

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.11 - phase: 0
(331 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
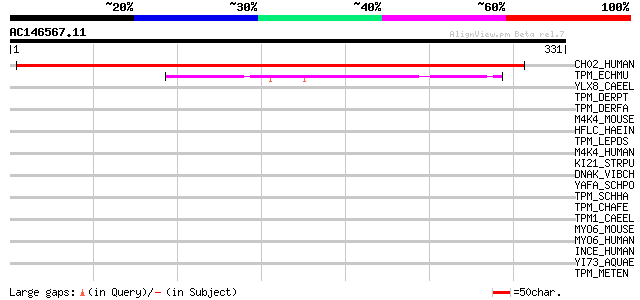
Score E
Sequences producing significant alignments: (bits) Value
CH02_HUMAN (O94905) Protein C8orf2 (UNQ2441/PRO9924/PRO5003) 348 1e-95
TPM_ECHMU (Q95PU1) Tropomyosin 45 3e-04
YLX8_CAEEL (P46504) Hypothetical protein F23F12.8 in chromosome ... 42 0.002
TPM_DERPT (O18416) Tropomyosin (Allergen Der p 10) 42 0.002
TPM_DERFA (Q23939) Tropomyosin (Allergen Der f 10) (Mag44) 42 0.002
M4K4_MOUSE (P97820) Mitogen-activated protein kinase kinase kina... 40 0.011
HFLC_HAEIN (P44545) HflC protein (EC 3.4.-.-) 40 0.011
TPM_LEPDS (Q9NFZ4) Tropomyosin (Allergen Lep d 10) 39 0.014
M4K4_HUMAN (O95819) Mitogen-activated protein kinase kinase kina... 39 0.014
KI21_STRPU (P46871) Kinesin-II 95 kDa subunit (KRP-85/95 95 kDa ... 39 0.018
DNAK_VIBCH (O34241) Chaperone protein dnaK (Heat shock protein 7... 38 0.031
YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I 38 0.040
TPM_SCHHA (Q26503) Tropomyosin 38 0.040
TPM_CHAFE (Q9N2R3) Tropomyosin (Allergen Cha f 1) (Cha f I) (Fra... 38 0.040
TPM1_CAEEL (Q22866) Tropomyosin isoforms 1/2 38 0.040
MYO6_MOUSE (Q64331) Myosin VI 38 0.040
MYO6_HUMAN (Q9UM54) Myosin VI 38 0.040
INCE_HUMAN (Q9NQS7) Inner centromere protein 38 0.040
YI73_AQUAE (O67720) Hypothetical protein AQ_1873 37 0.052
TPM_METEN (Q25456) Tropomyosin (Allergen Met e 1) (Met e I) 37 0.052
>CH02_HUMAN (O94905) Protein C8orf2 (UNQ2441/PRO9924/PRO5003)
Length = 339
Score = 348 bits (892), Expect = 1e-95
Identities = 174/304 (57%), Positives = 231/304 (75%), Gaps = 1/304 (0%)
Query: 5 LQVLVPSASPSF-KNTMAIVHQVPEGHVGVYWRGGALLKTITEPGFHMKMPFLTQFEPVQ 63
L +V AS F + + VH++ EGH+GVY+RGGALL + + PGFH+ +PF+T ++ VQ
Sbjct: 4 LGAVVAVASSFFCASLFSAVHKIEEGHIGVYYRGGALLTSTSGPGFHLMLPFITSYKSVQ 63
Query: 64 VTLQTDEVTDIPCGTKGGVMIVFGKIEVVNRLHKESVYETLLNYGVQYDKTWIYDKIHHE 123
TLQTDEV ++PCGT GGVMI F +IEVVN L +VY+ + NY YDK I++KIHHE
Sbjct: 64 TTLQTDEVKNVPCGTSGGVMIYFDRIEVVNFLVPNAVYDIVKNYTADYDKALIFNKIHHE 123
Query: 124 INQFCSSHSLQQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNF 183
+NQFCS H+LQ+VYI++FDQIDE +K ALQ D T APG+ I VRVTKPNIPE+IR N+
Sbjct: 124 LNQFCSVHTLQEVYIELFDQIDENLKLALQQDLTSMAPGLVIQAVRVTKPNIPEAIRRNY 183
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQE 243
E ME E+TK+LIA +KQKV EKEAET +K A+ EAEK A V++I QK+ EK++ ++
Sbjct: 184 ELMESEKTKLLIAAQKQKVVEKEAETERKKALIEAEKVAQVAEITYGQKVMEKETEKKIS 243
Query: 244 EIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTKIFFGDKIPN 303
EIE+A +LAREK+ ADA+ Y +K AEAN+LKLTPE+L+L ++IA+N+KI+FG IPN
Sbjct: 244 EIEDAAFLAREKAKADAECYTAMKIAEANKLKLTPEYLQLMKYKAIASNSKIYFGKDIPN 303
Query: 304 MILD 307
M +D
Sbjct: 304 MFMD 307
>TPM_ECHMU (Q95PU1) Tropomyosin
Length = 284
Score = 45.1 bits (105), Expect = 3e-04
Identities = 46/206 (22%), Positives = 92/206 (44%), Gaps = 17/206 (8%)
Query: 94 RLHKESVYETLLNYGVQY-DKTWIYDKIHHEINQFCSSHSLQQVYIDVFDQIDEKMKDAL 152
+L KE+ E +N Q +K ++K E+N + S Q +D + E +++A+
Sbjct: 12 KLEKENALEKAINLENQLKEKAKDFEKKEEEMNDWLSKVKNIQTEVDT---VQESLQEAI 68
Query: 153 QV--DCTRYAPGIEIIGVRVTKPN--IPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAE 208
+ + A E +T+ + E + ++ E TK+ A + + SE+ +
Sbjct: 69 SKLEETEKRATNAEAEVAAMTRRIRLLEEDFEQSSGRLTETSTKLDDASKAAEESERNRK 128
Query: 209 TMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKE 268
T++ +IS+ E+ A + + + E K +D+ R+ +E AR ++ + D R
Sbjct: 129 TLETRSISDDERMAQLEEQVKEAKYIAEDAERKYDE------AARRLAVTEVDLERAESR 182
Query: 269 AEANRLKLTPEFLELKFIESIANNTK 294
E + K+ EL+ + NN K
Sbjct: 183 LETSESKIVELEEELRI---VGNNMK 205
>YLX8_CAEEL (P46504) Hypothetical protein F23F12.8 in chromosome III
precursor
Length = 887
Score = 42.4 bits (98), Expect = 0.002
Identities = 37/145 (25%), Positives = 74/145 (50%), Gaps = 11/145 (7%)
Query: 134 QQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFE---QMEEER 190
Q+V ++ Q +E ++ L+V A +E RV + + +H E Q EE++
Sbjct: 442 QKVEMEQIRQQEEARQEQLRVLEEERARELE----RVRQEELER--QHQMEILRQQEEDQ 495
Query: 191 TKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMY 250
K + ++++ ++EAE + +M I E E N K++ E+K K + E+ +NA+Y
Sbjct: 496 KKKKLEKDREQREQQEAEELNRMII-EKEMKENKQKMI-EEKNKRKMLEKEMEDRQNAIY 553
Query: 251 LAREKSLADADFYRVIKEAEANRLK 275
E+ +A+ + + I+ E R++
Sbjct: 554 EEEERRIAEEERRKQIEIEERRRIQ 578
Score = 35.8 bits (81), Expect = 0.15
Identities = 26/98 (26%), Positives = 47/98 (47%), Gaps = 10/98 (10%)
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQE 243
++M+E + K++ K+K+ EKE E + E E+ +I E++ + + R+
Sbjct: 522 KEMKENKQKMIEEKNKRKMLEKEMEDRQNAIYEEEER-----RIAEEERRKQIEIEERRR 576
Query: 244 EIENAMYLAREKSLADA-----DFYRVIKEAEANRLKL 276
+ M E+S DA + R IKE+E R +L
Sbjct: 577 IQQQIMIATEERSRLDAMEREREMLRQIKESEKQRKEL 614
Score = 30.4 bits (67), Expect = 6.4
Identities = 29/118 (24%), Positives = 50/118 (41%), Gaps = 29/118 (24%)
Query: 180 RHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEA--------------------- 218
+ FE+ME+ER + ++++ +E E +K+ SE
Sbjct: 314 QEKFEKMEQERLR-----QEKEEKARELERRRKLEESETARQAELDRQATIYAEQERMAM 368
Query: 219 EKNANVSKILMEQKLSEKDSARRQE---EIENAMYLAREKSLADADFYRVIKEAEANR 273
E+N + +I +E+K E + R++E EI L R + RV +E EA R
Sbjct: 369 ERNRELERIRLEEKKRENERVRQEEIAMEISKIRELERLQLERQRKNERVRQELEAAR 426
>TPM_DERPT (O18416) Tropomyosin (Allergen Der p 10)
Length = 284
Score = 42.4 bits (98), Expect = 0.002
Identities = 32/115 (27%), Positives = 58/115 (49%), Gaps = 14/115 (12%)
Query: 185 QMEEERTKVLIA-IEKQKVSEKEAETMKKM----AISEAEKNANVSKILMEQKLSEKDSA 239
+ EER K+ A +E+ S E+E M+KM +I++ E+ + L E ++ +D+
Sbjct: 100 ERSEERLKIATAKLEEASQSADESERMRKMLEHRSITDEERMEGLENQLKEARMMAEDAD 159
Query: 240 RRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
R+ +E+ AR+ ++ +AD R + AE K+ EL+ + NN K
Sbjct: 160 RKYDEV------ARKLAMVEADLERAEERAETGESKIVELEEELRV---VGNNLK 205
>TPM_DERFA (Q23939) Tropomyosin (Allergen Der f 10) (Mag44)
Length = 284
Score = 42.0 bits (97), Expect = 0.002
Identities = 32/115 (27%), Positives = 58/115 (49%), Gaps = 14/115 (12%)
Query: 185 QMEEERTKVLIA-IEKQKVSEKEAETMKKM----AISEAEKNANVSKILMEQKLSEKDSA 239
+ EER K+ A +E+ S E+E M+KM +I++ E+ + L E ++ +D+
Sbjct: 100 ERSEERLKIATAKLEEASQSADESERMRKMLEHRSITDEERMDGLENQLKEARMMAEDAD 159
Query: 240 RRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
R+ +E+ AR+ ++ +AD R + AE K+ EL+ + NN K
Sbjct: 160 RKYDEV------ARKLAMVEADLERAEERAETGESKIVELEEELRV---VGNNLK 205
>M4K4_MOUSE (P97820) Mitogen-activated protein kinase kinase kinase
kinase 4 (EC 2.7.1.37) (MAPK/ERK kinase kinase kinase 4)
(MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase
HGK) (Nck interacting kinase)
Length = 1233
Score = 39.7 bits (91), Expect = 0.011
Identities = 43/192 (22%), Positives = 78/192 (40%), Gaps = 42/192 (21%)
Query: 134 QQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPE---------------S 178
+QV I + D ID K + D T Y E G + +PE +
Sbjct: 295 RQVRIQLKDHIDRTRKKRGEKDETEY----EYSGSEEEEEEVPEQEGEPSSIVNVPGEST 350
Query: 179 IRHNFEQMEEERTKVLIAIEKQKVSE----KEAETMKKMAISEAEKNANVSKILM----- 229
+R +F ++++E + A+ +Q++ + +E E K+ ++E +K K
Sbjct: 351 LRRDFLRLQQENKERSEALRRQQLLQEQQLREQEEYKRQLLAERQKRIEQQKEQRRRLEE 410
Query: 230 --------------EQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLK 275
EQ+ E++ RR EE+E E+ A+ + RV +E E R +
Sbjct: 411 QQRREREARRQQEREQRRREQEEKRRLEELERRRKEEEERRRAEEEKRRVEREQEYIRRQ 470
Query: 276 LTPEFLELKFIE 287
L E L+ ++
Sbjct: 471 LEEEQRHLEILQ 482
>HFLC_HAEIN (P44545) HflC protein (EC 3.4.-.-)
Length = 295
Score = 39.7 bits (91), Expect = 0.011
Identities = 55/249 (22%), Positives = 98/249 (39%), Gaps = 45/249 (18%)
Query: 44 ITEPGFHMKMPFLTQFEPVQVTLQTDEVTDIPCGTKGGVMIVFGKIEVVNRLHKESVYET 103
+ EPG H K+P + + V D T G F +E + L V
Sbjct: 47 VYEPGLHFKVPLIDSIK----------VLDARIRTLDGSATRFVTVEKKDLLVDSYVKWK 96
Query: 104 LLNYGVQYDKTWIYD----------KIHHEINQFCSSHSLQQVYIDVFDQIDEKMKDALQ 153
+ ++G Y T D K++ + S +++ + ++ E K AL
Sbjct: 97 ISDFGRFYTSTGGGDYAQAANLLSRKVNDRLRSEIGSRTIKDIVSGTRGELMEGAKKALS 156
Query: 154 VDCTRYAP-GIEIIGVRVTKPNIPESIRHN-FEQMEEERTKVLIAIEKQKVSEKEAETMK 211
A GIE+I VRV + N+P+ + + +++M ER V E ++ +
Sbjct: 157 SGQDSTAELGIEVIDVRVKQINLPDEVSSSIYQRMRAERDAV--------AREHRSQGKE 208
Query: 212 KMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADA--------DFY 263
K A +A+ + V+ IL ++ + +E+ + A K +DA F
Sbjct: 209 KAAFIQADVDRKVTLIL-------ANANKTAQELRGSGDAAAAKLYSDAFAQEPQFFTFV 261
Query: 264 RVIKEAEAN 272
R +K EA+
Sbjct: 262 RSLKAYEAS 270
>TPM_LEPDS (Q9NFZ4) Tropomyosin (Allergen Lep d 10)
Length = 284
Score = 39.3 bits (90), Expect = 0.014
Identities = 31/118 (26%), Positives = 58/118 (48%), Gaps = 16/118 (13%)
Query: 184 EQMEEERTKVLIA---IEKQKVSEKEAETMKKM----AISEAEKNANVSKILMEQKLSEK 236
E +E ++ IA +E+ S E+E M+KM +I++ E+ + L E ++ +
Sbjct: 97 EDLERSEGRLKIATSKLEEASQSADESERMRKMLEHRSITDEERMEGLESQLKEARMMAE 156
Query: 237 DSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
D+ R+ +E+ AR+ ++ +AD R + AE K+ EL+ + NN K
Sbjct: 157 DADRKYDEV------ARKLAMVEADLERAEERAETGESKIVELEEELRV---VGNNLK 205
>M4K4_HUMAN (O95819) Mitogen-activated protein kinase kinase kinase
kinase 4 (EC 2.7.1.37) (MAPK/ERK kinase kinase kinase 4)
(MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase
HGK) (Nck interacting kinase)
Length = 1239
Score = 39.3 bits (90), Expect = 0.014
Identities = 43/192 (22%), Positives = 78/192 (40%), Gaps = 42/192 (21%)
Query: 134 QQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPE---------------S 178
+QV I + D ID K + D T Y E G + +PE +
Sbjct: 296 RQVRIQLKDHIDRTRKKRGEKDETEY----EYSGSEEEEEEVPEQEGEPSSIVNVPGEST 351
Query: 179 IRHNFEQMEEERTKVLIAIEKQKVSE----KEAETMKKMAISEAEKNANVSKILM----- 229
+R +F ++++E + A+ +Q++ + +E E K+ ++E +K K
Sbjct: 352 LRRDFLRLQQENKERSEALRRQQLLQEQQLREQEEYKRQLLAERQKRIEQQKEQRRRLEE 411
Query: 230 --------------EQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLK 275
EQ+ E++ RR EE+E E+ A+ + RV +E E R +
Sbjct: 412 QQRREREARRQQEREQRRREQEEKRRLEELERRRKEEEERRRAEEEKRRVEREQEYIRRQ 471
Query: 276 LTPEFLELKFIE 287
L E L+ ++
Sbjct: 472 LEEEQRHLEVLQ 483
>KI21_STRPU (P46871) Kinesin-II 95 kDa subunit (KRP-85/95 95 kDa
subunit)
Length = 742
Score = 38.9 bits (89), Expect = 0.018
Identities = 28/108 (25%), Positives = 51/108 (46%), Gaps = 2/108 (1%)
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
E I N + EE+ K+L ++K++ K+ K+M E + A SK+L+ K
Sbjct: 423 EKIMANQSMIAEEKQKLLSEVQKRQGEIKKEHQQKEML--EGKIKAMESKLLVGGKSIVD 480
Query: 237 DSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELK 284
+ +Q +IE L E+ + D R +KE + +++ F L+
Sbjct: 481 HTNEQQRKIEEQRLLLAEEKNRERDMERKLKEQDDKTVEIEGTFSSLQ 528
>DNAK_VIBCH (O34241) Chaperone protein dnaK (Heat shock protein 70)
(Heat shock 70 kDa protein) (HSP70)
Length = 635
Score = 38.1 bits (87), Expect = 0.031
Identities = 35/142 (24%), Positives = 63/142 (43%), Gaps = 18/142 (12%)
Query: 147 KMKDALQVDCTRYAPGIEIIGVRVTK-----PNIPESIRHNFEQMEEERTKVLIAI---- 197
++KD L +D T + GIE +G +TK IP F E+ ++ V I +
Sbjct: 384 EVKDVLLLDVTPLSLGIETMGGVMTKLIEKNTTIPTKANQVFSTAEDNQSAVTIHVLQGE 443
Query: 198 EKQKVSEKEAETMKKMAISEAEKNANVSKILMEQK------LSEKDSARRQEEIENAMYL 251
KQ + K I+ A + +++ + +S KD +Q E + +
Sbjct: 444 RKQAMYNKSLGQFNLEGINPAPRGMPQIEVIFDLDADGILHVSAKD---KQTGKEQKITI 500
Query: 252 AREKSLADADFYRVIKEAEANR 273
L+DA+ ++++EAEAN+
Sbjct: 501 QASGGLSDAEIEKMVQEAEANK 522
>YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I
Length = 1085
Score = 37.7 bits (86), Expect = 0.040
Identities = 29/146 (19%), Positives = 70/146 (47%), Gaps = 11/146 (7%)
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSAR--- 240
++++ E+ K +E+QK EK+ + ++ + + ++ A+ K+ EQ+L E++ R
Sbjct: 644 QRLKREQEKKQQELERQKREEKQKQKEREKKLKKQQQEADREKMAREQRLREEEEKRILE 703
Query: 241 ---RQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIE---SIANNTK 294
R+E+++ R + L + + KE K+ F + E +++
Sbjct: 704 ERKRREKLDKEEEERRRRELLEKESEE--KERRLREAKIAAFFAPNQTKEGSDGCTTSSQ 761
Query: 295 IFFGDKIPNMILDQRLLGNFLVEEVP 320
+ +K +++ D+ L + L++ VP
Sbjct: 762 LGLFEKKGDLVNDEDKLSSHLLDSVP 787
>TPM_SCHHA (Q26503) Tropomyosin
Length = 284
Score = 37.7 bits (86), Expect = 0.040
Identities = 47/208 (22%), Positives = 87/208 (41%), Gaps = 15/208 (7%)
Query: 88 KIEVVNRLHKESVYETLLNYGVQYDKTWIYDKIHHEINQFCSSHSLQQVYID-VFDQIDE 146
K+E N + + YE LL K +K EI + + Q+ D V + + E
Sbjct: 12 KLEKENAMERAVQYEELLK-----KKEEEREKRESEIAELNTKMKQAQIDCDEVQETLQE 66
Query: 147 KMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKE 206
+M + D E+ + + E + + ++ E TK+ A + + SE+
Sbjct: 67 QMNKLEETDKRATNAEAEVAAMTRRIRLLEEDLEVSSSRLTETLTKLEEASKTAEESERG 126
Query: 207 AETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVI 266
+ ++ +I++ E+ + E K +D+ R+ +E AR+ ++A+ DF R
Sbjct: 127 RKDLEIRSIADDERLNQLEDQQKEAKYIAEDADRKYDE------AARKLAIAEVDFKRAE 180
Query: 267 KEAEANRLKLTPEFLELKFIESIANNTK 294
EA K+ EL+ I NN K
Sbjct: 181 ARLEAAESKIVELEEELRV---IGNNMK 205
>TPM_CHAFE (Q9N2R3) Tropomyosin (Allergen Cha f 1) (Cha f I)
(Fragment)
Length = 264
Score = 37.7 bits (86), Expect = 0.040
Identities = 37/153 (24%), Positives = 73/153 (47%), Gaps = 18/153 (11%)
Query: 143 QIDEKMKDALQVDCTRYAPG-IEIIGVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQK 201
++DEK K ALQ A G + + R+ P E + + E++ TK+ A +
Sbjct: 70 KLDEKEK-ALQ-----NAEGEVAALNRRIQLPE--EDLERSEERLNTATTKLAEASQAAD 121
Query: 202 VSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADAD 261
SE+ + ++ ++S+ E+ + L E + +++ R+ +E+ AR+ ++ +AD
Sbjct: 122 ESERMRKVLENRSLSDEERMDALENQLKEARFLAEEADRKYDEV------ARKLAMVEAD 175
Query: 262 FYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
R + AE+ K+ EL+ + NN K
Sbjct: 176 LERAEERAESGESKIVELEEELRV---VGNNLK 205
>TPM1_CAEEL (Q22866) Tropomyosin isoforms 1/2
Length = 284
Score = 37.7 bits (86), Expect = 0.040
Identities = 30/118 (25%), Positives = 57/118 (47%), Gaps = 16/118 (13%)
Query: 184 EQMEEERTKVLIAIEKQKV-------SEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
E++E ++ IA EK + SE+ + M+ ++ + E+ V L E +L +
Sbjct: 97 EELERAEERLKIATEKLEEATHNVDESERVRKVMENRSLQDEERANTVEAQLKEAQLLAE 156
Query: 237 DSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
++ R+ +E+ AR+ ++ +AD R + AEA K+ EL+ + NN K
Sbjct: 157 EADRKYDEV------ARKLAMVEADLERAEERAEAGENKIVELEEELRV---VGNNLK 205
>MYO6_MOUSE (Q64331) Myosin VI
Length = 1265
Score = 37.7 bits (86), Expect = 0.040
Identities = 33/143 (23%), Positives = 67/143 (46%), Gaps = 15/143 (10%)
Query: 138 IDVFDQIDEKMKDALQVDCTRYAPGIEI----IGVRVTKPNIP-ESIRHNFEQMEEERTK 192
+D F+++ +KD + + R +EI + + T + E I+ ++ + +
Sbjct: 853 LDKFNEVVSALKDG-KPEVNRQIKNLEISIDALMAKFTSTMMTREQIQKEYDALVKSSED 911
Query: 193 VLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLA 252
+L A++K+K E+EAE ++++ E EK ++ E + RR+EE E M L
Sbjct: 912 LLSALQKKKQQEEEAERLRRIQ-EEMEKE--------RKRREEDEERRRKEEEERRMKLE 962
Query: 253 REKSLADADFYRVIKEAEANRLK 275
E + R +E + R++
Sbjct: 963 MEPKRKQEEEERKKREDDEKRIQ 985
>MYO6_HUMAN (Q9UM54) Myosin VI
Length = 1285
Score = 37.7 bits (86), Expect = 0.040
Identities = 26/103 (25%), Positives = 50/103 (48%), Gaps = 9/103 (8%)
Query: 177 ESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEK 236
E I+ ++ + + ++L A++K+K E+EAE ++++ E EK ++ E
Sbjct: 893 EQIQKEYDALVKSSEELLSALQKKKQQEEEAERLRRIQ-EEMEKE--------RKRREED 943
Query: 237 DSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPE 279
+ RR+EE E M L E + R +E + R++ E
Sbjct: 944 EKRRRKEEEERRMKLEMEAKRKQEEEERKKREDDEKRIQAEVE 986
>INCE_HUMAN (Q9NQS7) Inner centromere protein
Length = 919
Score = 37.7 bits (86), Expect = 0.040
Identities = 36/137 (26%), Positives = 64/137 (46%), Gaps = 15/137 (10%)
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQM----EEERTKVLIAIE 198
++D K K+ +++ R E ++ + + E R E++ EE KVL A E
Sbjct: 527 RMDPKEKERQRLENLRRKEEAE----QLRRQKVEEDKRRRLEEVKLKREERLRKVLQARE 582
Query: 199 K--QKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAREKS 256
+ Q EK+ + +K A + + + L E+K +K +A++ EE+E AR K
Sbjct: 583 RVEQMKEEKKKQIEQKFAQIDEKTEKAKEERLAEEKAKKKAAAKKMEEVE-----ARRKQ 637
Query: 257 LADADFYRVIKEAEANR 273
DA R +++ E R
Sbjct: 638 EEDARRLRWLQQEEEER 654
Score = 32.7 bits (73), Expect = 1.3
Identities = 32/140 (22%), Positives = 64/140 (44%), Gaps = 9/140 (6%)
Query: 141 FDQIDEKMKDA-----LQVDCTRYAPGIEIIGVRVTKPNIPESIRHNFEQMEEERTKVLI 195
F QIDEK + A + + A ++ V + ++ R + Q EEE +
Sbjct: 599 FAQIDEKTEKAKEERLAEEKAKKKAAAKKMEEVEARRKQEEDARRLRWLQQEEEERRHQE 658
Query: 196 AIEKQKVSEKEAETMKKMAISE--AEKNANVSKILMEQKLSEKDSARRQEEIENAMYLAR 253
++K+K E+E E ++K A ++ AE+ + ++ ++ RR++E +
Sbjct: 659 LLQKKK--EEEQERLRKAAEAKRLAEQREQERREQERREQERREQERREQERREQERREQ 716
Query: 254 EKSLADADFYRVIKEAEANR 273
E+ LA+ + R + +A R
Sbjct: 717 ERQLAEQERRREQERLQAER 736
>YI73_AQUAE (O67720) Hypothetical protein AQ_1873
Length = 407
Score = 37.4 bits (85), Expect = 0.052
Identities = 47/175 (26%), Positives = 77/175 (43%), Gaps = 20/175 (11%)
Query: 123 EINQFCSSHSLQQVYIDVFDQIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIPESIRHN 182
EIN+ S Q Y+ ++D++ VD P +++ +R + P E IR
Sbjct: 58 EINELYSKKFKNQKYVILYDEL-------FPVDPKNILPPNKVLVLRDSPPE--EIIRRA 108
Query: 183 FEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEK--NANVSKILMEQKLSEKDS-- 238
E+ +EE + I I E+E +T KK E E+ A V + ++KL++ +
Sbjct: 109 LEEKKEETKEEGIEIGASVPVEEELKTEKKEEKVEKEETGKAKVLVVSFDKKLTDNITNA 168
Query: 239 --ARRQEEIENAMYLAREKSL-ADADFYRVIKE--AEANRLKL--TPEFLELKFI 286
+ E+ + +EK L AD Y I AE N ++L P+ E FI
Sbjct: 169 LKGKFSVEVVRNIKSVKEKGLEADIIVYDAISGSIAEKNLMELAEVPQLREKHFI 223
Score = 32.0 bits (71), Expect = 2.2
Identities = 35/137 (25%), Positives = 62/137 (44%), Gaps = 17/137 (12%)
Query: 172 KPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQ 231
+ IPE + E+++EE+ E ++ EK AIS EK ++L E
Sbjct: 284 REEIPEEVPQEVEEIKEEKP------EMSEIKEKTE------AISTKEKTPEAVQVLAE- 330
Query: 232 KLSEKDSARR--QEEIENAMYLAREKSLADA-DFYRVIKEAEANRLKLTPEFLELKFIES 288
KLS++ R E I ++ RE+ A+ D+ + + E+ R ++ F E+ +
Sbjct: 331 KLSDEKFIRSLLVEAISKELHGVREEIKAEVRDYIKNVLES-IIREEIERAFAEVGVAKI 389
Query: 289 IANNTKIFFGDKIPNMI 305
I TK +KI ++
Sbjct: 390 IREETKRIVEEKIKELL 406
>TPM_METEN (Q25456) Tropomyosin (Allergen Met e 1) (Met e I)
Length = 274
Score = 37.4 bits (85), Expect = 0.052
Identities = 26/120 (21%), Positives = 57/120 (46%), Gaps = 9/120 (7%)
Query: 175 IPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLS 234
+ E + + E++ TK+ A + SE+ + ++ ++S+ E+ + L E +
Sbjct: 85 LEEDLERSEERLNTATTKLAEASQAADESERMRKVLENRSLSDEERMDALENQLKEARFL 144
Query: 235 EKDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTK 294
+++ R+ +E+ AR+ ++ +AD R + AE K+ EL+ + NN K
Sbjct: 145 AEEADRKYDEV------ARKLAMVEADLERAEERAETGESKIVELEEELRV---VGNNLK 195
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,721,492
Number of Sequences: 164201
Number of extensions: 1523640
Number of successful extensions: 7194
Number of sequences better than 10.0: 331
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 307
Number of HSP's that attempted gapping in prelim test: 6663
Number of HSP's gapped (non-prelim): 758
length of query: 331
length of database: 59,974,054
effective HSP length: 111
effective length of query: 220
effective length of database: 41,747,743
effective search space: 9184503460
effective search space used: 9184503460
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146567.11


