

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.10 + phase: 0
(393 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
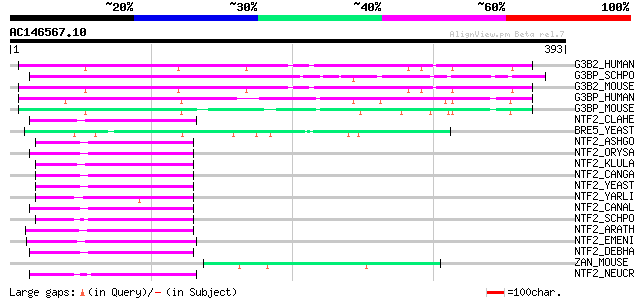
Sequences producing significant alignments: (bits) Value
G3B2_HUMAN (Q9UN86) Ras-GTPase-activating protein binding protei... 119 2e-26
G3BP_SCHPO (O94260) Putative G3BP-like protein 116 9e-26
G3B2_MOUSE (P97379) Ras-GTPase-activating protein binding protei... 115 2e-25
G3BP_HUMAN (Q13283) Ras-GTPase-activating protein binding protei... 114 6e-25
G3BP_MOUSE (P97855) Ras-GTPase-activating protein binding protei... 112 1e-24
NTF2_CLAHE (Q8NJ52) Nuclear transport factor 2 (NTF-2) (Allergen... 61 4e-09
BRE5_YEAST (P53741) UBP3-associated protein BRE5 59 3e-08
NTF2_ASHGO (Q75AA5) Nuclear transport factor 2 (NTF-2) 58 4e-08
NTF2_ORYSA (Q9XJ54) Nuclear transport factor 2 (NTF-2) 57 1e-07
NTF2_KLULA (Q6CQX4) Nuclear transport factor 2 (NTF-2) 57 1e-07
NTF2_CANGA (Q6FRC6) Nuclear transport factor 2 (NTF-2) 56 2e-07
NTF2_YEAST (P33331) Nuclear transport factor 2 (NTF-2) (Nuclear ... 54 5e-07
NTF2_YARLI (Q6CC82) Nuclear transport factor 2 (NTF-2) 54 9e-07
NTF2_CANAL (Q9P926) Nuclear transport factor 2 (NTF-2) 52 2e-06
NTF2_SCHPO (Q10100) Nuclear transport factor 2 (NTF-2) 52 3e-06
NTF2_ARATH (Q9C7F5) Nuclear transport factor 2 (NTF-2) 52 3e-06
NTF2_EMENI (Q96VN3) Nuclear transport factor 2 (NTF-2) 52 3e-06
NTF2_DEBHA (Q6BWC0) Nuclear transport factor 2 (NTF-2) 51 4e-06
ZAN_MOUSE (O88799) Zonadhesin precursor 50 1e-05
NTF2_NEUCR (P87102) Nuclear transport factor 2 (NTF-2) 49 3e-05
>G3B2_HUMAN (Q9UN86) Ras-GTPase-activating protein binding protein 2
(GAP SH3-domain binding protein 2) (G3BP-2)
Length = 482
Score = 119 bits (297), Expect = 2e-26
Identities = 108/394 (27%), Positives = 166/394 (41%), Gaps = 37/394 (9%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT---MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ P+ +HRFY +S D + V +I
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNKAPEYLHRFYGRNSSYVHGGVDASGKPQEAVYGQNDIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ---DK 120
K+ SL ++ ++ DA + ++GV+V V G L+ + +RKF Q+F LAP+
Sbjct: 63 HKVLSLNFSECHTKIRHVDAHATLSDGVVVQVMGLLSNSGQPERKFMQTFVLAPEGSVPN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVH------VPEVIPPT 174
FYV ND+FRY D + DE + P PEPV E P T
Sbjct: 123 KFYVHNDMFRYEDEVFGDSEPELDEESEDEVEEEQEERQPSPEPVQENANSGYYEAHPVT 182
Query: 175 QTV-IPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKE 233
+ P ++ P ++T+T +E+ +E L E + + + V
Sbjct: 183 NGIEEPLEESSHEPEPEPESETKT---EELKPQVEEKNL---EELEEKSTTPPPAEPVSL 236
Query: 234 PEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAV--HPPRVHSVP- 290
P++P AS T ++ P S AP AKP V PPRV
Sbjct: 237 PQEPPKAFSWASVTSKNLPPSGTVSSSGIPSHVKAPVSQPRVEAKPEVQSQPPRVREQRP 296
Query: 291 ------APEAPTPNMDIPLEKNNENAGR------AHAIFVANLPMSATVEQLDRAFKKFG 338
P P P +E+N+ + R +H +FV NLP +L F FG
Sbjct: 297 RERPGFPPRGPRPGRG-DMEQNDSDNRRIIRYPDSHQLFVGNLPHDIDENELKEFFMSFG 355
Query: 339 PIKRDGIQVRSNKGSC--FGFVEFESAASMQSAL 370
+ I + G FGFV F+ + +Q L
Sbjct: 356 NVVELRINTKGVGGKLPNFGFVVFDDSEPVQRIL 389
>G3BP_SCHPO (O94260) Putative G3BP-like protein
Length = 434
Score = 116 bits (291), Expect = 9e-26
Identities = 101/373 (27%), Positives = 181/373 (48%), Gaps = 22/373 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
+G FV++YY+ L+++P+++H FY S + +E +++ EI KI L++ +
Sbjct: 18 IGWMFVQEYYTYLNKEPNRLHCFYTKKSTLIHGDEGESISLCHGQQEIHNKILDLDFQNC 77
Query: 75 RVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFRYVDA 134
+V + + D+ S N G+++ V G ++ + RKFAQ+FFLA Q G++VLND+FR++
Sbjct: 78 KVLISNVDSLASSNGGIVIQVLGEMSNKGKLSRKFAQTFFLAEQPNGYFVLNDIFRFLRE 137
Query: 135 YKSIDIESVPANDAD-ESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIA 193
+ ES A + + + SE + H+P P A T +I+
Sbjct: 138 DVEEEEESPDAVEKEKKDVASEPYVNGVQSQEHLPSAKEEGHYQDPAATENNFATAALIS 197
Query: 194 -DTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEK-----VASNT 247
+T+++ +++P E + VTE +P + V + + +++ E TS K AS+
Sbjct: 198 NETDSLNQATLAVP-EEPVIQVTEASVP-SFVSQQENQLQD-EALTSNSKNADAIGASDA 254
Query: 248 QEDTPKKSFASIVNALKDNSAPFHLRASPAKPA-VHPPRVHSVPAPEAPTPNMDIPLEKN 306
T KS+A ++ N +AS + A V V A + P P ++
Sbjct: 255 NVATAPKSWADLI---ARNHPDVKSQASVSSTASTTGQTVKGVNADQTQQPT--APYTQS 309
Query: 307 NENAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASM 366
NE ++FV N+P + L A FGP+K I+ KG+ +V+F + +
Sbjct: 310 NELL--ETSVFVKNIPPETSDVSLKSAMSIFGPVK--AIEFARRKGT--AYVDFVNHECV 363
Query: 367 QSALEVWSISINS 379
Q AL ++ IN+
Sbjct: 364 QLALNKKTLQINN 376
>G3B2_MOUSE (P97379) Ras-GTPase-activating protein binding protein 2
(GAP SH3-domain binding protein 2) (G3BP-2)
Length = 482
Score = 115 bits (288), Expect = 2e-25
Identities = 106/394 (26%), Positives = 165/394 (40%), Gaps = 37/394 (9%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT---MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ P+ +HRFY +S D + V +I
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNKAPEYLHRFYGRNSSYVHGGVDASGKPQEAVYGQNDIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ---DK 120
K+ SL ++ ++ DA + ++GV+V V G L+ + +RKF Q+F LAP+
Sbjct: 63 HKVLSLNFSECHTKIRHVDAHATLSDGVVVQVMGLLSNSGQPERKFMQTFVLAPEGSVPN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVH------VPEVIPPT 174
FYV ND+FRY D + DE + P PEPV + P T
Sbjct: 123 KFYVHNDMFRYEDEVFGDSEPELDEESEDEVEEEQEDRQPSPEPVQENANSAYYDAHPVT 182
Query: 175 QTV-IPTAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKE 233
+ P ++ P ++T+T +E+ +E L E + + +
Sbjct: 183 NGIEEPLEESSHEPEPEPESETKT---EELKPQVEEKHL---EELEEKSATPPPAEPASL 236
Query: 234 PEQPTSIEKVASNTQEDTPKKSFASIVNALKDNSAPFHLRASPAKPAV--HPPRVHSVP- 290
P++P AS T ++ P S AP AKP V PPRV
Sbjct: 237 PQEPPKAFSWASVTSKNLPPSGTVSSSGIPPHVKAPVSQPRVDAKPEVQSQPPRVREQRP 296
Query: 291 ------APEAPTPNMDIPLEKNNENAGR------AHAIFVANLPMSATVEQLDRAFKKFG 338
P P P +E+N+ + R +H +FV NLP +L F FG
Sbjct: 297 RERPGFPPRGPRPGRG-DMEQNDSDNRRIIRYPDSHQLFVGNLPHDIDENELKEFFMSFG 355
Query: 339 PIKRDGIQVRSNKGSC--FGFVEFESAASMQSAL 370
+ I + G FGFV F+ + +Q L
Sbjct: 356 NVVELRINTKGVGGKLPNFGFVVFDDSEPVQRIL 389
>G3BP_HUMAN (Q13283) Ras-GTPase-activating protein binding protein 1
(GAP SH3-domain binding protein 1) (G3BP-1) (DNA
helicase VIII) (HDH-VIII)
Length = 466
Score = 114 bits (284), Expect = 6e-25
Identities = 106/414 (25%), Positives = 173/414 (41%), Gaps = 71/414 (17%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFY--HDSSVMSRPEEDGTMT-TVTTTAEID 63
++ P+P +VG FV QYY++L++ PD +HRFY + S V + +G V EI
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNQAPDMLHRFYGKNSSYVHGGLDSNGKPADAVYGQKEIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK--- 120
+K+ S +T+ ++ DA + N+GV+V V G L+ + R+F Q+F LAP+
Sbjct: 63 RKVMSQNFTNCHTKIRHVDAHATLNDGVVVQVMGLLSNNNQALRRFMQTFVLAPEGSVAN 122
Query: 121 GFYVLNDVFRYVD-AYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIP 179
FYV ND+FRY D + E ++ + P E TPE V+P
Sbjct: 123 KFYVHNDIFRYQDEVFGGFVTEPQEESEEEVEEPEERQQTPE---------------VVP 167
Query: 180 TAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHH-VKEPEQPT 238
V D E + + V+ P + + + PV+ ++E V E P
Sbjct: 168 DDSGTFYDQAVVSNDMEEHLEEPVAEPEPDPEPEPEQE--PVSEIQEEKPEPVLEETAPE 225
Query: 239 SIEK--------VASNTQEDTPKKSFASIVN-------ALKDNSAPFHLRASPAKPAVHP 283
+K +A QED S+AS+ + A+ P H+ PA
Sbjct: 226 DAQKSSSPAPADIAQTVQEDLRTFSWASVTSKNLPPSGAVPVTGIPPHVVKVPASQPRPE 285
Query: 284 PRVHSVPAPEAPTPNMDIPLEKNN----------ENAGR--------------AHAIFVA 319
+ S P+ P + + ++ N AG +H +F+
Sbjct: 286 SKPESQIPPQRPQRDQRVREQRINIPPQRGPRPIREAGEQGDIEPRRMVRHPDSHQLFIG 345
Query: 320 NLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS---CFGFVEFESAASMQSAL 370
NLP +L F+ +G + +++R N G FGFV F+ + +Q L
Sbjct: 346 NLPHEVDKSELKDFFQSYGNV----VELRINSGGKLPNFGFVVFDDSEPVQKVL 395
>G3BP_MOUSE (P97855) Ras-GTPase-activating protein binding protein 1
(GAP SH3-domain binding protein 1) (G3BP-1) (DNA
helicase VIII) (HDH-VIII)
Length = 465
Score = 112 bits (281), Expect = 1e-24
Identities = 109/412 (26%), Positives = 166/412 (39%), Gaps = 69/412 (16%)
Query: 7 VQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGT---MTTVTTTAEID 63
++ P+P +VG FV QYY++L++ PD +HRFY +S + D V EI
Sbjct: 3 MEKPSPLLVGREFVRQYYTLLNQAPDMLHRFYGKNSSYAHGGLDSNGKPADAVYGQKEIH 62
Query: 64 KKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDK--- 120
+K+ S +T+ ++ DA + N+GV+V V G L+ + R+F Q+F LAP+
Sbjct: 63 RKVMSQNFTNCHTKIRHVDAHATLNDGVVVQVMGLLSNNNQALRRFMQTFVLAPEGSVAN 122
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT 180
FYV ND+FRY D E + SE + E PEV+P
Sbjct: 123 KFYVHNDIFRYQD-------EVFGGFVTEPQEESEEEVEEPEERQQTPEVVPDDSGTFYD 175
Query: 181 AQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVK-EPEQPTS 239
QTV D E + + V P + PV+ ++E E P
Sbjct: 176 --------QTVSNDLEEHLEEPVVEPEPEPEPEPEPE--PVSDIQEDKPEAALEEAAPDD 225
Query: 240 IEKVASNT-------QEDTPKKSFASIVNALKDNSAPFHLRASP---AKPAVHPPRVHSV 289
++K S QED S+AS+ + S + +P K PR S
Sbjct: 226 VQKSTSPAPADVAPAQEDLRTFSWASVTSKNLPPSGAVPVTGTPPHVVKVPASQPRPESK 285
Query: 290 PAPEAPT--PNMDIPLEKNNEN------------AGR--------------AHAIFVANL 321
P + P P D + + N AG +H +F+ NL
Sbjct: 286 PDSQIPPQRPQRDQRVREQRINIPPQRGPRPIREAGEPGDVEPRRMVRHPDSHQLFIGNL 345
Query: 322 PMSATVEQLDRAFKKFGPIKRDGIQVRSNKGS---CFGFVEFESAASMQSAL 370
P +L F+ FG + +++R N G FGFV F+ + +Q L
Sbjct: 346 PHEVDKSELKDFFQNFGNV----VELRINSGGKLPNFGFVVFDDSEPVQKVL 393
>NTF2_CLAHE (Q8NJ52) Nuclear transport factor 2 (NTF-2) (Allergen
Cla h ?)
Length = 125
Score = 61.2 bits (147), Expect = 4e-09
Identities = 39/120 (32%), Positives = 61/120 (50%), Gaps = 7/120 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
+ F E YY D Q+ Y ++S+++ + + TA I K+Q L +
Sbjct: 7 IAQQFTEFYYKTFDTDRAQLAPLYRENSMLTFEQ-----SPFLGTANIVGKLQELPFQRI 61
Query: 75 RVEVLSADAQPSYNNG-VMVVVTGCLTGTDNIK-RKFAQSFFLAPQDKGFYVLNDVFRYV 132
+V + DAQPS +G ++VVV+G L + + + Q+F L P D +YV NDVFR V
Sbjct: 62 EHQVATVDAQPSNESGGILVVVSGALLVEEERRPMSYTQTFQLLPADGAYYVFNDVFRLV 121
>BRE5_YEAST (P53741) UBP3-associated protein BRE5
Length = 515
Score = 58.5 bits (140), Expect = 3e-08
Identities = 80/365 (21%), Positives = 143/365 (38%), Gaps = 68/365 (18%)
Query: 11 TPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVM--------SRPEEDGTMTTVTTT--- 59
T Q + AF++ YY + DP ++ FY ++ + S E+D + TV T
Sbjct: 4 TVQDICFAFLQNYYERMRTDPSKLAYFYASTAELTHTNYQSKSTNEKDDVLPTVKVTGRE 63
Query: 60 ------AEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSF 113
+ D K++SL+ +++ + + ++++ TG + T KF Q+F
Sbjct: 64 NINKFFSRNDAKVRSLK---LKLDTIDFQYTGHLHKSILIMATGEMFWTGTPVYKFCQTF 120
Query: 114 FLAPQDKG--FYVLNDVFRYV-DAYKSIDIESVPANDADESAPSEAI----------ITP 160
L P G F + ND+ R++ +++K + + ++E A+ +
Sbjct: 121 ILLPSSNGSTFDITNDIIRFISNSFKPYVLTDASLSQSNEENSVSAVEEDKIRHESGVEK 180
Query: 161 EPEPVHVPEVIPP-------TQTVIPTAQT-------------VIPPTQTVIADTETIIS 200
E E PE+ P T PT + IP T+ TET+ S
Sbjct: 181 EKEKEKSPEISKPKAKKETVKDTTAPTESSTQEKPIVDHSQPRAIPVTKESKIHTETVPS 240
Query: 201 KEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPT-----SIEKVAS-------NTQ 248
++ ++S TE + V + E SH ++ P S+E + + N +
Sbjct: 241 STKGNHKQD-EVS-TEELGNVTKLNEKSHKAEKKAAPIKTKEGSVEAINAVNNSSLPNGK 298
Query: 249 EDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRV-HSVPAPEAPTPNMDIPLEKNN 307
E + +K V + P + S AK + + S P P A P M + N
Sbjct: 299 EVSDEKPVPGGVKEAETEIKPIEPQVSDAKESGNNASTPSSSPEPVANPPKMTWASKLMN 358
Query: 308 ENAGR 312
EN+ R
Sbjct: 359 ENSDR 363
>NTF2_ASHGO (Q75AA5) Nuclear transport factor 2 (NTF-2)
Length = 125
Score = 58.2 bits (139), Expect = 4e-08
Identities = 36/114 (31%), Positives = 59/114 (51%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F E YY+ D Q+ Y D S+++ + +I +K+ SL + + +
Sbjct: 12 FTEFYYNQFDTDRSQLGNLYRDQSMLTFETSQ-----LQGAKDIVEKLVSLPFQKVQHRI 66
Query: 79 LSADAQPSYNNG-VMVVVTGCLTGTDNIK-RKFAQSFFLAPQDKGFYVLNDVFR 130
+ DAQP+ NG V+V++TG L D ++F+Q F L P+ +YV ND+FR
Sbjct: 67 TTLDAQPASPNGDVLVMITGDLLIDDEQNAQRFSQVFHLMPEGNSYYVFNDIFR 120
>NTF2_ORYSA (Q9XJ54) Nuclear transport factor 2 (NTF-2)
Length = 122
Score = 56.6 bits (135), Expect = 1e-07
Identities = 40/118 (33%), Positives = 56/118 (46%), Gaps = 7/118 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
V AFVE YY + + Y D S+++ + A I K+ SL +
Sbjct: 6 VAKAFVEHYYRTFDTNRPALVSLYQDGSMLTFEGQQ-----FLGAAAIAGKLGSLPFAQC 60
Query: 75 RVEVLSADAQPSY-NNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
++ + D QPS G++V V+G L TG D KF+Q F L P FYV ND+FR
Sbjct: 61 HHDINTVDCQPSGPQGGMLVFVSGSLRTGPDEHPLKFSQMFQLLPAGGNFYVQNDMFR 118
>NTF2_KLULA (Q6CQX4) Nuclear transport factor 2 (NTF-2)
Length = 125
Score = 56.6 bits (135), Expect = 1e-07
Identities = 35/114 (30%), Positives = 59/114 (51%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F E YY+ D Q+ Y + S+++ T + +I +K+ SL + +
Sbjct: 12 FTEFYYNQFDSDRTQLGNLYREQSMLTFET-----TQLQGAKDIVEKLVSLPFQKVAHRI 66
Query: 79 LSADAQPSYNNG-VMVVVTG-CLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
+ DAQP+ NG V+V++TG L + ++F+Q F L P+ +YV ND+FR
Sbjct: 67 TTLDAQPASPNGDVLVMITGDLLIDEEQNPQRFSQVFHLMPEGSSYYVYNDIFR 120
>NTF2_CANGA (Q6FRC6) Nuclear transport factor 2 (NTF-2)
Length = 125
Score = 55.8 bits (133), Expect = 2e-07
Identities = 36/114 (31%), Positives = 56/114 (48%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F E YY+ D Q+ Y D S+++ + I +K+ SL + +
Sbjct: 12 FTEFYYNQFDSDRSQLGNLYRDESMLTFETSQ-----LQGAKSIVEKLVSLPFQKVAHRI 66
Query: 79 LSADAQPSYNNG-VMVVVTGCLTGTDNIK-RKFAQSFFLAPQDKGFYVLNDVFR 130
+ DAQP+ NG V+V++TG L D ++F+Q F L P +YV ND+FR
Sbjct: 67 TTLDAQPASPNGDVLVMITGDLLIDDEQNPQRFSQVFHLIPDGNSYYVFNDIFR 120
>NTF2_YEAST (P33331) Nuclear transport factor 2 (NTF-2) (Nuclear
transport factor P10)
Length = 125
Score = 54.3 bits (129), Expect = 5e-07
Identities = 33/114 (28%), Positives = 58/114 (49%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F + YY+ D Q+ Y + S+++ + +I +K+ SL + + +
Sbjct: 12 FTQFYYNQFDTDRSQLGNLYRNESMLTFETSQ-----LQGAKDIVEKLVSLPFQKVQHRI 66
Query: 79 LSADAQPSYNNG-VMVVVTG-CLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
+ DAQP+ NG V+V++TG L + ++F+Q F L P +YV ND+FR
Sbjct: 67 TTLDAQPASPNGDVLVMITGDLLIDEEQNPQRFSQVFHLIPDGNSYYVFNDIFR 120
>NTF2_YARLI (Q6CC82) Nuclear transport factor 2 (NTF-2)
Length = 123
Score = 53.5 bits (127), Expect = 9e-07
Identities = 37/114 (32%), Positives = 53/114 (46%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F E YY D Q+ Y D S+++ T T I +K+ L + R ++
Sbjct: 12 FCEFYYQTFDTDRSQLGNLYRDHSMLTF-----TGTQHQGAQAIVEKLVGLPFGQVRHKI 66
Query: 79 LSADAQPSYNNG--VMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
DAQP+ G V+V+VTG L + +AQ F L P +YV ND+FR
Sbjct: 67 SDIDAQPASAQGGDVIVLVTGELCVDGDNPLPYAQVFHLIPDGSSYYVFNDIFR 120
>NTF2_CANAL (Q9P926) Nuclear transport factor 2 (NTF-2)
Length = 124
Score = 52.4 bits (124), Expect = 2e-06
Identities = 33/118 (27%), Positives = 58/118 (48%), Gaps = 7/118 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
V F YY+ D Q+ Y + S+++ + +I +K+ SL +
Sbjct: 8 VATEFCNFYYNQFDSDRSQLGNLYRNESMLTFETSQ-----LQGARDIVEKLASLPFQKV 62
Query: 75 RVEVLSADAQPSYNNG-VMVVVTG-CLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
+ + DAQP+ NG ++V+VTG L + ++++Q F L P + +YV ND+FR
Sbjct: 63 AHRISTLDAQPASANGDILVMVTGELLIDEEQNAQRYSQVFHLIPDNGSYYVFNDIFR 120
>NTF2_SCHPO (Q10100) Nuclear transport factor 2 (NTF-2)
Length = 123
Score = 52.0 bits (123), Expect = 3e-06
Identities = 35/114 (30%), Positives = 60/114 (51%), Gaps = 7/114 (6%)
Query: 19 FVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEV 78
F + YY D Q+ Y + S++S +G + T I +K+ SL + + +
Sbjct: 11 FTQFYYQTFDSDRSQLSSLYREESMLSF---EGAQ--LQGTKAIVEKLVSLPFQRVQHRI 65
Query: 79 LSADAQPSYNNG-VMVVVTG-CLTGTDNIKRKFAQSFFLAPQDKGFYVLNDVFR 130
+ DAQP+ G V+V+VTG L + + ++++Q F L + +YVLND+FR
Sbjct: 66 STLDAQPTGTTGSVIVMVTGELLLDEEQMAQRYSQVFHLVNNNGNYYVLNDLFR 119
>NTF2_ARATH (Q9C7F5) Nuclear transport factor 2 (NTF-2)
Length = 126
Score = 52.0 bits (123), Expect = 3e-06
Identities = 38/122 (31%), Positives = 60/122 (49%), Gaps = 8/122 (6%)
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P V AFVE YYS + + Y ++S+++ + + I K+ SL +
Sbjct: 6 PDAVSKAFVEHYYSTFDTNRVGLAGLYQEASMLTFEGQ-----KIQGVQSIVAKLTSLPF 60
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + + D QPS +G++V V+G L + KF+Q F L P +G FYV ND+
Sbjct: 61 QQCKHHISTVDCQPSGPASGMLVFVSGNLQLAGEEHALKFSQMFHLMPTPQGSFYVFNDI 120
Query: 129 FR 130
FR
Sbjct: 121 FR 122
>NTF2_EMENI (Q96VN3) Nuclear transport factor 2 (NTF-2)
Length = 125
Score = 51.6 bits (122), Expect = 3e-06
Identities = 37/123 (30%), Positives = 61/123 (49%), Gaps = 8/123 (6%)
Query: 13 QMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYT 72
Q + FV YY + + Y D S+++ + + A I +K+ SL +
Sbjct: 5 QSIAQQFVTFYYQTFDGNRAGLAPLYRDHSMLTFET-----SAIQGVAGIIEKLTSLPFQ 59
Query: 73 SFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDVF 129
+ +V + DAQPS + G++V+VTG L + + Q+F L P G ++VLNDVF
Sbjct: 60 KVQHQVSTLDAQPSGEHGGILVLVTGALLVDEEKNPMNYTQTFQLMPDGAGSYFVLNDVF 119
Query: 130 RYV 132
R +
Sbjct: 120 RLI 122
>NTF2_DEBHA (Q6BWC0) Nuclear transport factor 2 (NTF-2)
Length = 124
Score = 51.2 bits (121), Expect = 4e-06
Identities = 34/118 (28%), Positives = 56/118 (46%), Gaps = 7/118 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
V + F YY D Q+ Y + S+++ + +I +K+ SL +
Sbjct: 8 VASEFCNFYYQQFDSDRTQLGNLYREQSMLTFETSQ-----LQGAKDIVEKLVSLPFQKV 62
Query: 75 RVEVLSADAQPSYNNG-VMVVVTGCLTGTDNIK-RKFAQSFFLAPQDKGFYVLNDVFR 130
+ + DAQP NG ++V+VTG L D ++++Q F L P +YV ND+FR
Sbjct: 63 AHRISTLDAQPGSPNGDILVMVTGELIIDDEQNAQRYSQVFHLIPDGNSYYVFNDIFR 120
>ZAN_MOUSE (O88799) Zonadhesin precursor
Length = 5376
Score = 50.1 bits (118), Expect = 1e-05
Identities = 42/181 (23%), Positives = 72/181 (39%), Gaps = 13/181 (7%)
Query: 138 IDIESVPANDADESAPSEAIITPE-----PEPVHVPEVIPPTQTVIPTA---QTVIPPTQ 189
+ E + + S P+E I+ E PE +P +P T + +T +PP +
Sbjct: 867 VHTEVTNVSPEETSVPTEETISTEVTTVSPEETTLPTEVPTVSTEVTNVSPEETSVPPEE 926
Query: 190 TVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEP-EQPT-SIEKVASNT 247
T++ + T+ +E P+E L +P+ + P E PT S E +T
Sbjct: 927 TILTEITTVSPEETVFPIEGTTLPTEVLTVPIEVTTFPTGETTVPTEVPTVSTEMTGVHT 986
Query: 248 QEDT---PKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNMDIPLE 304
+ T + S + V + S P +P + PP ++PA P IP E
Sbjct: 987 EVTTVFPEETSIPTEVATVLPASIPPEETTTPTEVTTTPPEETTIPAEVTTVPPASIPPE 1046
Query: 305 K 305
+
Sbjct: 1047 E 1047
Score = 41.2 bits (95), Expect = 0.005
Identities = 29/134 (21%), Positives = 58/134 (42%), Gaps = 18/134 (13%)
Query: 137 SIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQ-TVIPT--------------A 181
++ E + + + S P E I E V E+I PT+ T +PT
Sbjct: 601 TVPTEVINVSPKETSIPPEVTIPTEVITVSPEEIISPTEVTPVPTDVTAAYVEATNASPE 660
Query: 182 QTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIE 241
+T +PP T++ + T+ +E ++P E + + P E++ + + P PT +
Sbjct: 661 ETSVPPEVTILTEVTTVSPEETTVPTEVPIVLIEATAFPTG---ETTLYTEVPTVPTEVT 717
Query: 242 KVASNTQEDTPKKS 255
V + +P+++
Sbjct: 718 GVHTEVTNVSPEET 731
Score = 37.0 bits (84), Expect = 0.087
Identities = 25/83 (30%), Positives = 38/83 (45%), Gaps = 16/83 (19%)
Query: 149 DESAPSEAIITPEPEPVHVPE---VIPPTQTVIPTA------------QTVIPPTQTVIA 193
+E+A + T PE P +PP +T IPT +T +PP +T IA
Sbjct: 1046 EETASLTEVTTTPPEETTTPTEVTTVPPEKTTIPTEVTTVPPASIFPEETTVPPEETTIA 1105
Query: 194 DTETIIS-KEVSLPLENGKLSVT 215
ET +S +E +L E ++ T
Sbjct: 1106 SEETTVSTQETTLLTEQSAVTQT 1128
Score = 35.4 bits (80), Expect = 0.25
Identities = 25/86 (29%), Positives = 32/86 (37%), Gaps = 1/86 (1%)
Query: 153 PSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIADTETIISKEVSLPLENGKL 212
P E I E V +P IPP +T PT T PP +T I T + P E L
Sbjct: 993 PEETSIPTEVATV-LPASIPPEETTTPTEVTTTPPEETTIPAEVTTVPPASIPPEETASL 1051
Query: 213 SVTENVIPVNHVKESSHHVKEPEQPT 238
+ P + PE+ T
Sbjct: 1052 TEVTTTPPEETTTPTEVTTVPPEKTT 1077
Score = 35.0 bits (79), Expect = 0.33
Identities = 44/191 (23%), Positives = 77/191 (40%), Gaps = 21/191 (10%)
Query: 47 PEEDGTMTTVTTTAEIDKKIQSLEYTSFRVEVLSADAQ-PSYNNGVMVVVTGCLTGTDNI 105
PEE T+ T TT ++ + +E T+ EVL+ + ++ G V T T + +
Sbjct: 924 PEE--TILTEITTVSPEETVFPIEGTTLPTEVLTVPIEVTTFPTGETTVPTEVPTVSTEM 981
Query: 106 KRKFAQSFFLAPQDKGF-----YVLNDVFRYVDAYKSIDIESVPAND----ADESAPSEA 156
+ + P++ VL + ++ + P + A+ + A
Sbjct: 982 TGVHTEVTTVFPEETSIPTEVATVLPASIPPEETTTPTEVTTTPPEETTIPAEVTTVPPA 1041
Query: 157 IITPEPEPVHVPEVI--PPTQTVIPTAQTVIPPTQTVIADTET------IISKEVSLPLE 208
I PE E + EV PP +T PT T +PP +T I T I +E ++P E
Sbjct: 1042 SIPPE-ETASLTEVTTTPPEETTTPTEVTTVPPEKTTIPTEVTTVPPASIFPEETTVPPE 1100
Query: 209 NGKLSVTENVI 219
++ E +
Sbjct: 1101 ETTIASEETTV 1111
Score = 33.5 bits (75), Expect = 0.96
Identities = 26/123 (21%), Positives = 49/123 (39%), Gaps = 5/123 (4%)
Query: 138 IDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPTQTVIADTE- 196
+ E + + S P+E I+ E V E PT+ I + PT + TE
Sbjct: 719 VHTEVTNVSPEETSVPTEETISTEVTTVSPEETTVPTEVPIVLIEATASPTGEITLYTEV 778
Query: 197 -TIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQ---PTSIEKVASNTQEDTP 252
T+ ++ + E +S E +P + PE+ PT + V++ +P
Sbjct: 779 PTVPTEVTGVHTEVTNVSPEETSVPTEETISTEVTTVSPEETTLPTEVPTVSTEVTNVSP 838
Query: 253 KKS 255
+++
Sbjct: 839 EET 841
Score = 32.7 bits (73), Expect = 1.6
Identities = 30/122 (24%), Positives = 46/122 (37%), Gaps = 10/122 (8%)
Query: 140 IESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPTAQTVIPPT--QTVIADTET 197
+E+ A+ + S P E I E V E PT+ I + PT T+ + T
Sbjct: 652 VEATNASPEETSVPPEVTILTEVTTVSPEETTVPTEVPIVLIEATAFPTGETTLYTEVPT 711
Query: 198 IISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPT--------SIEKVASNTQE 249
+ ++ + E +S E +P + PE+ T IE AS T E
Sbjct: 712 VPTEVTGVHTEVTNVSPEETSVPTEETISTEVTTVSPEETTVPTEVPIVLIEATASPTGE 771
Query: 250 DT 251
T
Sbjct: 772 IT 773
Score = 32.3 bits (72), Expect = 2.1
Identities = 30/133 (22%), Positives = 49/133 (36%), Gaps = 16/133 (12%)
Query: 144 PANDADESAPSEAIITPEPEPVHVPEV--IPPTQTVIPTAQTVIPPTQTVIADTETIISK 201
P+ + P E +P E+ P + IPT T +P ++ ET I
Sbjct: 559 PSESTVPTLPMEQPTSPTKATTVTIEIPTTPTEEATIPTETTTVPTEVINVSPKETSIPP 618
Query: 202 EVSLPLENGKLSVTENVIP--------------VNHVKESSHHVKEPEQPTSIEKVASNT 247
EV++P E +S E + P V S P + T + +V + +
Sbjct: 619 EVTIPTEVITVSPEEIISPTEVTPVPTDVTAAYVEATNASPEETSVPPEVTILTEVTTVS 678
Query: 248 QEDTPKKSFASIV 260
E+T + IV
Sbjct: 679 PEETTVPTEVPIV 691
Score = 30.4 bits (67), Expect = 8.2
Identities = 35/190 (18%), Positives = 70/190 (36%), Gaps = 36/190 (18%)
Query: 138 IDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPTQTVIPT--------------AQT 183
+ E + + S P+E I+ E V P +T +PT +T
Sbjct: 788 VHTEVTNVSPEETSVPTEETISTEVTTVS------PEETTLPTEVPTVSTEVTNVSPEET 841
Query: 184 VIPPTQTVI----ADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQ--- 236
+PP +T++ + T+ ++ + E +S E +P + PE+
Sbjct: 842 SVPPEETILTTLYTEVPTVPTEVTGVHTEVTNVSPEETSVPTEETISTEVTTVSPEETTL 901
Query: 237 ----PTSIEKVASNTQEDT---PKKSFASIVNALKDNSAPFHLRAS--PAKPAVHPPRVH 287
PT +V + + E+T P+++ + + + F + + P + P V
Sbjct: 902 PTEVPTVSTEVTNVSPEETSVPPEETILTEITTVSPEETVFPIEGTTLPTEVLTVPIEVT 961
Query: 288 SVPAPEAPTP 297
+ P E P
Sbjct: 962 TFPTGETTVP 971
>NTF2_NEUCR (P87102) Nuclear transport factor 2 (NTF-2)
Length = 124
Score = 48.5 bits (114), Expect = 3e-05
Identities = 34/120 (28%), Positives = 60/120 (49%), Gaps = 7/120 (5%)
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
+ FV YYS D + Y D+S+++ +G + I +K+ SL +
Sbjct: 8 IATQFVAHYYSTFDSDRKNLAGLYRDNSMLTF---EGAQSL--GAQGITEKLTSLPFQKV 62
Query: 75 RVEVLSADAQPSYNNGVMVVVTGCLTGTDNIK-RKFAQSFFLAPQDKG-FYVLNDVFRYV 132
+ E DAQP+ G++++VTG L D + ++Q+F L+ G ++V ND+F+ V
Sbjct: 63 KHEYGPPDAQPTATGGIIILVTGQLIVDDEQRPLGYSQAFQLSQDASGQWFVFNDIFKLV 122
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,280,993
Number of Sequences: 164201
Number of extensions: 2131259
Number of successful extensions: 7473
Number of sequences better than 10.0: 255
Number of HSP's better than 10.0 without gapping: 58
Number of HSP's successfully gapped in prelim test: 198
Number of HSP's that attempted gapping in prelim test: 6680
Number of HSP's gapped (non-prelim): 585
length of query: 393
length of database: 59,974,054
effective HSP length: 112
effective length of query: 281
effective length of database: 41,583,542
effective search space: 11684975302
effective search space used: 11684975302
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146567.10


