

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
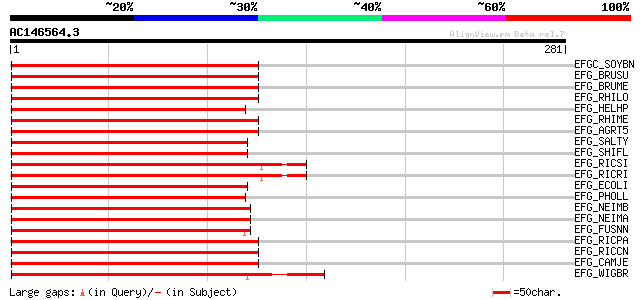
Score E
Sequences producing significant alignments: (bits) Value
EFGC_SOYBN (P34811) Elongation factor G, chloroplast precursor (... 186 7e-47
EFG_BRUSU (Q8G075) Elongation factor G (EF-G) 158 2e-38
EFG_BRUME (Q8YHP3) Elongation factor G (EF-G) 158 2e-38
EFG_RHILO (Q98N59) Elongation factor G (EF-G) 152 9e-37
EFG_HELHP (Q7VJ85) Elongation factor G (EF-G) 150 4e-36
EFG_RHIME (Q92QH2) Elongation factor G (EF-G) 149 6e-36
EFG_AGRT5 (Q8UE15) Elongation factor G (EF-G) 148 1e-35
EFG_SALTY (P26229) Elongation factor G (EF-G) 147 4e-35
EFG_SHIFL (Q83JC3) Elongation factor G (EF-G) 146 6e-35
EFG_RICSI (Q8KTB8) Elongation factor G (EF-G) 146 6e-35
EFG_RICRI (Q8KTC1) Elongation factor G (EF-G) 146 6e-35
EFG_ECOLI (P02996) Elongation factor G (EF-G) 146 6e-35
EFG_PHOLL (Q7N9B2) Elongation factor G (EF-G) 145 8e-35
EFG_NEIMB (Q9K1I8) Elongation factor G (EF-G) 145 8e-35
EFG_NEIMA (Q9JX07) Elongation factor G (EF-G) 145 8e-35
EFG_FUSNN (Q8R602) Elongation factor G (EF-G) 145 8e-35
EFG_RICPA (Q8KTB9) Elongation factor G (EF-G) 145 1e-34
EFG_RICCN (Q92J93) Elongation factor G (EF-G) 145 1e-34
EFG_CAMJE (Q9PI16) Elongation factor G (EF-G) 145 1e-34
EFG_WIGBR (Q8D3H2) Elongation factor G (EF-G) 145 1e-34
>EFGC_SOYBN (P34811) Elongation factor G, chloroplast precursor
(EF-G)
Length = 788
Score = 186 bits (471), Expect = 7e-47
Identities = 91/126 (72%), Positives = 108/126 (85%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 465 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 524
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS+++EV
Sbjct: 525 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKISEV 584
Query: 121 RYVFNK 126
+YV K
Sbjct: 585 KYVHKK 590
Score = 37.4 bits (85), Expect = 0.041
Identities = 16/28 (57%), Positives = 22/28 (78%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 754 MTKGRASYTMQLAMFDVVPQHIQNQLAT 781
>EFG_BRUSU (Q8G075) Elongation factor G (EF-G)
Length = 694
Score = 158 bits (399), Expect = 2e-38
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>EFG_BRUME (Q8YHP3) Elongation factor G (EF-G)
Length = 694
Score = 158 bits (399), Expect = 2e-38
Identities = 80/126 (63%), Positives = 95/126 (74%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTKAD EKM L +LA
Sbjct: 377 DIVALAGLKETTTGDTLCDPLKPVILERMEFPDPVIEIAIEPKTKADQEKMGIALNRLAA 436
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR+KREFKVEANVGAPQ YRESI+ E+
Sbjct: 437 EDPSFRVKSDEESGQTIIAGMGELHLDILVDRMKREFKVEANVGAPQVAYRESITRAAEI 496
Query: 121 RYVFNK 126
Y K
Sbjct: 497 DYTHKK 502
>EFG_RHILO (Q98N59) Elongation factor G (EF-G)
Length = 696
Score = 152 bits (384), Expect = 9e-37
Identities = 79/126 (62%), Positives = 93/126 (73%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDT G+TLCDP P +L++M+FP VI+IAIEPKTK D EKM L +LA
Sbjct: 378 DIVALAGLKDTTTGDTLCDPLHPVILERMEFPDPVIQIAIEPKTKNDQEKMGLALHRLAA 437
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEANVGAPQ YRE+I+ E
Sbjct: 438 EDPSFRVKTDEESGQTIISGMGELHLDIIVDRMRREFKVEANVGAPQVAYRETITRKHEQ 497
Query: 121 RYVFNK 126
Y K
Sbjct: 498 DYTHKK 503
>EFG_HELHP (Q7VJ85) Elongation factor G (EF-G)
Length = 692
Score = 150 bits (378), Expect = 4e-36
Identities = 77/119 (64%), Positives = 90/119 (74%), Gaps = 1/119 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
+I VGLKDT+ G+TLCD K P +L++MDFP VI+IA+EPKTKAD EKM L KLAE
Sbjct: 373 EICAFVGLKDTVTGDTLCDEKVPVILERMDFPEPVIQIAVEPKTKADQEKMSIALSKLAE 432
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTE 119
EDPSFRVS +E +T+I G GELHLE IVDRLKREFKVEA VG PQ +RE+I E
Sbjct: 433 EDPSFRVSTHEETGQTLIGGMGELHLEIIVDRLKREFKVEAEVGEPQVAFRETIRTAVE 491
>EFG_RHIME (Q92QH2) Elongation factor G (EF-G)
Length = 699
Score = 149 bits (377), Expect = 6e-36
Identities = 77/126 (61%), Positives = 93/126 (73%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTK D EKM L +LA
Sbjct: 378 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIQIAIEPKTKGDQEKMGLALNRLAA 437
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ IVDR++REFKVEA+VGAPQ YRE+I+ E
Sbjct: 438 EDPSFRVKTDEESGQTIIAGMGELHLDIIVDRMRREFKVEASVGAPQVAYRETITRKHEE 497
Query: 121 RYVFNK 126
Y K
Sbjct: 498 DYTHKK 503
>EFG_AGRT5 (Q8UE15) Elongation factor G (EF-G)
Length = 699
Score = 148 bits (374), Expect = 1e-35
Identities = 76/126 (60%), Positives = 92/126 (72%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLK+T G+TLCDP P +L++M+FP VI+IAIEPKTK D EKM L +LA
Sbjct: 378 DIVALAGLKETTTGDTLCDPLKPVILERMEFPEPVIQIAIEPKTKGDQEKMGLALNRLAA 437
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHL+ +VDR++REFKVEA VGAPQ YRE+I+ E
Sbjct: 438 EDPSFRVKTDEESGQTIIAGMGELHLDILVDRMRREFKVEATVGAPQVAYRETITRQHEE 497
Query: 121 RYVFNK 126
Y K
Sbjct: 498 DYTHKK 503
Score = 30.0 bits (66), Expect = 6.6
Identities = 11/26 (42%), Positives = 17/26 (65%)
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNEL 278
+++GRA YSM FD + V N+ E+
Sbjct: 666 MSQGRAQYSMTFDHYSPVPSNVAQEI 691
>EFG_SALTY (P26229) Elongation factor G (EF-G)
Length = 703
Score = 147 bits (370), Expect = 4e-35
Identities = 74/121 (61%), Positives = 94/121 (77%), Gaps = 2/121 (1%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD G+TLCDP+ P +L++M+FP VI IA+EPKTKAD EKM L +LA+
Sbjct: 380 DIAAAIGLKDVTTGDTLCDPENPIILERMEFPEPVISIAVEPKTKADQEKMGLALGRLAK 439
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESI-SEVTE 119
EDPSFRV D+E ++TII G GELHL+ IVDR+KREF VEANVG PQ YRE+I ++VT+
Sbjct: 440 EDPSFRVWTDEESNQTIIAGMGELHLDIIVDRMKREFNVEANVGKPQVAYREAIRAKVTD 499
Query: 120 V 120
+
Sbjct: 500 I 500
>EFG_SHIFL (Q83JC3) Elongation factor G (EF-G)
Length = 703
Score = 146 bits (368), Expect = 6e-35
Identities = 75/121 (61%), Positives = 92/121 (75%), Gaps = 2/121 (1%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD G+TLCDP P +L++M+FP VI IA+EPKTKAD EKM L +LA+
Sbjct: 380 DIAAAIGLKDVTTGDTLCDPDAPIILERMEFPEPVISIAVEPKTKADQEKMGLALGRLAK 439
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESI-SEVTE 119
EDPSFRV D+E ++TII G GELHL+ IVDR+KREF VEANVG PQ YRE+I +VT+
Sbjct: 440 EDPSFRVWTDEESNQTIIAGMGELHLDIIVDRMKREFNVEANVGKPQVAYRETIRQKVTD 499
Query: 120 V 120
V
Sbjct: 500 V 500
>EFG_RICSI (Q8KTB8) Elongation factor G (EF-G)
Length = 699
Score = 146 bits (368), Expect = 6e-35
Identities = 81/161 (50%), Positives = 106/161 (65%), Gaps = 14/161 (8%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDT G+TL D +L++M+FP VI++A+EPK+ AD EKM L +LA
Sbjct: 373 DIVALAGLKDTTTGDTLSDIDQQVILERMEFPEPVIELAVEPKSTADQEKMGLALSRLAA 432
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +T+I+G GELHLE I+DR++REFKVEAN+GAPQ YRE+I++V E+
Sbjct: 433 EDPSFRVSTDYETGQTVIKGMGELHLEIIIDRMRREFKVEANIGAPQVAYRETITKVCEI 492
Query: 121 RYVFNK-----------PYILNPWTQVVDTNLRIKSKKKHF 150
Y K I P +V D L+ + K K+F
Sbjct: 493 DYTHKKQSGGAGQFARVKIIFEPLKEVKD--LKDEDKNKNF 531
>EFG_RICRI (Q8KTC1) Elongation factor G (EF-G)
Length = 699
Score = 146 bits (368), Expect = 6e-35
Identities = 81/161 (50%), Positives = 106/161 (65%), Gaps = 14/161 (8%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDT G+TL D +L++M+FP VI++A+EPK+ AD EKM L +LA
Sbjct: 373 DIVALAGLKDTTTGDTLSDIDQQVILERMEFPEPVIELAVEPKSTADQEKMGLALSRLAA 432
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +T+I+G GELHLE I+DR++REFKVEAN+GAPQ YRE+I++V E+
Sbjct: 433 EDPSFRVSTDYETGQTVIKGMGELHLEIIIDRMRREFKVEANIGAPQVAYRETITKVCEI 492
Query: 121 RYVFNK-----------PYILNPWTQVVDTNLRIKSKKKHF 150
Y K I P +V D L+ + K K+F
Sbjct: 493 DYTHKKQSGGAGQFARVKIIFEPLKEVKD--LKDEDKNKNF 531
>EFG_ECOLI (P02996) Elongation factor G (EF-G)
Length = 703
Score = 146 bits (368), Expect = 6e-35
Identities = 75/121 (61%), Positives = 92/121 (75%), Gaps = 2/121 (1%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD G+TLCDP P +L++M+FP VI IA+EPKTKAD EKM L +LA+
Sbjct: 380 DIAAAIGLKDVTTGDTLCDPDAPIILERMEFPEPVISIAVEPKTKADQEKMGLALGRLAK 439
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESI-SEVTE 119
EDPSFRV D+E ++TII G GELHL+ IVDR+KREF VEANVG PQ YRE+I +VT+
Sbjct: 440 EDPSFRVWTDEESNQTIIAGMGELHLDIIVDRMKREFNVEANVGKPQVAYRETIRQKVTD 499
Query: 120 V 120
V
Sbjct: 500 V 500
>EFG_PHOLL (Q7N9B2) Elongation factor G (EF-G)
Length = 702
Score = 145 bits (367), Expect = 8e-35
Identities = 72/119 (60%), Positives = 88/119 (73%), Gaps = 1/119 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD G+TLCD P +L++M+FP VI +AIEPKTKAD EKM L +LA+
Sbjct: 381 DIAAAIGLKDVTTGDTLCDINAPIILERMEFPEPVISVAIEPKTKADQEKMGIALGRLAQ 440
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTE 119
EDPSFRVS D+E +TII G GELHL+ +VDR++REFKVEANVG PQ YRE+I E
Sbjct: 441 EDPSFRVSTDEESGQTIIAGMGELHLDVLVDRMRREFKVEANVGKPQVAYRETIRSTVE 499
>EFG_NEIMB (Q9K1I8) Elongation factor G (EF-G)
Length = 701
Score = 145 bits (367), Expect = 8e-35
Identities = 73/122 (59%), Positives = 88/122 (71%), Gaps = 1/122 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD GETLC P +L++M+FP VI IA+EPKTKAD EKM L +LA+
Sbjct: 380 DIAAAIGLKDVTTGETLCAESAPIILERMEFPEPVIHIAVEPKTKADQEKMGIALNRLAK 439
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHLE IVDR+KREF VEAN+GAPQ YRE+I + +
Sbjct: 440 EDPSFRVRTDEESGQTIISGMGELHLEIIVDRMKREFGVEANIGAPQVAYRETIRKAVKA 499
Query: 121 RY 122
Y
Sbjct: 500 EY 501
>EFG_NEIMA (Q9JX07) Elongation factor G (EF-G)
Length = 701
Score = 145 bits (367), Expect = 8e-35
Identities = 73/122 (59%), Positives = 88/122 (71%), Gaps = 1/122 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD GETLC P +L++M+FP VI IA+EPKTKAD EKM L +LA+
Sbjct: 380 DIAAAIGLKDVTTGETLCAESAPIILERMEFPEPVIHIAVEPKTKADQEKMGIALNRLAK 439
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRV D+E +TII G GELHLE IVDR+KREF VEAN+GAPQ YRE+I + +
Sbjct: 440 EDPSFRVRTDEESGQTIISGMGELHLEIIVDRMKREFGVEANIGAPQVAYRETIRKAVKA 499
Query: 121 RY 122
Y
Sbjct: 500 EY 501
>EFG_FUSNN (Q8R602) Elongation factor G (EF-G)
Length = 693
Score = 145 bits (367), Expect = 8e-35
Identities = 76/124 (61%), Positives = 91/124 (73%), Gaps = 3/124 (2%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI VGLKDT G+TLC P VL+QM+FP VI +A+EPKTK D EKM L KLAE
Sbjct: 375 DIAAAVGLKDTATGDTLCAENAPIVLEQMEFPEPVISVAVEPKTKNDQEKMGIALSKLAE 434
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEV--T 118
EDP+F+V D+E +TII G GELHLE IVDR+KREFKVE+NVG PQ YRE+I++
Sbjct: 435 EDPTFKVRTDEETGQTIISGMGELHLEIIVDRMKREFKVESNVGKPQVAYRETITQSCDQ 494
Query: 119 EVRY 122
EV+Y
Sbjct: 495 EVKY 498
>EFG_RICPA (Q8KTB9) Elongation factor G (EF-G)
Length = 699
Score = 145 bits (366), Expect = 1e-34
Identities = 73/126 (57%), Positives = 94/126 (73%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDT G+TL D +L++M+FP VI++A+EPK+ AD EKM L +LA
Sbjct: 373 DIVALAGLKDTTTGDTLSDIDQQVILERMEFPEPVIELAVEPKSTADQEKMGLALSRLAA 432
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +T+I+G GELHLE I+DR++REFKVEAN+GAPQ YRE+I++V E+
Sbjct: 433 EDPSFRVSTDYETGQTVIKGMGELHLEIIIDRMRREFKVEANIGAPQVAYRETITKVCEI 492
Query: 121 RYVFNK 126
Y K
Sbjct: 493 DYTHKK 498
>EFG_RICCN (Q92J93) Elongation factor G (EF-G)
Length = 699
Score = 145 bits (366), Expect = 1e-34
Identities = 73/126 (57%), Positives = 94/126 (73%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DIV L GLKDT G+TL D +L++M+FP VI++A+EPK+ AD EKM L +LA
Sbjct: 373 DIVALAGLKDTTTGDTLSDIDQQVILERMEFPEPVIELAVEPKSTADQEKMGLALSRLAA 432
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D E +T+I+G GELHLE I+DR++REFKVEAN+GAPQ YRE+I++V E+
Sbjct: 433 EDPSFRVSTDYETGQTVIKGMGELHLEIIIDRMRREFKVEANIGAPQVAYRETITKVCEI 492
Query: 121 RYVFNK 126
Y K
Sbjct: 493 DYTHKK 498
>EFG_CAMJE (Q9PI16) Elongation factor G (EF-G)
Length = 691
Score = 145 bits (366), Expect = 1e-34
Identities = 75/126 (59%), Positives = 91/126 (71%), Gaps = 1/126 (0%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
+I +VGLKDT+ G+TL K +L++MDFP VI +A+EPKTKAD EKM L KLA+
Sbjct: 372 EIGAVVGLKDTLTGDTLASEKDKVILERMDFPDPVISVAVEPKTKADQEKMSIALNKLAQ 431
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTEV 120
EDPSFRVS D+E +TII G GELHLE IVDR+ REFKVEA VG PQ YRE+I + E
Sbjct: 432 EDPSFRVSTDEESGQTIISGMGELHLEIIVDRMLREFKVEAEVGQPQVAYRETIRKTVEQ 491
Query: 121 RYVFNK 126
Y + K
Sbjct: 492 EYKYAK 497
Score = 36.2 bits (82), Expect = 0.092
Identities = 13/25 (52%), Positives = 21/25 (84%)
Query: 254 TEGRASYSMQFDRFDTVSRNIQNEL 278
T+GRA+YSM+FD +D V +N+ +E+
Sbjct: 661 TQGRATYSMEFDHYDEVPKNVADEI 685
>EFG_WIGBR (Q8D3H2) Elongation factor G (EF-G)
Length = 705
Score = 145 bits (365), Expect = 1e-34
Identities = 77/164 (46%), Positives = 103/164 (61%), Gaps = 13/164 (7%)
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI +GLKD G+TLCDP P +L++M+FP VI +A+EPKTK+D EKM + L +LA+
Sbjct: 380 DIAAAIGLKDVTTGDTLCDPLHPIILEKMEFPEPVISVAVEPKTKSDQEKMSSALNRLAK 439
Query: 62 EDPSFRVSQDKED-RTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTE- 119
EDPSFRVS D+E +TII G GELHLE ++DR+ REF V +NVG PQ +YRE+I E
Sbjct: 440 EDPSFRVSSDEESGQTIISGMGELHLEILIDRMYREFNVRSNVGKPQVSYRETIKSCVEE 499
Query: 120 ----VRYVFNKPYILNPWTQVVDTNLRIKSKKKHFQKNSLYWGF 159
+R + + W LRI+ K + KN + F
Sbjct: 500 EGKFIRQSGGRGQFGHVW-------LRIEPNKSNIAKNKNNYTF 536
Score = 29.6 bits (65), Expect = 8.6
Identities = 12/25 (48%), Positives = 18/25 (72%)
Query: 254 TEGRASYSMQFDRFDTVSRNIQNEL 278
T+GRASYSM+F R++ + NI +
Sbjct: 674 TQGRASYSMEFLRYNELPNNITESI 698
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,244,934
Number of Sequences: 164201
Number of extensions: 1160612
Number of successful extensions: 5121
Number of sequences better than 10.0: 350
Number of HSP's better than 10.0 without gapping: 269
Number of HSP's successfully gapped in prelim test: 81
Number of HSP's that attempted gapping in prelim test: 4467
Number of HSP's gapped (non-prelim): 512
length of query: 281
length of database: 59,974,054
effective HSP length: 109
effective length of query: 172
effective length of database: 42,076,145
effective search space: 7237096940
effective search space used: 7237096940
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146564.3


