: 414 EELPSVPSL-QFKNLKVLIIASC-----QLRGTVP--QWLSNSPSLQLLDLSWNQ-LSGT 464
Query: 161 IPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI-------- 212
IP G L +L L L N G IP +L L+ LV N PD
Sbjct: 465 IPPWLGSLNSLFYLDLSNNTFIGEIPH---SLTSLQSLVSKENAVEEPSPDFPFFKKKNT 521
Query: 213 -FGGL------SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLK 265
GGL S ++DLS NSL+G++ G L + L+L +N L G + +
Sbjct: 522 NAGGLQYNQPSSFPPMIDLSYNSLNGSIWPEFGDLRQLHVLNLKNNNLSGNIPANLSGMT 581
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRT 307
+L ++DL +N L + SL +++ L ++ N L G I T
Sbjct: 582 SLEVLDLSHNNLSGNIPPSLVKLSFLSTFSVAYNKLSGPIPT 623
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 97.1 bits (240), Expect = 9e-20
Identities = 89/298 (29%), Positives = 128/298 (42%), Gaps = 50/298 (16%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF--- 213
L GNIP LK L L + EN L+G++ +EI NL L RL +S N FSG IPD+F
Sbjct: 208 LTGNIPEDLFHLKRLNLLGIQENRLSGSLSREIRNLSSLVRLDVSWNLFSGEIPDVFDEL 267
Query: 214 --------------GGL-------SDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNF 252
GG+ L +L+L NSLSG L + +I++ LDL N
Sbjct: 268 PQLKFFLGQTNGFIGGIPKSLANSPSLNLLNLRNNSLSGRLMLNCTAMIALNSLDLGTNR 327
Query: 253 LEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGD-------- 304
G+L + K L ++L N + S + SL LSN+ L
Sbjct: 328 FNGRLPENLPDCKRLKNVNLARNTFHGQVPESFKNFESLSYFSLSNSSLANISSALGILQ 387
Query: 305 --------IRTLKWE----------NLQNLVILELSNMELIGEIPESLSQLKKLRFLGLS 346
+ TL + + + L +L ++N L G +P LS +L+ L LS
Sbjct: 388 HCKNLTTLVLTLNFHGEALPDDSSLHFEKLKVLVVANCRLTGSMPRWLSSSNELQLLDLS 447
Query: 347 DNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLCYPF 404
N +TG + + +L L LS N+ GEI S L R + + P +PF
Sbjct: 448 WNRLTGAIPSWIGDFKALFYLDLSNNSFTGEIPKSLTKLESLTSRNISVNEPSPDFPF 505
Score = 91.7 bits (226), Expect = 4e-18
Identities = 89/329 (27%), Positives = 144/329 (43%), Gaps = 21/329 (6%)
Query: 48 GFVGNSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGP-IHENSLPCANEKLEFK 106
G NS N + G+ + G + L + V+N+ ++S+P +
Sbjct: 67 GITCNSNNTGRVIRLELGNKKLSGKLSESLGKLDEIRVLNLSRNFIKDSIPLS------- 119
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNPGLIGNIPSTF- 165
+F LK+L+ + S N L IPT +L+S + SN G++PS
Sbjct: 120 --IFNLKNLQTLDL-----SSNDLSGGIPTSI---NLPALQSFDLSSNK-FNGSLPSHIC 168
Query: 166 GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLS 225
++ + L N GN G V L+ L L N+ +GNIP+ L L +L +
Sbjct: 169 HNSTQIRVVKLAVNYFAGNFTSGFGKCVLLEHLCLGMNDLTGNIPEDLFHLKRLNLLGIQ 228
Query: 226 RNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL 285
N LSG+L + L S+++LD+S N G++ + F L L + N G+ SL
Sbjct: 229 ENRLSGSLSREIRNLSSLVRLDVSWNLFSGEIPDVFDELPQLKFFLGQTNGFIGGIPKSL 288
Query: 286 QEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGL 345
SL + L NN L G + L + L L+L G +PE+L K+L+ + L
Sbjct: 289 ANSPSLNLLNLRNNSLSGRL-MLNCTAMIALNSLDLGTNRFNGRLPENLPDCKRLKNVNL 347
Query: 346 SDNNITGNLSPKLETLPSLNALYLSGNNL 374
+ N G + + SL+ LS ++L
Sbjct: 348 ARNTFHGQVPESFKNFESLSYFSLSNSSL 376
Score = 88.6 bits (218), Expect = 3e-17
Identities = 79/242 (32%), Positives = 105/242 (42%), Gaps = 42/242 (17%)
Query: 153 SNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
+N L G++P LQ L L N LTG IP IG+ L L LS N+F+G IP
Sbjct: 423 ANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSFTGEIPKS 482
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKL------------DLSHNFLEGKLLNE 260
L L ++S N S P + R S L +L HN L G + E
Sbjct: 483 LTKLESLTSRNISVNEPSPDFPFFMKRNESARALQYNQIFGFPPTIELGHNNLSGPIWEE 542
Query: 261 FGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILE 320
FGNLK L + DL+ N L + SL M SLE + LSNN
Sbjct: 543 FGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNN--------------------- 581
Query: 321 LSNMELIGEIPESLSQLKKLRFLGLSDNNITGNL--SPKLETLPSLNALYLSGNNLKGEI 378
L G IP SL QL L ++ NN++G + + +T P+ + N+L GE
Sbjct: 582 ----RLSGSIPVSLQQLSFLSKFSVAYNNLSGVIPSGGQFQTFPNSS---FESNHLCGEH 634
Query: 379 QF 380
+F
Sbjct: 635 RF 636
Score = 85.5 bits (210), Expect = 3e-16
Identities = 76/259 (29%), Positives = 120/259 (45%), Gaps = 18/259 (6%)
Query: 144 ESLESIEFRS-NPGLIGNIPSTFGVL---KNLQSLVLLENGLTGNIPQEIG-NLVKLKRL 198
++ ES+ + S + + NI S G+L KNL +LVL N +P + + KLK L
Sbjct: 361 KNFESLSYFSLSNSSLANISSALGILQHCKNLTTLVLTLNFHGEALPDDSSLHFEKLKVL 420
Query: 199 VLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLL 258
V++ +G++P ++L +LDLS N L+G +P +G ++ LDLS+N G++
Sbjct: 421 VVANCRLTGSMPRWLSSSNELQLLDLSWNRLTGAIPSWIGDFKALFYLDLSNNSFTGEIP 480
Query: 259 NEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVI 318
L++LT ++ N ++ S L N + G T+
Sbjct: 481 KSLTKLESLTSRNISVNEPSPDFPFFMKRNESAR--ALQYNQIFGFPPTI---------- 528
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
EL + L G I E LKKL L N ++G++ L + SL AL LS N L G I
Sbjct: 529 -ELGHNNLSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGSI 587
Query: 379 QFSKGFFGKLGRRFGAWSN 397
S L + A++N
Sbjct: 588 PVSLQQLSFLSKFSVAYNN 606
Score = 81.6 bits (200), Expect = 4e-15
Identities = 60/192 (31%), Positives = 96/192 (49%), Gaps = 9/192 (4%)
Query: 191 NLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSH 250
N ++ RL L SG + + G L ++ +L+LSRN + ++P+++ L ++ LDLS
Sbjct: 74 NTGRVIRLELGNKKLSGKLSESLGKLDEIRVLNLSRNFIKDSIPLSIFNLKNLQTLDLSS 133
Query: 251 NFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL-QEMNSLEEMVLSNNPLGGDIRTLK 309
N L G + NL L DL +N+ L + + + L+ N G+ +
Sbjct: 134 NDLSGGIPTSI-NLPALQSFDLSSNKFNGSLPSHICHNSTQIRVVKLAVNYFAGNFTS-- 190
Query: 310 WENLQNLVILE---LSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNA 366
V+LE L +L G IPE L LK+L LG+ +N ++G+LS ++ L SL
Sbjct: 191 --GFGKCVLLEHLCLGMNDLTGNIPEDLFHLKRLNLLGIQENRLSGSLSREIRNLSSLVR 248
Query: 367 LYLSGNNLKGEI 378
L +S N GEI
Sbjct: 249 LDVSWNLFSGEI 260
Score = 66.6 bits (161), Expect = 1e-10
Identities = 45/133 (33%), Positives = 65/133 (48%), Gaps = 1/133 (0%)
Query: 173 SLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGT 232
++ L N L+G I +E GNL KL L N SG+IP G++ L LDLS N LSG+
Sbjct: 527 TIELGHNNLSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSGS 586
Query: 233 LPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLE 292
+PV+L +L + K +++N L G ++ G + +N LC E
Sbjct: 587 IPVSLQQLSFLSKFSVAYNNLSG-VIPSGGQFQTFPNSSFESNHLCGEHRFPCSEGTESA 645
Query: 293 EMVLSNNPLGGDI 305
+ S GGDI
Sbjct: 646 LIKRSRRSRGGDI 658
Score = 51.2 bits (121), Expect = 6e-06
Identities = 34/87 (39%), Positives = 45/87 (51%), Gaps = 1/87 (1%)
Query: 148 SIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSG 207
+IE N L G I FG LK L L N L+G+IP + + L+ L LS N SG
Sbjct: 527 TIELGHN-NLSGPIWEEFGNLKKLHVFDLKWNALSGSIPSSLSGMTSLEALDLSNNRLSG 585
Query: 208 NIPDIFGGLSDLLILDLSRNSLSGTLP 234
+IP LS L ++ N+LSG +P
Sbjct: 586 SIPVSLQQLSFLSKFSVAYNNLSGVIP 612
Score = 46.2 bits (108), Expect = 2e-04
Identities = 28/75 (37%), Positives = 41/75 (54%)
Query: 307 TLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNA 366
T N ++ LEL N +L G++ ESL +L ++R L LS N I ++ + L +L
Sbjct: 69 TCNSNNTGRVIRLELGNKKLSGKLSESLGKLDEIRVLNLSRNFIKDSIPLSIFNLKNLQT 128
Query: 367 LYLSGNNLKGEIQFS 381
L LS N+L G I S
Sbjct: 129 LDLSSNDLSGGIPTS 143
>PGI1_ARATH (Q9M5J9) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 330
Score = 96.3 bits (238), Expect = 2e-19
Identities = 77/221 (34%), Positives = 112/221 (49%), Gaps = 7/221 (3%)
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
LE++ FR L G I T LKNL+ L L LTG IP I L L+ L LS N+
Sbjct: 96 LETLVFRKLSNLTGTIQPTIAKLKNLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDL 155
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLI-SVLKLDLSHNFLEGKLLNEFGNL 264
SG+IP L +L L+LSRN L+G++P + G +V L LSHN L G + GN+
Sbjct: 156 SGSIPSSLSTLPKILALELSRNKLTGSIPESFGSFPGTVPDLRLSHNQLSGPIPKSLGNI 215
Query: 265 KNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNM 324
+ +DL N+L + + + LS N DI K + + L IL+L++
Sbjct: 216 -DFNRIDLSRNKLQGDASMLFGSNKTTWSIDLSRNMFQFDIS--KVDIPKTLGILDLNHN 272
Query: 325 ELIGEIPESLSQLKKLRFLGLSDNNITGNL--SPKLETLPS 363
+ G IP ++ L+F +S N + G++ KL+T S
Sbjct: 273 GITGNIPVQWTE-APLQFFNVSYNKLCGHIPTGGKLQTFDS 312
Score = 75.5 bits (184), Expect = 3e-13
Identities = 74/233 (31%), Positives = 106/233 (44%), Gaps = 32/233 (13%)
Query: 173 SLVLLENGLTGNIPQEIGNLVKLKRLV-------------------------LSGNNFSG 207
+L + ++G IP E+G+L L+ LV LS N +G
Sbjct: 74 ALTIFSGQISGQIPAEVGDLPYLETLVFRKLSNLTGTIQPTIAKLKNLRMLRLSWTNLTG 133
Query: 208 NIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL 267
IPD L +L L+LS N LSG++P +L L +L L+LS N L G + FG+
Sbjct: 134 PIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRNKLTGSIPESFGSFPG- 192
Query: 268 TLMDLR--NNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
T+ DLR +N+L + SL ++ + LS N L GD L N + ++LS
Sbjct: 193 TVPDLRLSHNQLSGPIPKSLGNID-FNRIDLSRNKLQGDASMLFGSN-KTTWSIDLSRNM 250
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+I + K L L L+ N ITGN+ P T L +S N L G I
Sbjct: 251 FQFDI-SKVDIPKTLGILDLNHNGITGNI-PVQWTEAPLQFFNVSYNKLCGHI 301
Score = 62.4 bits (150), Expect = 2e-09
Identities = 36/73 (49%), Positives = 46/73 (62%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L+NL +L LS L G IP+ +SQLK L FL LS N+++G++ L TLP + AL LS N
Sbjct: 118 LKNLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFNDLSGSIPSSLSTLPKILALELSRN 177
Query: 373 NLKGEIQFSKGFF 385
L G I S G F
Sbjct: 178 KLTGSIPESFGSF 190
Score = 44.7 bits (104), Expect = 5e-04
Identities = 30/76 (39%), Positives = 42/76 (54%), Gaps = 2/76 (2%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L+ LV +LSN L G I ++++LK LR L LS N+TG + + L +L L LS N
Sbjct: 96 LETLVFRKLSN--LTGTIQPTIAKLKNLRMLRLSWTNLTGPIPDFISQLKNLEFLELSFN 153
Query: 373 NLKGEIQFSKGFFGKL 388
+L G I S K+
Sbjct: 154 DLSGSIPSSLSTLPKI 169
>PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor
(Polygalacturonase-inhibiting protein)
Length = 330
Score = 95.9 bits (237), Expect = 2e-19
Identities = 71/212 (33%), Positives = 109/212 (50%), Gaps = 9/212 (4%)
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
LE++EF P L G I LK L+SL L L+G++P + L L L LS NN
Sbjct: 96 LETLEFHKQPNLTGPIQPAIAKLKGLKSLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNL 155
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLI-SVLKLDLSHNFLEGKLLNEFGNL 264
+G IP L +L L L RN L+G +P++ G+ I +V L LSHN L G + F +
Sbjct: 156 TGAIPSSLSELPNLGALRLDRNKLTGHIPISFGQFIGNVPDLYLSHNQLSGNIPTSFAQM 215
Query: 265 KNLTLMDLRNNRL--CCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELS 322
+ T +DL N+L ++ L + + + LS N L + K E +L L+++
Sbjct: 216 -DFTSIDLSRNKLEGDASVIFGLNKTTQIVD--LSRNLL--EFNLSKVEFPTSLTSLDIN 270
Query: 323 NMELIGEIPESLSQLKKLRFLGLSDNNITGNL 354
+ ++ G IP +QL +FL +S N + G +
Sbjct: 271 HNKIYGSIPVEFTQL-NFQFLNVSYNRLCGQI 301
Score = 75.5 bits (184), Expect = 3e-13
Identities = 70/233 (30%), Positives = 114/233 (48%), Gaps = 8/233 (3%)
Query: 171 LQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGN-NFSGNIPDIFGGLSDLLILDLSRNSL 229
+ SL + ++G IP +G+L L+ L N +G I L L L LS +L
Sbjct: 72 INSLTIFAGQVSGQIPALVGDLPYLETLEFHKQPNLTGPIQPAIAKLKGLKSLRLSWTNL 131
Query: 230 SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSL-QEM 288
SG++P L +L ++ LDLS N L G + + L NL + L N+L + +S Q +
Sbjct: 132 SGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSELPNLGALRLDRNKLTGHIPISFGQFI 191
Query: 289 NSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDN 348
++ ++ LS+N L G+I T + + ++LS +L G+ K + + LS N
Sbjct: 192 GNVPDLYLSHNQLSGNIPTSFAQ--MDFTSIDLSRNKLEGDASVIFGLNKTTQIVDLSRN 249
Query: 349 NITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKLGRRFGAWSNPKLC 401
+ NLS K+E SL +L ++ N + G I F +L +F S +LC
Sbjct: 250 LLEFNLS-KVEFPTSLTSLDINHNKIYGSIPVE---FTQLNFQFLNVSYNRLC 298
>PGI3_PHAVU (P58823) Polygalacturonase inhibitor 3 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2) (PGIP-3)
Length = 342
Score = 91.3 bits (225), Expect = 5e-18
Identities = 73/227 (32%), Positives = 112/227 (49%), Gaps = 11/227 (4%)
Query: 160 NIPSTFGVLKNLQSLVLLE-------NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
N+P + + +L +L L N L G IP I L +L L ++ N SG IPD
Sbjct: 90 NLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDF 149
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL-TLMD 271
+ L+ LD S N+LSGTLP ++ L +++ + N + G + + +G+ L T M
Sbjct: 150 LSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGNRISGAIPDSYGSFSKLFTSMT 209
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
+ NRL + + +N L + LS N L GD L + + +N + L+ L ++
Sbjct: 210 ISRNRLTGKIPPTFANLN-LAFVDLSRNMLQGDASVL-FGSDKNTQKIHLAKNSLDFDL- 266
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
E + K L L L +N I G L L L L++L +S NNL GEI
Sbjct: 267 EKVGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEI 313
Score = 82.8 bits (203), Expect = 2e-15
Identities = 59/199 (29%), Positives = 94/199 (46%), Gaps = 4/199 (2%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L+G IP L L L + ++G IP + + L L S N SG +P L
Sbjct: 118 LVGPIPPAIAKLTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSL 177
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVL-KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
+L+ + N +SG +P + G + + +S N L GK+ F NL NL +DL N
Sbjct: 178 PNLVGITFDGNRISGAIPDSYGSFSKLFTSMTISRNRLTGKIPPTFANL-NLAFVDLSRN 236
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLS 335
L + + +++ L+ N L D+ + +NL L+L N + G +P+ L+
Sbjct: 237 MLQGDASVLFGSDKNTQKIHLAKNSLDFDLEKVGLS--KNLNGLDLRNNRIYGTLPQGLT 294
Query: 336 QLKKLRFLGLSDNNITGNL 354
QLK L L +S NN+ G +
Sbjct: 295 QLKFLHSLNVSFNNLCGEI 313
Score = 52.4 bits (124), Expect = 3e-06
Identities = 29/76 (38%), Positives = 42/76 (55%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L L L +++ + G IP+ LSQ+K L L S N ++G L P + +LP+L + GN
Sbjct: 129 LTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGN 188
Query: 373 NLKGEIQFSKGFFGKL 388
+ G I S G F KL
Sbjct: 189 RISGAIPDSYGSFSKL 204
Score = 34.3 bits (77), Expect = 0.72
Identities = 19/45 (42%), Positives = 24/45 (53%)
Query: 166 GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
G+ KNL L L N + G +PQ + L L L +S NN G IP
Sbjct: 270 GLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEIP 314
>PGI2_PHAVU (P58822) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 342
Score = 90.9 bits (224), Expect = 6e-18
Identities = 74/229 (32%), Positives = 112/229 (48%), Gaps = 15/229 (6%)
Query: 160 NIPSTFGVLKNLQSLVLLE-------NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
N+P + + +L +L L N L G IP I L +L L ++ N SG IPD
Sbjct: 90 NLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDF 149
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL-TLMD 271
+ L+ LD S N+LSGTLP ++ L +++ + N + G + + +G+ L T M
Sbjct: 150 LSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGNRISGAIPDSYGSFSKLFTSMT 209
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTL--KWENLQNLVILELSNMELIGE 329
+ NRL + + +N L + LS N L GD L +N Q + + + S +G+
Sbjct: 210 ISRNRLTGKIPPTFANLN-LAFVDLSRNMLEGDASVLFGSDKNTQKIHLAKNSLAFDLGK 268
Query: 330 IPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ S K L L L +N I G L L L L++L +S NNL GEI
Sbjct: 269 VGLS----KNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEI 313
Score = 82.0 bits (201), Expect = 3e-15
Identities = 59/199 (29%), Positives = 94/199 (46%), Gaps = 4/199 (2%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L+G IP L L L + ++G IP + + L L S N SG +P L
Sbjct: 118 LVGPIPPAIAKLTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSL 177
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVL-KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
+L+ + N +SG +P + G + + +S N L GK+ F NL NL +DL N
Sbjct: 178 PNLVGITFDGNRISGAIPDSYGSFSKLFTSMTISRNRLTGKIPPTFANL-NLAFVDLSRN 236
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLS 335
L + + +++ L+ N L D+ + +NL L+L N + G +P+ L+
Sbjct: 237 MLEGDASVLFGSDKNTQKIHLAKNSLAFDLGKVGLS--KNLNGLDLRNNRIYGTLPQGLT 294
Query: 336 QLKKLRFLGLSDNNITGNL 354
QLK L L +S NN+ G +
Sbjct: 295 QLKFLHSLNVSFNNLCGEI 313
Score = 52.4 bits (124), Expect = 3e-06
Identities = 29/76 (38%), Positives = 42/76 (55%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L L L +++ + G IP+ LSQ+K L L S N ++G L P + +LP+L + GN
Sbjct: 129 LTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLVGITFDGN 188
Query: 373 NLKGEIQFSKGFFGKL 388
+ G I S G F KL
Sbjct: 189 RISGAIPDSYGSFSKL 204
Score = 34.3 bits (77), Expect = 0.72
Identities = 19/45 (42%), Positives = 24/45 (53%)
Query: 166 GVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIP 210
G+ KNL L L N + G +PQ + L L L +S NN G IP
Sbjct: 270 GLSKNLNGLDLRNNRIYGTLPQGLTQLKFLHSLNVSFNNLCGEIP 314
>PGI1_PHAVU (P35334) Polygalacturonase inhibitor 1 precursor
(Polygalacturonase-inhibiting protein) (PGIP-1)
Length = 342
Score = 87.8 bits (216), Expect = 5e-17
Identities = 72/227 (31%), Positives = 110/227 (47%), Gaps = 11/227 (4%)
Query: 160 NIPSTFGVLKNLQSLVLLE-------NGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDI 212
N+P + + +L +L L N L G IP I L +L L ++ N SG IPD
Sbjct: 90 NLPKPYPIPSSLANLPYLNFLYIGGINNLVGPIPPAIAKLTQLHYLYITHTNVSGAIPDF 149
Query: 213 FGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNL-TLMD 271
+ L+ LD S N+LSGTLP ++ L ++ + N + G + + +G+ L T M
Sbjct: 150 LSQIKTLVTLDFSYNALSGTLPPSISSLPNLGGITFDGNRISGAIPDSYGSFSKLFTAMT 209
Query: 272 LRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIP 331
+ NRL + + +N L + LS N L GD L + + +N + L+ L ++
Sbjct: 210 ISRNRLTGKIPPTFANLN-LAFVDLSRNMLEGDASVL-FGSDKNTKKIHLAKNSLAFDLG 267
Query: 332 ESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+ + K L L L +N I G L L L L +L +S NNL GEI
Sbjct: 268 K-VGLSKNLNGLDLRNNRIYGTLPQGLTQLKFLQSLNVSFNNLCGEI 313
Score = 79.7 bits (195), Expect = 1e-14
Identities = 59/199 (29%), Positives = 94/199 (46%), Gaps = 4/199 (2%)
Query: 157 LIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGL 216
L+G IP L L L + ++G IP + + L L S N SG +P L
Sbjct: 118 LVGPIPPAIAKLTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSL 177
Query: 217 SDLLILDLSRNSLSGTLPVTLGRLISVL-KLDLSHNFLEGKLLNEFGNLKNLTLMDLRNN 275
+L + N +SG +P + G + + +S N L GK+ F NL NL +DL N
Sbjct: 178 PNLGGITFDGNRISGAIPDSYGSFSKLFTAMTISRNRLTGKIPPTFANL-NLAFVDLSRN 236
Query: 276 RLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLS 335
L + + +++ L+ N L D+ + +NL L+L N + G +P+ L+
Sbjct: 237 MLEGDASVLFGSDKNTKKIHLAKNSLAFDLGKVGLS--KNLNGLDLRNNRIYGTLPQGLT 294
Query: 336 QLKKLRFLGLSDNNITGNL 354
QLK L+ L +S NN+ G +
Sbjct: 295 QLKFLQSLNVSFNNLCGEI 313
Score = 55.8 bits (133), Expect = 2e-07
Identities = 56/177 (31%), Positives = 88/177 (49%), Gaps = 16/177 (9%)
Query: 107 PELFQLKHLKAISFFNCFQSPNKLPVSIPT--GNWEKLAESLESIEFRSNPGLIGNIPST 164
P + L +L I+F N++ +IP G++ KL ++ R L G IP T
Sbjct: 172 PSISSLPNLGGITF-----DGNRISGAIPDSYGSFSKLFTAMTISRNR----LTGKIPPT 222
Query: 165 FGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLS-DLLILD 223
F L NL + L N L G+ G+ K++ L+ N+ + ++ + GLS +L LD
Sbjct: 223 FANL-NLAFVDLSRNMLEGDASVLFGSDKNTKKIHLAKNSLAFDLGKV--GLSKNLNGLD 279
Query: 224 LSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCG 280
L N + GTLP L +L + L++S N L G++ + GNLK + NN+ CG
Sbjct: 280 LRNNRIYGTLPQGLTQLKFLQSLNVSFNNLCGEI-PQGGNLKRFDVSSYANNKCLCG 335
Score = 53.5 bits (127), Expect = 1e-06
Identities = 29/76 (38%), Positives = 42/76 (55%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L L L +++ + G IP+ LSQ+K L L S N ++G L P + +LP+L + GN
Sbjct: 129 LTQLHYLYITHTNVSGAIPDFLSQIKTLVTLDFSYNALSGTLPPSISSLPNLGGITFDGN 188
Query: 373 NLKGEIQFSKGFFGKL 388
+ G I S G F KL
Sbjct: 189 RISGAIPDSYGSFSKL 204
>PGI2_ARATH (Q9M5J8) Polygalacturonase inhibitor 2 precursor
(Polygalacturonase-inhibiting protein) (PGIP-2)
Length = 330
Score = 81.6 bits (200), Expect = 4e-15
Identities = 69/215 (32%), Positives = 102/215 (47%), Gaps = 6/215 (2%)
Query: 146 LESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNF 205
L S+ FR L G+I T LKNL L L LTG +P+ + L L+ + LS N+
Sbjct: 96 LTSLIFRKLTNLTGHIQPTIAKLKNLTFLRLSWTNLTGPVPEFLSQLKNLEYIDLSFNDL 155
Query: 206 SGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLI-SVLKLDLSHNFLEGKLLNEFGNL 264
SG+IP L L L+LSRN L+G +P + G V L LSHN L G + GN
Sbjct: 156 SGSIPSSLSSLRKLEYLELSRNKLTGPIPESFGTFSGKVPSLFLSHNQLSGTIPKSLGN- 214
Query: 265 KNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNM 324
+ +DL N+L + + + +S N D+ +K N L++++
Sbjct: 215 PDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMFQFDLSKVKLAKTLN--NLDMNHN 272
Query: 325 ELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLE 359
+ G IP S+ + L +S N + G + PK E
Sbjct: 273 GITGSIPAEWSK-AYFQLLNVSYNRLCGRI-PKGE 305
Score = 73.2 bits (178), Expect = 1e-12
Identities = 61/217 (28%), Positives = 108/217 (49%), Gaps = 7/217 (3%)
Query: 173 SLVLLENGLTGNIPQEIGNLVKLKRLVLSG-NNFSGNIPDIFGGLSDLLILDLSRNSLSG 231
SL++ + ++G IP E+G+L L L+ N +G+I L +L L LS +L+G
Sbjct: 74 SLIIQDGEISGQIPPEVGDLPYLTSLIFRKLTNLTGHIQPTIAKLKNLTFLRLSWTNLTG 133
Query: 232 TLPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMN-S 290
+P L +L ++ +DLS N L G + + +L+ L ++L N+L + S +
Sbjct: 134 PVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRNKLTGPIPESFGTFSGK 193
Query: 291 LEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNI 350
+ + LS+N L G I K + ++LS +L G+ K + +S N
Sbjct: 194 VPSLFLSHNQLSGTIP--KSLGNPDFYRIDLSRNKLQGDASILFGAKKTTWIVDISRNMF 251
Query: 351 TGNLSPKLETLPSLNALYLSGNNLKGEI--QFSKGFF 385
+LS K++ +LN L ++ N + G I ++SK +F
Sbjct: 252 QFDLS-KVKLAKTLNNLDMNHNGITGSIPAEWSKAYF 287
Score = 57.8 bits (138), Expect = 6e-08
Identities = 36/81 (44%), Positives = 47/81 (57%), Gaps = 1/81 (1%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L+NL L LS L G +PE LSQLK L ++ LS N+++G++ L +L L L LS N
Sbjct: 118 LKNLTFLRLSWTNLTGPVPEFLSQLKNLEYIDLSFNDLSGSIPSSLSSLRKLEYLELSRN 177
Query: 373 NLKGEIQFSKG-FFGKLGRRF 392
L G I S G F GK+ F
Sbjct: 178 KLTGPIPESFGTFSGKVPSLF 198
Score = 42.7 bits (99), Expect = 0.002
Identities = 29/76 (38%), Positives = 42/76 (55%), Gaps = 2/76 (2%)
Query: 313 LQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGN 372
L +L+ +L+N L G I ++++LK L FL LS N+TG + L L +L + LS N
Sbjct: 96 LTSLIFRKLTN--LTGHIQPTIAKLKNLTFLRLSWTNLTGPVPEFLSQLKNLEYIDLSFN 153
Query: 373 NLKGEIQFSKGFFGKL 388
+L G I S KL
Sbjct: 154 DLSGSIPSSLSSLRKL 169
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 80.9 bits (198), Expect = 7e-15
Identities = 46/104 (44%), Positives = 61/104 (58%)
Query: 154 NPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIF 213
N L G + G L NLQ L L N +TG IP+++GNL +L L L NN SG IP
Sbjct: 77 NANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTL 136
Query: 214 GGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGKL 257
G L L L L+ NSLSG +P +L ++++ LDLS+N L G +
Sbjct: 137 GRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDI 180
Score = 72.0 bits (175), Expect = 3e-12
Identities = 39/97 (40%), Positives = 59/97 (60%)
Query: 181 LTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRL 240
L+G + ++G L L+ L L NN +G IP+ G L++L+ LDL N+LSG +P TLGRL
Sbjct: 80 LSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRL 139
Query: 241 ISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRL 277
+ L L++N L G++ + L ++DL NN L
Sbjct: 140 KKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPL 176
Score = 72.0 bits (175), Expect = 3e-12
Identities = 43/109 (39%), Positives = 61/109 (55%)
Query: 197 RLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLEGK 256
R+ L N SG + G L +L L+L N+++GT+P LG L ++ LDL N L G
Sbjct: 72 RVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGP 131
Query: 257 LLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDI 305
+ + G LK L + L NN L + SL + +L+ + LSNNPL GDI
Sbjct: 132 IPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDI 180
Score = 67.8 bits (164), Expect = 6e-11
Identities = 44/119 (36%), Positives = 69/119 (57%), Gaps = 1/119 (0%)
Query: 266 NLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNME 325
++T +DL N L LV+ L ++ +L+ + L +N + G I + NL LV L+L
Sbjct: 69 SVTRVDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPE-QLGNLTELVSLDLYLNN 127
Query: 326 LIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGF 384
L G IP +L +LKKLRFL L++N+++G + L + +L L LS N L G+I + F
Sbjct: 128 LSGPIPSTLGRLKKLRFLRLNNNSLSGEIPRSLTAVLTLQVLDLSNNPLTGDIPVNGSF 186
Score = 65.5 bits (158), Expect = 3e-10
Identities = 39/91 (42%), Positives = 52/91 (56%), Gaps = 1/91 (1%)
Query: 145 SLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNN 204
+L+ +E SN + G IP G L L SL L N L+G IP +G L KL+ L L+ N+
Sbjct: 93 NLQYLELYSN-NITGTIPEQLGNLTELVSLDLYLNNLSGPIPSTLGRLKKLRFLRLNNNS 151
Query: 205 FSGNIPDIFGGLSDLLILDLSRNSLSGTLPV 235
SG IP + L +LDLS N L+G +PV
Sbjct: 152 LSGEIPRSLTAVLTLQVLDLSNNPLTGDIPV 182
Score = 46.6 bits (109), Expect = 1e-04
Identities = 28/70 (40%), Positives = 38/70 (54%)
Query: 319 LELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
++L N L G++ L QL L++L L NNITG + +L L L +L L NNL G I
Sbjct: 73 VDLGNANLSGQLVMQLGQLPNLQYLELYSNNITGTIPEQLGNLTELVSLDLYLNNLSGPI 132
Query: 379 QFSKGFFGKL 388
+ G KL
Sbjct: 133 PSTLGRLKKL 142
>LAP4_HUMAN (Q14160) LAP4 protein (Scribble homolog protein)
(hScrib)
Length = 1630
Score = 79.7 bits (195), Expect = 1e-14
Identities = 79/254 (31%), Positives = 124/254 (48%), Gaps = 39/254 (15%)
Query: 145 SLESIEFRSNPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNN 204
+L + ++ PG +GN L NL +L L EN L ++P + LVKL++L L GN+
Sbjct: 134 ALNDVSLQALPGDVGN-------LANLVTLELREN-LLKSLPASLSFLVKLEQLDLGGND 185
Query: 205 FSGNIPDIFGGLSDLLILDLSRNSLSGTLPVTLGRLISVLKLDLSHNFLE---------- 254
+PD G L +L L L RN LS LP LG L ++ LD+S N LE
Sbjct: 186 LEV-LPDTLGALPNLRELWLDRNQLS-ALPPELGNLRRLVCLDVSENRLEELPAELGGLV 243
Query: 255 ------------GKLLNEFGNLKNLTLMDLRNNRLCCGLVLSLQEMNSLEEMVLSNNPLG 302
+L + G LK L+++ + NRL C + ++ + +L E++L+ N L
Sbjct: 244 LLTDLLLSQNLLRRLPDGIGQLKQLSILKVDQNRL-CEVTEAIGDCENLSELILTENLLM 302
Query: 303 GDIRTL-KWENLQNLVILELSNMELIGEIPESLSQLKKLRFLGLSDNNITGNLSPKLETL 361
R+L K L NL + + +++E +P + L L L DN + L P+L
Sbjct: 303 ALPRSLGKLTKLTNLNV-DRNHLE---ALPPEIGGCVALSVLSLRDNRL-AVLPPELAHT 357
Query: 362 PSLNALYLSGNNLK 375
L+ L ++GN L+
Sbjct: 358 TELHVLDVAGNRLQ 371
Score = 32.3 bits (72), Expect = 2.7
Identities = 31/124 (25%), Positives = 52/124 (41%), Gaps = 22/124 (17%)
Query: 329 EIPESLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSKGFFGKL 388
E+P+ +L LR LGLSDN I L P++ L L +S N++ EI
Sbjct: 50 ELPKPFFRLLNLRKLGLSDNEIQ-RLPPEVANFMQLVELDVSRNDIP-EIP--------- 98
Query: 389 GRRFGAWSNPKLCYPFELMSTNNVPYGVKPCHQEEI----HLVKSNAKTEVINGDINHNS 444
+ K C E+ + P P ++ HL ++ + + GD+ + +
Sbjct: 99 -------ESIKFCKALEIADFSGNPLSRLPDGFTQLRSLAHLALNDVSLQALPGDVGNLA 151
Query: 445 NFIT 448
N +T
Sbjct: 152 NLVT 155
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 79.3 bits (194), Expect = 2e-14
Identities = 104/404 (25%), Positives = 157/404 (38%), Gaps = 97/404 (24%)
Query: 52 NSWNGSDLYPDPCGSTSIEGVSCDIFNGLWYVTVINIGPIHENSLPCANEKLEFKPELFQ 111
+S+ SD PDPC T I + G VT I IG H + L EL +
Sbjct: 43 SSFGWSD--PDPCKWTHI------VCTGTKRVTRIQIG--HSGLQGTLSPDLRNLSELER 92
Query: 112 LK-----------HLKAISFFNCFQSPNKLPVSIPTGNWEKLAESLESIEFRSNP----- 155
L+ L ++ N SIP+ ++ L SL+S+E +NP
Sbjct: 93 LELQWNNISGPVPSLSGLASLQVLMLSNNNFDSIPSDVFQGLT-SLQSVEIDNNPFKSWE 151
Query: 156 -------------------GLIGNIPSTFG--VLKNLQSLVLLENGLTGNIPQEIG---- 190
+ G++P G L L L N L G +P +
Sbjct: 152 IPESLRNASALQNFSANSANVSGSLPGFLGPDEFPGLSILHLAFNNLEGELPMSLAGSQV 211
Query: 191 ------------------NLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSLSGT 232
N+ LK + L N FSG +PD F GL +L L L NS +G
Sbjct: 212 QSLWLNGQKLTGDITVLQNMTGLKEVWLHSNKFSGPLPD-FSGLKELESLSLRDNSFTGP 270
Query: 233 LPVTLGRLISVLKLDLSHNFLEGKLLNEFGNLKNLTLMDLRNNRLCCG------------ 280
+P +L L S+ ++L++N L+G + F + ++ L D +N C
Sbjct: 271 VPASLLSLESLKVVNLTNNHLQGP-VPVFKSSVSVDL-DKDSNSFCLSSPGECDPRVKSL 328
Query: 281 --LVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQ----NLVILELSNMELIGEIPESL 334
+ S L E N+P W + N+ ++ L MEL G I
Sbjct: 329 LLIASSFDYPPRLAESWKGNDP------CTNWIGIACSNGNITVISLEKMELTGTISPEF 382
Query: 335 SQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEI 378
+K L+ + L NN+TG + +L TLP+L L +S N L G++
Sbjct: 383 GAIKSLQRIILGINNLTGMIPQELTTLPNLKTLDVSSNKLFGKV 426
Score = 79.0 bits (193), Expect = 3e-14
Identities = 67/230 (29%), Positives = 111/230 (48%), Gaps = 8/230 (3%)
Query: 156 GLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGG 215
GL G + L L+ L L N ++G +P + L L+ L+LS NNF D+F G
Sbjct: 75 GLQGTLSPDLRNLSELERLELQWNNISGPVPS-LSGLASLQVLMLSNNNFDSIPSDVFQG 133
Query: 216 LSDLLILDLSRNSL-SGTLPVTLGRLISVLKLDLSHNFLEGKLLNEFG--NLKNLTLMDL 272
L+ L +++ N S +P +L ++ + + G L G L+++ L
Sbjct: 134 LTSLQSVEIDNNPFKSWEIPESLRNASALQNFSANSANVSGSLPGFLGPDEFPGLSILHL 193
Query: 273 RNNRLCCGLVLSLQEMNSLEEMVLSNNPLGGDIRTLKWENLQNLVILELSNMELIGEIPE 332
N L L +SL + ++ + L+ L GDI L +N+ L + L + + G +P+
Sbjct: 194 AFNNLEGELPMSLAG-SQVQSLWLNGQKLTGDITVL--QNMTGLKEVWLHSNKFSGPLPD 250
Query: 333 SLSQLKKLRFLGLSDNNITGNLSPKLETLPSLNALYLSGNNLKGEIQFSK 382
S LK+L L L DN+ TG + L +L SL + L+ N+L+G + K
Sbjct: 251 -FSGLKELESLSLRDNSFTGPVPASLLSLESLKVVNLTNNHLQGPVPVFK 299
Score = 50.8 bits (120), Expect = 7e-06
Identities = 32/78 (41%), Positives = 41/78 (52%), Gaps = 2/78 (2%)
Query: 135 PTGNWEKLAESLESIEFRS--NPGLIGNIPSTFGVLKNLQSLVLLENGLTGNIPQEIGNL 192
P NW +A S +I S L G I FG +K+LQ ++L N LTG IPQE+ L
Sbjct: 350 PCTNWIGIACSNGNITVISLEKMELTGTISPEFGAIKSLQRIILGINNLTGMIPQELTTL 409
Query: 193 VKLKRLVLSGNNFSGNIP 210
LK L +S N G +P
Sbjct: 410 PNLKTLDVSSNKLFGKVP 427
Score = 45.1 bits (105), Expect = 4e-04
Identities = 25/65 (38%), Positives = 36/65 (54%)
Query: 170 NLQSLVLLENGLTGNIPQEIGNLVKLKRLVLSGNNFSGNIPDIFGGLSDLLILDLSRNSL 229
N+ + L + LTG I E G + L+R++L NN +G IP L +L LD+S N L
Sbjct: 363 NITVISLEKMELTGTISPEFGAIKSLQRIILGINNLTGMIPQELTTLPNLKTLDVSSNKL 422
Query: 230 SGTLP 234
G +P
Sbjct: 423 FGKVP 427
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,814,801
Number of Sequences: 164201
Number of extensions: 2509715
Number of successful extensions: 8301
Number of sequences better than 10.0: 321
Number of HSP's better than 10.0 without gapping: 175
Number of HSP's successfully gapped in prelim test: 151
Number of HSP's that attempted gapping in prelim test: 5836
Number of HSP's gapped (non-prelim): 1225
length of query: 475
length of database: 59,974,054
effective HSP length: 114
effective length of query: 361
effective length of database: 41,255,140
effective search space: 14893105540
effective search space used: 14893105540
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC146570.9


