

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.9 + phase: 0
(80 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
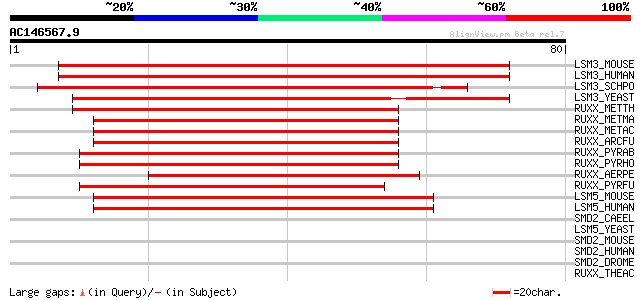
Sequences producing significant alignments: (bits) Value
LSM3_MOUSE (P62311) U6 snRNA-associated Sm-like protein LSm3 114 3e-26
LSM3_HUMAN (P62310) U6 snRNA-associated Sm-like protein LSm3 (MD... 114 3e-26
LSM3_SCHPO (Q9Y7M4) Probable U6 snRNA-associated Sm-like protein... 91 7e-19
LSM3_YEAST (P57743) U6 snRNA-associated Sm-like protein LSm3 (Sm... 58 5e-09
RUXX_METTH (O26745) Putative snRNP Sm-like protein 51 6e-07
RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein 47 9e-06
RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein 47 9e-06
RUXX_ARCFU (O29386) Putative snRNP Sm-like protein 47 9e-06
RUXX_PYRAB (Q9V0Y8) Putative snRNP Sm-like protein 46 2e-05
RUXX_PYRHO (O74016) Putative snRNP Sm-like protein 45 3e-05
RUXX_AERPE (Q9YEQ5) Putative snRNP Sm-like protein 45 3e-05
RUXX_PYRFU (Q8U0P4) Putative snRNP Sm-like protein 44 6e-05
LSM5_MOUSE (P62322) U6 snRNA-associated Sm-like protein LSm5 42 2e-04
LSM5_HUMAN (Q9Y4Y9) U6 snRNA-associated Sm-like protein LSm5 42 2e-04
SMD2_CAEEL (Q18786) Probable small nuclear ribonucleoprotein Sm ... 40 0.001
LSM5_YEAST (P40089) U6 snRNA-associated Sm-like protein LSm5 40 0.001
SMD2_MOUSE (P62317) Small nuclear ribonucleoprotein Sm D2 (snRNP... 39 0.002
SMD2_HUMAN (P62316) Small nuclear ribonucleoprotein Sm D2 (snRNP... 39 0.002
SMD2_DROME (Q9VI10) Probable small nuclear ribonucleoprotein Sm ... 39 0.002
RUXX_THEAC (P57670) Putative snRNP Sm-like protein 38 0.004
>LSM3_MOUSE (P62311) U6 snRNA-associated Sm-like protein LSm3
Length = 101
Score = 114 bits (286), Expect = 3e-26
Identities = 54/65 (83%), Positives = 62/65 (95%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDLIRLSLDERIYVK+R+DRELRG+LHAYDQHLNMILGDVEE VTT+EID+ETY
Sbjct: 11 NTVEEPLDLIRLSLDERIYVKMRNDRELRGRLHAYDQHLNMILGDVEETVTTIEIDEETY 70
Query: 68 EEIVR 72
EEI +
Sbjct: 71 EEIYK 75
>LSM3_HUMAN (P62310) U6 snRNA-associated Sm-like protein LSm3
(MDS017)
Length = 101
Score = 114 bits (286), Expect = 3e-26
Identities = 54/65 (83%), Positives = 62/65 (95%)
Query: 8 SAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETY 67
+ V+EPLDLIRLSLDERIYVK+R+DRELRG+LHAYDQHLNMILGDVEE VTT+EID+ETY
Sbjct: 11 NTVEEPLDLIRLSLDERIYVKMRNDRELRGRLHAYDQHLNMILGDVEETVTTIEIDEETY 70
Query: 68 EEIVR 72
EEI +
Sbjct: 71 EEIYK 75
>LSM3_SCHPO (Q9Y7M4) Probable U6 snRNA-associated Sm-like protein
LSm3
Length = 93
Score = 90.5 bits (223), Expect = 7e-19
Identities = 44/62 (70%), Positives = 51/62 (81%), Gaps = 1/62 (1%)
Query: 5 EEESAVKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDD 64
E AV EPLDL+RLSLDE +YVKLR DREL G+LHAYD+HLNM+LGD EEIVT + D+
Sbjct: 2 ESAQAVAEPLDLVRLSLDEIVYVKLRGDRELNGRLHAYDEHLNMVLGDAEEIVTIFD-DE 60
Query: 65 ET 66
ET
Sbjct: 61 ET 62
>LSM3_YEAST (P57743) U6 snRNA-associated Sm-like protein LSm3
(SmX4 protein)
Length = 89
Score = 57.8 bits (138), Expect = 5e-09
Identities = 28/63 (44%), Positives = 42/63 (66%), Gaps = 2/63 (3%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEE 69
++ PLDL++L+LDER+Y+KLR R L G L A+D H N++L D E T ++++E E
Sbjct: 1 METPLDLLKLNLDERVYIKLRGARTLVGTLQAFDSHCNIVLSDAVE--TIYQLNNEELSE 58
Query: 70 IVR 72
R
Sbjct: 59 SER 61
>RUXX_METTH (O26745) Putative snRNP Sm-like protein
Length = 81
Score = 50.8 bits (120), Expect = 6e-07
Identities = 21/47 (44%), Positives = 33/47 (69%)
Query: 10 VKEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
V+ PLD + SL+ + +KL+ DRE RG L ++D H+N++L D EE+
Sbjct: 11 VQRPLDALGNSLNSPVIIKLKGDREFRGVLKSFDLHMNLVLNDAEEL 57
>RUXX_METMA (Q8PZZ9) Putative snRNP Sm-like protein
Length = 72
Score = 47.0 bits (110), Expect = 9e-06
Identities = 19/44 (43%), Positives = 31/44 (70%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
PLD++ +LD + V+L+ RE RG+L YD H+N++L + EE+
Sbjct: 5 PLDILNNALDTPVIVRLKGAREFRGELKGYDIHMNLVLDNAEEL 48
>RUXX_METAC (Q8TL47) Putative snRNP Sm-like protein
Length = 72
Score = 47.0 bits (110), Expect = 9e-06
Identities = 19/44 (43%), Positives = 31/44 (70%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
PLD++ +LD + V+L+ RE RG+L YD H+N++L + EE+
Sbjct: 5 PLDILNNALDTPVIVRLKGAREFRGELKGYDIHMNLVLDNAEEL 48
>RUXX_ARCFU (O29386) Putative snRNP Sm-like protein
Length = 77
Score = 47.0 bits (110), Expect = 9e-06
Identities = 21/44 (47%), Positives = 29/44 (65%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
PLD++ SL + V+L+ RE RG L YD H+N++L D EEI
Sbjct: 5 PLDVLNRSLKSPVIVRLKGGREFRGTLDGYDIHMNLVLLDAEEI 48
>RUXX_PYRAB (Q9V0Y8) Putative snRNP Sm-like protein
Length = 75
Score = 45.8 bits (107), Expect = 2e-05
Identities = 22/46 (47%), Positives = 30/46 (64%)
Query: 11 KEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
+ PLD+I SLD+ + V L+ E RG+L YD HLN++L D E I
Sbjct: 3 ERPLDVIHRSLDKDVLVILKKGFEFRGRLIGYDIHLNVVLADAEMI 48
>RUXX_PYRHO (O74016) Putative snRNP Sm-like protein
Length = 75
Score = 45.4 bits (106), Expect = 3e-05
Identities = 21/46 (45%), Positives = 30/46 (64%)
Query: 11 KEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEI 56
+ PLD+I SLD+ + V L+ E RG+L YD HLN++L D E +
Sbjct: 3 ERPLDVIHRSLDKDVLVILKKGFEFRGRLIGYDIHLNVVLADAEMV 48
>RUXX_AERPE (Q9YEQ5) Putative snRNP Sm-like protein
Length = 77
Score = 45.1 bits (105), Expect = 3e-05
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 21 LDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTT 59
+D + VKL+S ++G L YDQHLN+ILGD EEI T
Sbjct: 17 VDTPVLVKLKSGLRIKGVLKTYDQHLNIILGDAEEIGET 55
>RUXX_PYRFU (Q8U0P4) Putative snRNP Sm-like protein
Length = 76
Score = 44.3 bits (103), Expect = 6e-05
Identities = 21/44 (47%), Positives = 29/44 (65%)
Query: 11 KEPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVE 54
+ PLD+I SLD+ + V L+ E RGKL YD HLN++L + E
Sbjct: 3 ERPLDVIHKSLDKDVLVILKKGFEFRGKLIGYDIHLNVVLANAE 46
>LSM5_MOUSE (P62322) U6 snRNA-associated Sm-like protein LSm5
Length = 90
Score = 42.4 bits (98), Expect = 2e-04
Identities = 19/49 (38%), Positives = 31/49 (62%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
PL+L+ + RI++ ++SD+E+ G L +D +NM+L DV E T E
Sbjct: 13 PLELVDKCIGSRIHIVMKSDKEIVGTLLGFDDFVNMVLEDVTEFEITPE 61
>LSM5_HUMAN (Q9Y4Y9) U6 snRNA-associated Sm-like protein LSm5
Length = 90
Score = 42.4 bits (98), Expect = 2e-04
Identities = 19/49 (38%), Positives = 31/49 (62%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVE 61
PL+L+ + RI++ ++SD+E+ G L +D +NM+L DV E T E
Sbjct: 13 PLELVDKCIGSRIHIVMKSDKEIVGTLLGFDDFVNMVLEDVTEFEITPE 61
>SMD2_CAEEL (Q18786) Probable small nuclear ribonucleoprotein Sm
D2 (snRNP core protein D2) (Sm-D2)
Length = 118
Score = 39.7 bits (91), Expect = 0.001
Identities = 22/66 (33%), Positives = 41/66 (61%), Gaps = 9/66 (13%)
Query: 4 TEEESAVKE-------PLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVE 54
T EE A KE PL ++ S+ + ++ + R++++L G++ A+D+H NM+L +V+
Sbjct: 12 TAEELAAKEDEEFNVGPLSILTNSVKNNHQVLINCRNNKKLLGRVKAFDRHCNMVLENVK 71
Query: 55 EIVTTV 60
E+ T V
Sbjct: 72 EMWTEV 77
>LSM5_YEAST (P40089) U6 snRNA-associated Sm-like protein LSm5
Length = 93
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/63 (31%), Positives = 39/63 (61%), Gaps = 2/63 (3%)
Query: 13 PLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTVEIDDETYEEIVR 72
PL++I ++++++ + L+S+RE G L +D +N+IL D E + ++ +DE+ E V
Sbjct: 8 PLEVIDKTINQKVLIVLQSNREFEGTLVGFDDFVNVILEDAVEWL--IDPEDESRNEKVM 65
Query: 73 VSH 75
H
Sbjct: 66 QHH 68
>SMD2_MOUSE (P62317) Small nuclear ribonucleoprotein Sm D2 (snRNP
core protein D2) (Sm-D2)
Length = 118
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/58 (32%), Positives = 38/58 (64%), Gaps = 2/58 (3%)
Query: 5 EEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
EEE PL ++ S+ + ++ + R++++L G++ A+D+H NM+L +V+E+ T V
Sbjct: 20 EEEEFNTGPLSVLTQSVKNNTQVLINCRNNKKLLGRVKAFDRHCNMVLENVKEMWTEV 77
>SMD2_HUMAN (P62316) Small nuclear ribonucleoprotein Sm D2 (snRNP
core protein D2) (Sm-D2)
Length = 118
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/58 (32%), Positives = 38/58 (64%), Gaps = 2/58 (3%)
Query: 5 EEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVTTV 60
EEE PL ++ S+ + ++ + R++++L G++ A+D+H NM+L +V+E+ T V
Sbjct: 20 EEEEFNTGPLSVLTQSVKNNTQVLINCRNNKKLLGRVKAFDRHCNMVLENVKEMWTEV 77
>SMD2_DROME (Q9VI10) Probable small nuclear ribonucleoprotein Sm
D2 (snRNP core protein D2) (Sm-D2)
Length = 119
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/60 (31%), Positives = 39/60 (64%), Gaps = 2/60 (3%)
Query: 1 MAGTEEESAVKEPLDLIRLSL--DERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIVT 58
+A EEE PL ++ S+ + ++ + R++++L G++ A+D+H NM+L +V+E+ T
Sbjct: 15 LARQEEEEFNTGPLSVLTQSVKNNTQVLINCRNNKKLLGRVKAFDRHCNMVLENVKEMWT 74
>RUXX_THEAC (P57670) Putative snRNP Sm-like protein
Length = 83
Score = 38.1 bits (87), Expect = 0.004
Identities = 13/46 (28%), Positives = 30/46 (64%)
Query: 12 EPLDLIRLSLDERIYVKLRSDRELRGKLHAYDQHLNMILGDVEEIV 57
+P+D+++ +L + + ++ +RE G L YD ++N++L + EI+
Sbjct: 9 KPMDVLKSALSRNVLIDVKGNREYSGILEGYDVYMNIVLQNASEII 54
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.134 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,140,744
Number of Sequences: 164201
Number of extensions: 318259
Number of successful extensions: 834
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 782
Number of HSP's gapped (non-prelim): 65
length of query: 80
length of database: 59,974,054
effective HSP length: 56
effective length of query: 24
effective length of database: 50,778,798
effective search space: 1218691152
effective search space used: 1218691152
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146567.9


