

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146565.8 - phase: 0 /pseudo
(1145 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
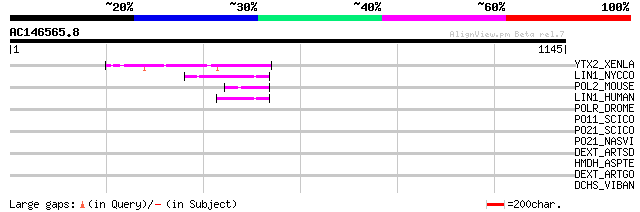
Sequences producing significant alignments: (bits) Value
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 79 9e-14
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 56 5e-07
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 53 5e-06
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 53 5e-06
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 36 0.68
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 35 1.2
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 34 2.0
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 34 2.6
DEXT_ARTSD (P39652) Dextranase precursor (EC 3.2.1.11) (Alpha-1,... 33 3.4
HMDH_ASPTE (Q9Y7D2) 3-hydroxy-3-methylglutaryl-coenzyme A reduct... 33 5.8
DEXT_ARTGO (P70744) Dextranase precursor (EC 3.2.1.11) (Alpha-1,... 32 7.5
DCHS_VIBAN (Q56581) Histidine decarboxylase (EC 4.1.1.22) (HDC) 32 7.5
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 78.6 bits (192), Expect = 9e-14
Identities = 89/366 (24%), Positives = 162/366 (43%), Gaps = 42/366 (11%)
Query: 197 RLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVDC 256
R+DR + ++ R S + SDH+ S++ +I+ KA H +
Sbjct: 203 RIDRIYISSHLMS--RAQSSTIRLAPFSDHN-------CVSLRMSIAPSLPKAAYWHFNN 253
Query: 257 RRLVLENWAKNVRGV--GMVRLQ------------GKLRILKTVFKQWNRSVFGDVDRQV 302
L E +AK+VR G Q GK+ LK + +++ +SV G + ++
Sbjct: 254 SLLEDEGFAKSVRDTWRGWRAFQDEFATLNQWWDVGKVH-LKLLCQEYTKSVSGQRNAEI 312
Query: 303 RLATDEVSRIQQIIDSVGFSDD--LYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGD 360
EV ++Q + S+D L + LE + L N Q + +SR Q D
Sbjct: 313 EALNGEVLDLEQRLSG---SEDQALQCEYLERKEALRNMEQRQARGAFVRSRMQLLCDMD 369
Query: 361 RNTAYFHRVARIKASSKNISLLY-DGATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLV 419
R + +F+ + + K + K I+ L+ + T L D I +++Q +F P+ +
Sbjct: 370 RGSRFFYALEKKKGNRKQITCLFAEDGTPLEDPEAIRDRARSFYQNLFS------PDPIS 423
Query: 420 ARSIHAL-----VTEDENNMLIRLPLR-DEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDI 473
+ L V + + P+ DE+ A+ + + +PG DG F+Q FWD
Sbjct: 424 PDACEELWDGLPVVSERRKERLETPITLDELSQALRLMPHNKSPGLDGLTIEFFQFFWDT 483
Query: 474 VGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILA 533
+G D + + F +G + +L L+PK + + ++RP++L + +KIV K ++
Sbjct: 484 LGPDFHRVLTEAFKKGELPLSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAIS 543
Query: 534 DRLASI 539
RL S+
Sbjct: 544 LRLKSV 549
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 56.2 bits (134), Expect = 5e-07
Identities = 51/181 (28%), Positives = 84/181 (46%), Gaps = 7/181 (3%)
Query: 360 DRNTAYFHRVARIKASSKNISLLYDGATVLSDAGDIAAHILNYFQGIFD-VVNNCIPNDL 418
D+ A R R+K+ +I D T +D +I + Y++ ++ N D
Sbjct: 373 DKPLANLTRKKRVKSLISSIRNGNDEIT--TDPSEIQKILNEYYKKLYSHKYENLKEIDQ 430
Query: 419 VARSIHA-LVTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSD 477
+ H +++ E ML R EI S + +L +PGPDGF FYQ+F + +
Sbjct: 431 YLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPKKKSPGPDGFTSEFYQTFKEELVPI 490
Query: 478 VVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQ--AMGDFRPIALANFQFKIVTKILADR 535
++ Q+ +G + + LIPK PG ++RPI+L N KI+ KIL +R
Sbjct: 491 LLNLFQNIEKEGILPNTFYEANITLIPK-PGKDPTRKENYRPISLMNIDAKILNKILTNR 549
Query: 536 L 536
+
Sbjct: 550 I 550
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 52.8 bits (125), Expect = 5e-06
Identities = 33/99 (33%), Positives = 54/99 (54%), Gaps = 9/99 (9%)
Query: 443 EIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFF----LQGSFADNINSN 498
EI++ + L +PGPDGF FYQ+F + D++ + F ++G+ ++
Sbjct: 483 EIEAVINSLPTKKSPGPDGFSAEFYQTFKE----DLIPILHKLFHKIEVEGTLPNSFYEA 538
Query: 499 LLVLIPK-IPGAQAMGDFRPIALANFQFKIVTKILADRL 536
+ LIPK + +FRPI+L N KI+ KILA+R+
Sbjct: 539 TITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRI 577
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 52.8 bits (125), Expect = 5e-06
Identities = 34/112 (30%), Positives = 58/112 (51%), Gaps = 3/112 (2%)
Query: 427 VTEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFF 486
+ ++E L R EI++ + L +PGP+GF FYQ + + + ++ Q
Sbjct: 440 LNQEEVESLNRPITSSEIEAIINSLPNKKSPGPEGFTAEFYQRYKEELVPFLLKLFQSIE 499
Query: 487 LQGSFADNINSNLLVLIPKIPGAQA--MGDFRPIALANFQFKIVTKILADRL 536
+G ++ ++LIPK PG +FRPI+L N KI+ KILA+++
Sbjct: 500 KEGILPNSFYEASIILIPK-PGRDTTKKENFRPISLMNIDAKILNKILANQI 550
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 35.8 bits (81), Expect = 0.68
Identities = 27/84 (32%), Positives = 41/84 (48%), Gaps = 5/84 (5%)
Query: 456 APGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQ-GSFADNINSNLLVLIPKIPGAQAMGD 514
+PGPDG + V S ++L + + L G+ +I V IPK A+ D
Sbjct: 359 SPGPDGITPKSARE----VPSGIMLRIMNLILWCGNLPHSIRLARTVFIPKTVTAKRPQD 414
Query: 515 FRPIALANFQFKIVTKILADRLAS 538
FRPI++ + + + ILA RL S
Sbjct: 415 FRPISVPSVLVRQLNAILATRLNS 438
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 35.0 bits (79), Expect = 1.2
Identities = 24/97 (24%), Positives = 41/97 (41%), Gaps = 2/97 (2%)
Query: 442 DEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLV 501
DE+ +V +PGPDG G ++ W + + + L+ F LV
Sbjct: 408 DEVSDSVRRCKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLYKQCLLESYFPQKWKIASLV 467
Query: 502 LIPKIPG--AQAMGDFRPIALANFQFKIVTKILADRL 536
++ K+ G +RPI L + K++ I+ RL
Sbjct: 468 ILLKLLDRIRSDPGSYRPICLLDNLGKVLEGIMVKRL 504
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 34.3 bits (77), Expect = 2.0
Identities = 32/96 (33%), Positives = 40/96 (41%), Gaps = 5/96 (5%)
Query: 443 EIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVL 502
EIKSA N GA GPDG + + D L F G I + V
Sbjct: 163 EIKSARAS-NEKGA-GPDGVTPRSWNALDDRYKR---LLYNIFVFYGRVPSPIKGSRTVF 217
Query: 503 IPKIPGAQAMGDFRPIALANFQFKIVTKILADRLAS 538
PKI G G FRP+++ + + KILA R S
Sbjct: 218 TPKIEGGPDPGVFRPLSICSVILREFNKILARRFVS 253
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 33.9 bits (76), Expect = 2.6
Identities = 22/81 (27%), Positives = 35/81 (43%), Gaps = 4/81 (4%)
Query: 456 APGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDF 515
A GPDG + W+ + + G + VLIPK PG F
Sbjct: 334 AAGPDGMT----TTAWNSIDECIKSLFNMIMYHGQCPRRYLDSRTVLIPKEPGTMDPACF 389
Query: 516 RPIALANFQFKIVTKILADRL 536
RP+++A+ + +ILA+R+
Sbjct: 390 RPLSIASVALRHFHRILANRI 410
>DEXT_ARTSD (P39652) Dextranase precursor (EC 3.2.1.11)
(Alpha-1,6-glucan-6-glucanohydrolase) (Endodextranase)
Length = 640
Score = 33.5 bits (75), Expect = 3.4
Identities = 27/108 (25%), Positives = 49/108 (45%), Gaps = 20/108 (18%)
Query: 29 NKPTFIFLAEPMISFSYVPSWYWHSIGVKKCCLNNRGPRLPNLWALWNDEVLVVVIFNSD 88
N+ TF E ++ V SWYW + G++ +G + N + ND+VL +++SD
Sbjct: 398 NEQTFHMNVE---NYKQVGSWYWQTDGIEL----YKGSTMKNTFFNANDDVL--KMYHSD 448
Query: 89 QCIALEVTWQHSSVFLA-----------AIYANTSYLFRRLLWADLTH 125
I V W++ + + ANT+ + R+ W D+ +
Sbjct: 449 VTIDNTVIWKNENGPVIQWGWTPRNIDNVNVANTTVIHNRMYWKDVKY 496
>HMDH_ASPTE (Q9Y7D2) 3-hydroxy-3-methylglutaryl-coenzyme A reductase
(EC 1.1.1.34) (HMG-CoA reductase)
Length = 1068
Score = 32.7 bits (73), Expect = 5.8
Identities = 29/96 (30%), Positives = 42/96 (43%), Gaps = 8/96 (8%)
Query: 72 WALWNDEVLVVVIFNSDQCIALEVTWQHSSVFLAAIYA-NTSYLF------RRLLWADLT 124
W L D VL+ + + C+ LE+T S L + A + S +F + W L
Sbjct: 358 WTLLIDAVLLFTFYATILCVKLEITRIRSPGGLGQVNAKHPSGIFGHKVKSTNITWWKLL 417
Query: 125 HLQGCFLGPWLFIGD-FNAVMGAHEKRGRWPPTAAS 159
+ G L +L + F VMG + G PPTA S
Sbjct: 418 TVGGFVLCHFLQLSPFFYRVMGEYMANGTLPPTAVS 453
>DEXT_ARTGO (P70744) Dextranase precursor (EC 3.2.1.11)
(Alpha-1,6-glucan-6-glucanohydrolase) (Endodextranase)
Length = 640
Score = 32.3 bits (72), Expect = 7.5
Identities = 26/108 (24%), Positives = 48/108 (44%), Gaps = 20/108 (18%)
Query: 29 NKPTFIFLAEPMISFSYVPSWYWHSIGVKKCCLNNRGPRLPNLWALWNDEVLVVVIFNSD 88
N+ TF E ++ V SWYW + G++ +G + N + ND+VL +++SD
Sbjct: 398 NEQTFHMNVE---NYKQVGSWYWQTDGIEL----YQGSTMKNTFFNANDDVL--KMYHSD 448
Query: 89 QCIALEVTWQHSSVFLA-----------AIYANTSYLFRRLLWADLTH 125
I V W++ + + NT+ + R+ W D+ +
Sbjct: 449 VSIDNTVVWKNENGPVVQWGWTPRNIDNVNVTNTTVIHNRMYWKDVKY 496
>DCHS_VIBAN (Q56581) Histidine decarboxylase (EC 4.1.1.22) (HDC)
Length = 386
Score = 32.3 bits (72), Expect = 7.5
Identities = 17/47 (36%), Positives = 26/47 (55%)
Query: 288 KQWNRSVFGDVDRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQL 334
KQ N +F ++ +R ATD + RIQQ + S+G + Y +A L
Sbjct: 158 KQKNPIIFANIGTTMRGATDNIQRIQQDLASIGLERNDYYIHADAAL 204
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.346 0.153 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 129,206,013
Number of Sequences: 164201
Number of extensions: 5341155
Number of successful extensions: 18558
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 18540
Number of HSP's gapped (non-prelim): 17
length of query: 1145
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1024
effective length of database: 40,105,733
effective search space: 41068270592
effective search space used: 41068270592
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC146565.8


