

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146569.8 - phase: 0
(166 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
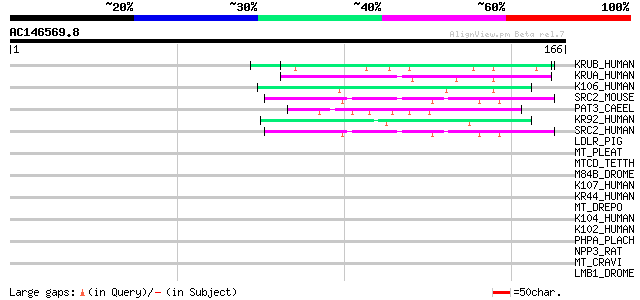
Score E
Sequences producing significant alignments: (bits) Value
KRUB_HUMAN (O75690) Keratin, ultra high-sulfur matrix protein B ... 48 1e-05
KRUA_HUMAN (P26371) Keratin, ultra high-sulfur matrix protein A ... 45 1e-04
K106_HUMAN (P60371) Keratin associated protein 10-6 (Keratin ass... 44 2e-04
SRC2_MOUSE (P59222) Scavenger receptor class F member 2 precurso... 43 4e-04
PAT3_CAEEL (Q27874) Integrin beta pat-3 precursor 43 4e-04
KR92_HUMAN (Q9BYQ4) Keratin associated protein 9-2 (Keratin asso... 42 6e-04
SRC2_HUMAN (Q96GP6) Scavenger receptor class F member 2 precurso... 42 8e-04
LDLR_PIG (Q28832) Low-density lipoprotein receptor (LDL receptor... 41 0.001
MT_PLEAT (Q6XUW5) Metallothionein (MT) 41 0.001
MTCD_TETTH (Q8WSW3) Cadmium metallothionein precursor (MT-CD) (C... 41 0.001
M84B_DROME (Q01643) Male specific sperm protein Mst84Db 40 0.002
K107_HUMAN (P60409) Keratin associated protein KAP10-7 (Keratin ... 40 0.002
KR44_HUMAN (Q9BYR3) Keratin associated protein 4-4 (Keratin asso... 40 0.002
MT_DREPO (Q94550) Metallothionein (MT) 40 0.003
K104_HUMAN (P60372) Keratin associated protein 10-4 (Keratin ass... 39 0.004
K102_HUMAN (P60368) Keratin associated protein 10-2 (Keratin ass... 39 0.004
PHPA_PLACH (Q02752) Acidic phosphoprotein precursor (50 kDa anti... 39 0.005
NPP3_RAT (P97675) Ectonucleotide pyrophosphatase/phosphodiestera... 39 0.005
MT_CRAVI (P23038) Metallothionein (MT) 39 0.005
LMB1_DROME (P11046) Laminin beta-1 chain precursor (Laminin B1 c... 39 0.005
>KRUB_HUMAN (O75690) Keratin, ultra high-sulfur matrix protein B
(UHS keratin B) (UHS KerB)
Length = 194
Score = 47.8 bits (112), Expect = 1e-05
Identities = 30/106 (28%), Positives = 40/106 (37%), Gaps = 25/106 (23%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-----------CGNSKCDGNKCN 130
GSC C+ +C+ C + C SS C + C CS CG+S C + C
Sbjct: 88 GSCGCSQCSCYKPCCCSSGCGSSCCQSSCCKPCCSQSSCCKPCSCSSGCGSSCCQSSCCK 147
Query: 131 GCTPKVT------------SPCFQR--CPQWCTCSKCCAPYQPYCK 162
C + + S C Q C C+ S CC P CK
Sbjct: 148 PCCSQSSCCKPCCCSSGCGSSCCQSSCCKPCCSQSSCCVPICCQCK 193
Score = 43.9 bits (102), Expect = 2e-04
Identities = 32/110 (29%), Positives = 39/110 (35%), Gaps = 19/110 (17%)
Query: 73 IEACHAVLHGSC---KCNNFNCHGKCNSCTKCESSK-------CDGTNCV--TICSFKCG 120
+ AC GSC K +C G C C SK C +C CS CG
Sbjct: 49 VPACSCSSCGSCGGSKGGRGSCGGSKGDCGSCGGSKGGCGSCGCSQCSCYKPCCCSSGCG 108
Query: 121 NSKCDGNKCNGCTPKVT----SPCFQRCPQWCTCSKCCAP---YQPYCKP 163
+S C + C C + + C C C S CC P CKP
Sbjct: 109 SSCCQSSCCKPCCSQSSCCKPCSCSSGCGSSCCQSSCCKPCCSQSSCCKP 158
>KRUA_HUMAN (P26371) Keratin, ultra high-sulfur matrix protein A
(UHS keratin A) (UHS KerA)
Length = 169
Score = 44.7 bits (104), Expect = 1e-04
Identities = 30/97 (30%), Positives = 40/97 (40%), Gaps = 17/97 (17%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKC------------GNSKCDGNKC 129
GSC C+ +C C + C SS C + C CS +C G+S C + C
Sbjct: 73 GSCGCSQCSCCKPCCCSSGCGSSCCQCSCCKPYCS-QCSCCKPCCSSSGRGSSCCQSSCC 131
Query: 130 NGC--TPKVTSPCFQR--CPQWCTCSKCCAPYQPYCK 162
C + S C Q C C+ S+CC P CK
Sbjct: 132 KPCCSSSGCGSSCCQSSCCKPCCSQSRCCVPVCYQCK 168
Score = 40.8 bits (94), Expect = 0.001
Identities = 25/80 (31%), Positives = 31/80 (38%), Gaps = 5/80 (6%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTP-KVTSPCF 141
SC + C C C S C C + K G C ++C+ C P +S C
Sbjct: 34 SCCAPVYCCKPVCCCVPACSCSSCGKRGCGSCGGSKGGCGSCGCSQCSCCKPCCCSSGCG 93
Query: 142 QRCPQWCTCSKCCAPYQPYC 161
C C CS CC PY C
Sbjct: 94 SSC---CQCS-CCKPYCSQC 109
Score = 34.7 bits (78), Expect = 0.10
Identities = 23/87 (26%), Positives = 31/87 (35%), Gaps = 5/87 (5%)
Query: 73 IEACHAVLHGSCKCNNFN-CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNG 131
+ AC G C + G C SC + S C C + C C C C+
Sbjct: 49 VPACSCSSCGKRGCGSCGGSKGGCGSCGCSQCSCCKPCCCSSGCGSSCCQCSCCKPYCSQ 108
Query: 132 CTPKVTSPCFQRCPQWCTC--SKCCAP 156
C+ PC + +C S CC P
Sbjct: 109 CS--CCKPCCSSSGRGSSCCQSSCCKP 133
Score = 32.0 bits (71), Expect = 0.66
Identities = 19/64 (29%), Positives = 25/64 (38%), Gaps = 14/64 (21%)
Query: 95 CNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCF-----QRCPQWCT 149
C C+ S C G C CG+ G+ C GC P +P + C C+
Sbjct: 3 CCGCSGGCGSSCGG------CDSSCGSC---GSGCRGCGPSCCAPVYCCKPVCCCVPACS 53
Query: 150 CSKC 153
CS C
Sbjct: 54 CSSC 57
>K106_HUMAN (P60371) Keratin associated protein 10-6 (Keratin
associated protein 10.6) (High sulfur keratin associated
protein 10.6) (Keratin associated protein 18-6) (Keratin
associated protein 18.6)
Length = 365
Score = 43.9 bits (102), Expect = 2e-04
Identities = 27/97 (27%), Positives = 38/97 (38%), Gaps = 15/97 (15%)
Query: 75 ACHAVLHGSCKCNNFNCHGKCNS--CTKCESSKCDGTNCVTICSFKCGNSKCDGNKC-NG 131
+C + S C C C C+ SS C ++CV+ S C + C+ + C +G
Sbjct: 140 SCQSACCTSSPCQQACCVPVCCKPVCSGISSSCCQQSSCVSCVSSPCCQAVCEPSPCQSG 199
Query: 132 CTPKVTSPCFQR------------CPQWCTCSKCCAP 156
CT T C Q+ C Q C CC P
Sbjct: 200 CTSSCTPSCCQQSSCQPTCCTSSPCQQACCVPVCCVP 236
Score = 38.5 bits (88), Expect = 0.007
Identities = 27/96 (28%), Positives = 38/96 (39%), Gaps = 14/96 (14%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC-------SFKCGNSK-CDGN 127
C + SC C +C C + + C+ + C C T+C S CG+S C +
Sbjct: 81 CTSSCTPSC-CQQSSCQLACCASSPCQQACCVPVCCKTVCCKPVCCVSVCCGDSSCCQQS 139
Query: 128 KCNG--CTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
C CT +SPC Q C C C+ C
Sbjct: 140 SCQSACCT---SSPCQQACCVPVCCKPVCSGISSSC 172
Score = 33.1 bits (74), Expect = 0.30
Identities = 25/100 (25%), Positives = 34/100 (34%), Gaps = 22/100 (22%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC---------SFKCGNSKCDG 126
C + SC C +C C + + C+ + C CV +C S C S C
Sbjct: 200 CTSSCTPSC-CQQSSCQPTCCTSSPCQQACCVPVCCVPVCCVPTCSEDSSSCCQQSSCQP 258
Query: 127 NKCNG-----------CTPKVTSPCFQ-RCPQWCTCSKCC 154
C C+ TS C Q C C + CC
Sbjct: 259 ACCTSSPCQHACCVPVCSGASTSCCQQSSCQPACCTASCC 298
Score = 30.8 bits (68), Expect = 1.5
Identities = 27/100 (27%), Positives = 36/100 (36%), Gaps = 29/100 (29%)
Query: 93 GKCNSCTK-----------CESSKCDGTNCVTIC-------SFKCGNSKCDGNKC-NGCT 133
G C+SC+ CE C C+++ S C C+ + C +GCT
Sbjct: 23 GSCDSCSDSWQVDDCPESCCEPPCCAPAPCLSLVCTPVSRVSSPCCPVTCEPSPCQSGCT 82
Query: 134 PKVTSPCFQR--CPQWCTCSK-----CCAPY---QPYCKP 163
T C Q+ C C S CC P CKP
Sbjct: 83 SSCTPSCCQQSSCQLACCASSPCQQACCVPVCCKTVCCKP 122
Score = 30.4 bits (67), Expect = 1.9
Identities = 16/55 (29%), Positives = 22/55 (39%), Gaps = 2/55 (3%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTK--CESSKCDGTNCVTICSFKCGNSKCDGNK 128
CH V +C +C +SC C ++ C C +CS S C G K
Sbjct: 308 CHPVCKSTCCVPVPSCGASASSCQPSCCRTASCVSLLCRPMCSRPACYSLCSGQK 362
Score = 28.5 bits (62), Expect = 7.3
Identities = 21/79 (26%), Positives = 28/79 (34%), Gaps = 18/79 (22%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSK---------CDGTNCVTICSFKCGNSKCDGNKCNG-- 131
SC C +C C + + C SS C T CV + S S C + C
Sbjct: 281 SC-CQQSSCQPACCTASCCRSSSSVSLLCHPVCKSTCCVPVPSCGASASSCQPSCCRTAS 339
Query: 132 -----CTPKVTSP-CFQRC 144
C P + P C+ C
Sbjct: 340 CVSLLCRPMCSRPACYSLC 358
>SRC2_MOUSE (P59222) Scavenger receptor class F member 2 precursor
(Scavenger receptor expressed by endothelial cells 2
protein) (SREC-II)
Length = 833
Score = 42.7 bits (99), Expect = 4e-04
Identities = 31/94 (32%), Positives = 45/94 (46%), Gaps = 10/94 (10%)
Query: 77 HAVLHGSCKCNNFNCHGKCNSC-TKCESSKCDGTNCVTICSFKCGNSKCD--GNKCNGCT 133
HA H + KC + N + C TKC S+ G +C +CS CG+ CD +C C+
Sbjct: 330 HACNHVTGKCTHCNAGWIGDRCETKC-SNGTYGEDCAFVCS-DCGSGHCDFQSGRCL-CS 386
Query: 134 PKVTSP-CFQRCP---QWCTCSKCCAPYQPYCKP 163
P V P C CP C++ C+ ++ C P
Sbjct: 387 PGVHGPHCNVTCPAGLHGVDCAQACSCHEESCDP 420
Score = 38.9 bits (89), Expect = 0.005
Identities = 32/119 (26%), Positives = 44/119 (36%), Gaps = 36/119 (30%)
Query: 76 CHAVLHGS-----CKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCD----- 125
CHA G+ C+C + CH + +C +CE G C + C + S+CD
Sbjct: 135 CHARRWGARCEHACQCQHGTCHPRSGAC-RCEPGWW-GAQCASAC-YCSATSRCDPQTGA 191
Query: 126 --------GNKCNGCTPKVTSPCFQ--------------RCPQWCTCSK-CCAPYQPYC 161
G CN +SPC Q RC ++C CS C P C
Sbjct: 192 CLCHVGWWGRSCNNQCACNSSPCEQQSGRCQCRERMFGARCDRYCQCSHGRCHPVDGTC 250
>PAT3_CAEEL (Q27874) Integrin beta pat-3 precursor
Length = 809
Score = 42.7 bits (99), Expect = 4e-04
Identities = 29/102 (28%), Positives = 46/102 (44%), Gaps = 33/102 (32%)
Query: 84 CKCNNFNC----------HGKCNSCTKC--------ESSKC--DGTNCVT----ICSFK- 118
C+C+NFNC HG+CN C KC + +C +C++ IC+ K
Sbjct: 563 CECDNFNCPRHDRKICAEHGECN-CGKCICAPGWTGRACECPISTDSCLSANGKICNGKG 621
Query: 119 ---CGNSKC----DGNKCNGCTPKVTSPCFQRCPQWCTCSKC 153
CG +C DGN+ +G ++ C +C ++ C C
Sbjct: 622 ECICGRCRCFDSPDGNRYSGAKCEICPTCPTKCVEYKNCVMC 663
>KR92_HUMAN (Q9BYQ4) Keratin associated protein 9-2 (Keratin
associated protein 9.2) (Ultrahigh sulfur keratin
associated protein 9.2)
Length = 174
Score = 42.0 bits (97), Expect = 6e-04
Identities = 26/87 (29%), Positives = 30/87 (33%), Gaps = 7/87 (8%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNC---VTICSFKCGNSKCDGNKCNGC 132
C GS C +C C + C C T C T+C C N C N C C
Sbjct: 86 CQPTCCGSSCCGQTSCGSSCGQSSSCAPVYCRRT-CYYPTTVCLPGCLNQSCGSNCCQPC 144
Query: 133 TPKV---TSPCFQRCPQWCTCSKCCAP 156
T+ C C Q S CC P
Sbjct: 145 CRPACCETTCCRTTCFQPTCVSSCCQP 171
Score = 36.2 bits (82), Expect = 0.035
Identities = 26/95 (27%), Positives = 34/95 (35%), Gaps = 3/95 (3%)
Query: 69 KKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNK 128
K + C + C +C C T C+++ C T C C C C
Sbjct: 25 KPTTVTTCSSTSCCQPACCVSSCCQPCCRPTSCQNTCCRTTCCQPTCVTSCCQPSCCSTP 84
Query: 129 CNGCTPKVTSPCFQ-RCPQWCTCSKCCAPYQPYCK 162
C T +S C Q C C S CAP YC+
Sbjct: 85 CCQPTCCGSSCCGQTSCGSSCGQSSSCAPV--YCR 117
Score = 30.8 bits (68), Expect = 1.5
Identities = 19/69 (27%), Positives = 26/69 (37%), Gaps = 11/69 (15%)
Query: 95 CNSCTKCESSKCDGTNCVTIC-----SFKCGNSKCDGNKCNGCTPKVTSPCFQ--RCPQW 147
C+ C C+ + C T C T C C ++ C C C PC + C
Sbjct: 5 CSPC--CQPTCCRTTCCRTTCWKPTTVTTCSSTSCCQPAC--CVSSCCQPCCRPTSCQNT 60
Query: 148 CTCSKCCAP 156
C + CC P
Sbjct: 61 CCRTTCCQP 69
>SRC2_HUMAN (Q96GP6) Scavenger receptor class F member 2 precursor
(Scavenger receptor expressed by endothelial cells 2
protein) (SREC-II) (SRECRP-1)
Length = 866
Score = 41.6 bits (96), Expect = 8e-04
Identities = 30/94 (31%), Positives = 44/94 (45%), Gaps = 10/94 (10%)
Query: 77 HAVLHGSCKCNNFNCHGKCNSC-TKCESSKCDGTNCVTICSFKCGNSKCD--GNKCNGCT 133
HA H + KC N + C TKC S+ G +C +C+ CG+ CD +C C+
Sbjct: 338 HACNHVTGKCTRCNAGWIGDRCETKC-SNGTYGEDCAFVCA-DCGSGHCDFQSGRCL-CS 394
Query: 134 PKVTSP-CFQRCP---QWCTCSKCCAPYQPYCKP 163
P V P C CP C++ C+ ++ C P
Sbjct: 395 PGVHGPHCNVTCPPGLHGADCAQACSCHEDTCDP 428
Score = 37.7 bits (86), Expect = 0.012
Identities = 31/119 (26%), Positives = 44/119 (36%), Gaps = 36/119 (30%)
Query: 76 CHAVLHGS-----CKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCD----- 125
CHA G+ C+C + CH + +C +CE G C + C + S+CD
Sbjct: 143 CHARRWGARCEHACQCQHGTCHPRSGAC-RCEPGWW-GAQCASAC-YCSATSRCDPQTGA 199
Query: 126 --------GNKCNGCTPKVTSPCFQ--------------RCPQWCTCSK-CCAPYQPYC 161
G CN +SPC Q RC ++C C + C P C
Sbjct: 200 CLCHAGWWGRSCNNQCACNSSPCEQQSGRCQCRERTFGARCDRYCQCFRGRCHPVDGTC 258
>LDLR_PIG (Q28832) Low-density lipoprotein receptor (LDL receptor)
(Fragment)
Length = 811
Score = 41.2 bits (95), Expect = 0.001
Identities = 30/93 (32%), Positives = 42/93 (44%), Gaps = 19/93 (20%)
Query: 64 EKEEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSK 123
E ++ + +E C +V +CK +F+C G+ N C ES +CDG C N
Sbjct: 20 ECKDGSDESLETCMSV---TCKIGDFSCGGRVNRCIP-ESWRCDGQQ-------DCEN-- 66
Query: 124 CDGNKCNGCTPKVTSPCFQRCPQWCTCSKCCAP 156
G+ GC+PK S RC KC AP
Sbjct: 67 --GSDEEGCSPKTCSQDEFRCQD----GKCIAP 93
>MT_PLEAT (Q6XUW5) Metallothionein (MT)
Length = 61
Score = 40.8 bits (94), Expect = 0.001
Identities = 24/62 (38%), Positives = 29/62 (46%), Gaps = 9/62 (14%)
Query: 73 IEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKC-DGNKCNG 131
++ C GSC NC G C SCT C + C T+C + C C SKC G C G
Sbjct: 1 MDPCECSKTGSC-----NCGGNC-SCTNCACTSCKKTSCCSCCPAGC--SKCASGCVCKG 52
Query: 132 CT 133
T
Sbjct: 53 KT 54
Score = 35.4 bits (80), Expect = 0.060
Identities = 20/56 (35%), Positives = 22/56 (38%), Gaps = 2/56 (3%)
Query: 101 CESSKCDGTNCVTICSF-KCGNSKCDGNKCNGCTPKVTSPCFQRCP-QWCTCSKCC 154
CE SK NC CS C + C C C P S C C + TC K C
Sbjct: 4 CECSKTGSCNCGGNCSCTNCACTSCKKTSCCSCCPAGCSKCASGCVCKGKTCDKTC 59
>MTCD_TETTH (Q8WSW3) Cadmium metallothionein precursor (MT-CD)
(CD-MT)
Length = 162
Score = 40.8 bits (94), Expect = 0.001
Identities = 28/90 (31%), Positives = 36/90 (39%), Gaps = 11/90 (12%)
Query: 74 EACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCT 133
E C G CKC NC C K +++ C G N C F + C +K N C
Sbjct: 40 ECCTGTGEG-CKC--VNC-----KCCKPQANCCCGVNAKPCC-FDPNSGCCCVSKTNNCC 90
Query: 134 PKVTSPCFQRCPQWCTCS--KCCAPYQPYC 161
T C + C C+ +CC P Q C
Sbjct: 91 KSDTKECCTGTGEGCKCTSCQCCKPVQQGC 120
Score = 29.6 bits (65), Expect = 3.3
Identities = 21/82 (25%), Positives = 28/82 (33%), Gaps = 5/82 (6%)
Query: 76 CHAVLHGSCKCNNFNC---HGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGC 132
C + + CK + C G+ CT C+ K C C K D N C
Sbjct: 82 CVSKTNNCCKSDTKECCTGTGEGCKCTSCQCCKPVQQGCC--CGDKAKACCTDPNSGCCC 139
Query: 133 TPKVTSPCFQRCPQWCTCSKCC 154
+ K C Q C +CC
Sbjct: 140 SNKANKCCDATSKQECQTCQCC 161
>M84B_DROME (Q01643) Male specific sperm protein Mst84Db
Length = 74
Score = 40.4 bits (93), Expect = 0.002
Identities = 22/71 (30%), Positives = 25/71 (34%), Gaps = 13/71 (18%)
Query: 91 CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQWCTC 150
C G C C C C + CS CG+ C C PC C C
Sbjct: 15 CGGPCGPCGPCGP-------CGSCCS-PCGSCCAPWGPCGPC-----GPCCGGCGPCGLC 61
Query: 151 SKCCAPYQPYC 161
CC P +PYC
Sbjct: 62 GPCCGPCRPYC 72
>K107_HUMAN (P60409) Keratin associated protein KAP10-7 (Keratin
associated protein 10.7) (High sulfur keratin associated
protein 10.7) (Keratin associated protein KAP18-7)
(Keratin associated protein 18.7)
Length = 370
Score = 40.4 bits (93), Expect = 0.002
Identities = 30/106 (28%), Positives = 42/106 (39%), Gaps = 28/106 (26%)
Query: 83 SCKCNNFNCHGKCNSCTKCE-----------------SSKCDGTNCVTICSFKCGNSKCD 125
SC C +C C + + C+ SS C ++CV+ S C + C+
Sbjct: 134 SC-CQQSSCQSACCTSSPCQQACCVPICCKPVCSGISSSCCQQSSCVSCVSSPCCQAVCE 192
Query: 126 GNKC-NGCTPKVTSPCFQRC---PQWCTCS----KCCAPYQPYCKP 163
+ C +GC T C Q+ P CT S CC P CKP
Sbjct: 193 PSPCQSGCISSCTPSCCQQSSCKPACCTSSPCQQACCVPV--CCKP 236
Score = 33.9 bits (76), Expect = 0.17
Identities = 24/99 (24%), Positives = 33/99 (33%), Gaps = 20/99 (20%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC-------------SFKCGNS 122
C + SC C +C C + + C+ + C C T+C S C S
Sbjct: 81 CTSSCTPSC-CQQSSCQLACCASSPCQQACCVPVCCKTVCCKPVYCVPVCSGDSSCCQQS 139
Query: 123 KCDGNKCNGCTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
C C +SPC Q C C C+ C
Sbjct: 140 SCQSACC------TSSPCQQACCVPICCKPVCSGISSSC 172
Score = 32.7 bits (73), Expect = 0.39
Identities = 30/107 (28%), Positives = 38/107 (35%), Gaps = 38/107 (35%)
Query: 93 GKCNSCTK------CESSKCDGTNC------------VTICSFKCGNSKCDGNKC-NGCT 133
G C+SC+ C S C+ C V+ S C C+ + C +GCT
Sbjct: 23 GSCDSCSDSWQVDDCPESCCEPPCCAPAPCLSLVCTPVSYVSSPCCRVTCEPSPCQSGCT 82
Query: 134 PKVTSPCFQR------------CPQWC---TCSK--CCAPYQPYCKP 163
T C Q+ C Q C C K CC P YC P
Sbjct: 83 SSCTPSCCQQSSCQLACCASSPCQQACCVPVCCKTVCCKPV--YCVP 127
Score = 31.2 bits (69), Expect = 1.1
Identities = 21/68 (30%), Positives = 24/68 (34%), Gaps = 7/68 (10%)
Query: 91 CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNG-CTPKVTSPCFQRCPQWCT 149
C G SC C+ S C C T C C S C C P P C +
Sbjct: 280 CSGASTSC--CQQSSCQPACCTTSC---CRPSSSVSLLCRPVCRPACCVPVPSCCAPTSS 334
Query: 150 C-SKCCAP 156
C + CC P
Sbjct: 335 CQASCCRP 342
Score = 28.5 bits (62), Expect = 7.3
Identities = 17/74 (22%), Positives = 27/74 (35%), Gaps = 7/74 (9%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQ 142
SC C +C C + + C+ + C C +C C + + C P +
Sbjct: 207 SC-CQQSSCKPACCTSSPCQQACCVPVCCKPVCCV----PTCSDDSGSCCQPACCTS--S 259
Query: 143 RCPQWCTCSKCCAP 156
+ Q C CC P
Sbjct: 260 QSQQGCCVPVCCKP 273
>KR44_HUMAN (Q9BYR3) Keratin associated protein 4-4 (Keratin
associated protein 4.4) (Ultrahigh sulfur keratin
associated protein 4.4)
Length = 166
Score = 40.0 bits (92), Expect = 0.002
Identities = 32/102 (31%), Positives = 37/102 (35%), Gaps = 18/102 (17%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNC-VTICSFKCGNSKCDGNKCNGCTP 134
CH S C C C T C C T C T C C +C + C C P
Sbjct: 61 CHPSCCVSSCCRPQCCQSVCCQPTCCRPQCCQTTCCRTTCCRPSCCRPQCCQSVC--CQP 118
Query: 135 KVTSP--CFQRC--PQWC--TCSK--CCAP-------YQPYC 161
P C C PQ C TC + CC P Y+P+C
Sbjct: 119 TCCCPSYCVSSCCRPQCCQTTCCRTTCCRPSCCVSRCYRPHC 160
Score = 34.7 bits (78), Expect = 0.10
Identities = 25/82 (30%), Positives = 34/82 (40%), Gaps = 13/82 (15%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCV-TICSFKCGNSKCDGNKCNGCTPKVTSPCF 141
SC C + C +C T C ++ C + CV + C +C S C C P P
Sbjct: 39 SC-CVSSCCRPQCCQTTCCRTTCCHPSCCVSSCCRPQCCQSVC-------CQPTCCRP-- 88
Query: 142 QRCPQWCTCSKCCAPYQPYCKP 163
Q C C + CC P C+P
Sbjct: 89 QCCQTTCCRTTCCRP--SCCRP 108
>MT_DREPO (Q94550) Metallothionein (MT)
Length = 73
Score = 39.7 bits (91), Expect = 0.003
Identities = 25/88 (28%), Positives = 33/88 (37%), Gaps = 21/88 (23%)
Query: 72 MIEACHAVLHGSCKCNNFNCHG----KCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGN 127
M + C+ V G C+C + +C KC KC C G N + KCG + G
Sbjct: 1 MSDPCNCVETGDCRCADGSCSDCSNCKCGDSCKCSKPNCCGKN----VTCKCGENCQCGV 56
Query: 128 KCNGCTPKVTSPCFQRCPQWCTCSKCCA 155
C G P CTC C+
Sbjct: 57 GCTG-------------PDSCTCDSGCS 71
Score = 29.6 bits (65), Expect = 3.3
Identities = 16/43 (37%), Positives = 21/43 (48%), Gaps = 7/43 (16%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKC-------DGTNCVTICSFK 118
SCKC+ NC GK +C E+ +C D C + CS K
Sbjct: 31 SCKCSKPNCCGKNVTCKCGENCQCGVGCTGPDSCTCDSGCSCK 73
>K104_HUMAN (P60372) Keratin associated protein 10-4 (Keratin
associated protein 10.4) (High sulfur keratin associated
protein 10.4) (Keratin associated protein 18-4) (Keratin
associated protein 18.4)
Length = 401
Score = 39.3 bits (90), Expect = 0.004
Identities = 30/109 (27%), Positives = 42/109 (38%), Gaps = 29/109 (26%)
Query: 83 SCKCNNFNCHGKCNSCTKCE-----------------SSKCDGTNCVTICSFKCGNSKCD 125
SC C +C C + + C+ SS C ++CV+ S C + C+
Sbjct: 134 SC-CQQSSCQSACCTSSPCQQACCVPICCKPVCSGISSSCCQQSSCVSCVSSPCCQAVCE 192
Query: 126 GNKC-NGCTPKVTSPCFQRC---PQWCTCSK----CCAPY---QPYCKP 163
+ C +GC T C Q+ P CT S CC P CKP
Sbjct: 193 PSPCQSGCISSCTPSCCQQSSCQPACCTSSSCQQACCVPVCCKTVCCKP 241
Score = 36.6 bits (83), Expect = 0.027
Identities = 26/96 (27%), Positives = 37/96 (38%), Gaps = 14/96 (14%)
Query: 76 CHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTIC-------SFKCGNSK-CDGN 127
C + SC C +C C + + C+ + C C T+C CG+S C +
Sbjct: 81 CTSSCTPSC-CQQSSCQLACCASSPCQQACCVPVCCKTVCCKPVCCVPVCCGDSSCCQQS 139
Query: 128 KCNG--CTPKVTSPCFQRCPQWCTCSKCCAPYQPYC 161
C CT +SPC Q C C C+ C
Sbjct: 140 SCQSACCT---SSPCQQACCVPICCKPVCSGISSSC 172
Score = 35.8 bits (81), Expect = 0.046
Identities = 22/78 (28%), Positives = 31/78 (39%), Gaps = 5/78 (6%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK-CGNSKCDGNKCNGCTPK--VTSP 139
SC C +C C + + C+ + C C T+C C + + C P +SP
Sbjct: 207 SC-CQQSSCQPACCTSSSCQQACCVPVCCKTVCCKPVCSEDSSSCCQQSSCQPACCTSSP 265
Query: 140 CFQRCPQWCTCSK-CCAP 156
C Q C C CC P
Sbjct: 266 CQQACCVPVCCKPVCCKP 283
Score = 31.6 bits (70), Expect = 0.87
Identities = 25/91 (27%), Positives = 33/91 (35%), Gaps = 13/91 (14%)
Query: 86 CNNFNCHGKCNSCTKCESSKCDGTNCVTIC--SFKCGNSKCDGNKC-NGCTPKVTSPCFQ 142
C C C + + C + C C + S C C+ + C +GCT T C Q
Sbjct: 32 CPESCCEPPCCAPSCCAPAPCLSLVCTPVSRVSSPCCPVTCEPSPCQSGCTSSCTPSCCQ 91
Query: 143 R--CPQWCTCSK-----CCAPY---QPYCKP 163
+ C C S CC P CKP
Sbjct: 92 QSSCQLACCASSPCQQACCVPVCCKTVCCKP 122
Score = 31.2 bits (69), Expect = 1.1
Identities = 21/68 (30%), Positives = 24/68 (34%), Gaps = 7/68 (10%)
Query: 91 CHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNG-CTPKVTSPCFQRCPQWCT 149
C G +SC C+ S C C T C C S C C P P C +
Sbjct: 332 CSGASSSC--CQQSSCQPACCTTSC---CRPSSSVSLLCRPVCRPACCVPVPSCCAPTSS 386
Query: 150 CS-KCCAP 156
C CC P
Sbjct: 387 CQPSCCRP 394
Score = 31.2 bits (69), Expect = 1.1
Identities = 25/88 (28%), Positives = 36/88 (40%), Gaps = 18/88 (20%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQ 142
SC C +C C + + C+ + C C +C G+ C+G +S C Q
Sbjct: 249 SC-CQQSSCQPACCTSSPCQQACCVPVCCKPVCCKPVGSVPI----CSG----ASSLCCQ 299
Query: 143 RC---PQWCTCSK----CCAPYQPYCKP 163
+ P CT S+ CC P CKP
Sbjct: 300 QSSCQPACCTSSQSQQGCCVPV--CCKP 325
>K102_HUMAN (P60368) Keratin associated protein 10-2 (Keratin
associated protein 10.2) (High sulfur keratin associated
protein 10.2) (Keratin associated protein 18-2) (Keratin
associated protein 18.2)
Length = 255
Score = 39.3 bits (90), Expect = 0.004
Identities = 24/76 (31%), Positives = 32/76 (41%), Gaps = 11/76 (14%)
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQ 142
SC C +C C + + C+ S C C +C S C C+G +SPC Q
Sbjct: 161 SC-CQQSSCQPACCTSSPCQQSCCVSVCCKPVCC----KSICCVPVCSG----ASSPCCQ 211
Query: 143 R--CPQWCTCSKCCAP 156
+ C C S CC P
Sbjct: 212 QSSCQPACCTSSCCRP 227
Score = 37.0 bits (84), Expect = 0.021
Identities = 27/88 (30%), Positives = 36/88 (40%), Gaps = 12/88 (13%)
Query: 86 CNNFNCHGKCNSCTKCESSKCDGTNC--VTICSFKCGNSKCDGNKC-NGCTPKVTSPCFQ 142
C C C + + C + C C V+ S C + C+ + C +GCT T C Q
Sbjct: 22 CPESCCELPCGTPSCCAPAPCLTLVCTPVSCVSSPCCQAACEPSACQSGCTSSCTPSCCQ 81
Query: 143 RC---PQWCTCS----KCCAPYQPYCKP 163
+ P CT S CC P CKP
Sbjct: 82 QSSCQPACCTSSPCQQACCVPV--CCKP 107
Score = 36.2 bits (82), Expect = 0.035
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 10/90 (11%)
Query: 75 ACHAVLHGSCK---CNNFNCHGKCNSCTKCESSKCDGTNCVTICSFK--CGNSKCDGNKC 129
AC + SC C +C C + + C+ + C C +C CG S C +
Sbjct: 66 ACQSGCTSSCTPSCCQQSSCQPACCTSSPCQQACCVPVCCKPVCCVPVCCGASSC--CQQ 123
Query: 130 NGCTPK--VTSPCFQRC-PQWCTCSKCCAP 156
+ C P +S C Q C C + CC P
Sbjct: 124 SSCQPACCASSSCQQSCRVPVCCKAVCCVP 153
>PHPA_PLACH (Q02752) Acidic phosphoprotein precursor (50 kDa
antigen)
Length = 441
Score = 38.9 bits (89), Expect = 0.005
Identities = 17/36 (47%), Positives = 27/36 (74%)
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIE 74
KKK N +TT++ KKK + +KKKEKE++K+K ++
Sbjct: 382 KKKVKNKPTMTTKKKKKKEKKKKKKEKEKKKEKKVK 417
>NPP3_RAT (P97675) Ectonucleotide pyrophosphatase/phosphodiesterase
3 (E-NPP 3) (Phosphodiesterase I/nucleotide
pyrophosphatase 3) (Phosphodiesterase I beta) (PD-Ibeta)
(RB13-6 antigen) (B10) [Includes: Alkaline
phosphodiesterase I (EC 3.1.4.1); Nucleotid
Length = 875
Score = 38.9 bits (89), Expect = 0.005
Identities = 24/94 (25%), Positives = 40/94 (42%), Gaps = 14/94 (14%)
Query: 69 KKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNK 128
+KK ++ H L G C+C++ C + + C E + T T SF+CG ++ +
Sbjct: 56 RKKCFDSSHRGLEG-CRCDS-GCTDRGDCCWDFEDTCVKSTQIWTCNSFRCGETRLEAAL 113
Query: 129 CNGCTPKVTSPCFQRCPQWCTCSKCCAPYQPYCK 162
C+ C QR CC Y+ C+
Sbjct: 114 CS-----CADDCLQR-------KDCCTDYKAVCQ 135
>MT_CRAVI (P23038) Metallothionein (MT)
Length = 74
Score = 38.9 bits (89), Expect = 0.005
Identities = 25/83 (30%), Positives = 36/83 (43%), Gaps = 14/83 (16%)
Query: 74 EACHAVLHGSCKCNNFNCHG---KCNSCTKC-ESSKCDGTNCVTICSFKCGNSKCDGNKC 129
+ C+ + G+C C++ +C KC KC + KC G C KC + G KC
Sbjct: 2 DPCNCIETGTCACSD-SCPATGCKCGPGCKCGDDCKCAG------CKVKCSCTSEGGCKC 54
Query: 130 NGCTPKVTSPCFQRCPQWCTCSK 152
K T P +C C+C K
Sbjct: 55 G---EKCTGPATCKCGSGCSCKK 74
>LMB1_DROME (P11046) Laminin beta-1 chain precursor (Laminin B1
chain)
Length = 1790
Score = 38.9 bits (89), Expect = 0.005
Identities = 28/100 (28%), Positives = 36/100 (36%), Gaps = 22/100 (22%)
Query: 82 GSCKCNNFNCHGKCNSCTK------------CESSKCD--GTNCVTICSFKCGNSKC--- 124
G+C C F +CN C CE C+ GT + C + G KC
Sbjct: 446 GACHCKAFVTGRRCNQCKDGYWNLQSDNPEGCEPCTCNPLGTLNNSGCVMRTGECKCKKY 505
Query: 125 -DGNKCNGCTPKVTSPCFQRCPQWCTCSKCCA--PYQPYC 161
G CN C P+ P+ C+ C A Y YC
Sbjct: 506 VTGKDCNQCMPETYG--LSESPEGCSLCNCDAGGSYDNYC 543
Score = 30.4 bits (67), Expect = 1.9
Identities = 18/56 (32%), Positives = 21/56 (37%), Gaps = 5/56 (8%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTI-CSFKCGNSKC----DGNKCNGC 132
G CN H N KC+ +CD C + GN C G KCN C
Sbjct: 1122 GGRACNQCQAHYWGNPNEKCQPCECDQFGAADFQCDRETGNCVCHEGIGGYKCNEC 1177
Score = 29.6 bits (65), Expect = 3.3
Identities = 21/82 (25%), Positives = 29/82 (34%), Gaps = 8/82 (9%)
Query: 82 GSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCF 141
G CKC + CN C + ++C+ G S N C+ ++ C
Sbjct: 498 GECKCKKYVTGKDCNQCMPETYGLSESPEGCSLCNCDAGGSY--DNYCD----VISGQC- 550
Query: 142 QRCPQWCTCSKCCAPYQPYCKP 163
RC T C P Q Y P
Sbjct: 551 -RCRPHMTGRSCSQPKQNYFIP 571
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.134 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,661,070
Number of Sequences: 164201
Number of extensions: 976051
Number of successful extensions: 11494
Number of sequences better than 10.0: 535
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 392
Number of HSP's that attempted gapping in prelim test: 9340
Number of HSP's gapped (non-prelim): 1798
length of query: 166
length of database: 59,974,054
effective HSP length: 102
effective length of query: 64
effective length of database: 43,225,552
effective search space: 2766435328
effective search space used: 2766435328
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146569.8


