

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.14 - phase: 0 /pseudo
(312 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
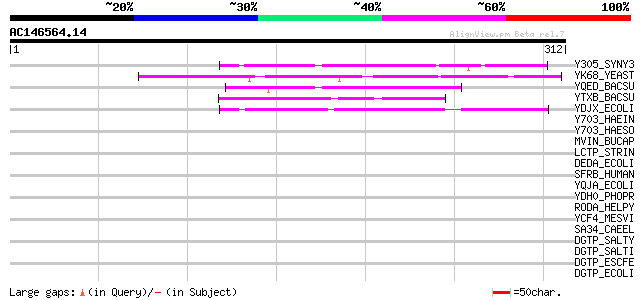
Score E
Sequences producing significant alignments: (bits) Value
Y305_SYNY3 (Q55909) Hypothetical UPF0043 protein slr0305 74 4e-13
YK68_YEAST (P36164) Hypothetical 38.3 kDa protein in PRP16-SRP40... 52 2e-06
YQED_BACSU (P54449) Hypothetical protein yqeD 46 1e-04
YTXB_BACSU (P06568) Hypothetical UPF0043 protein ytxB (ORF-213) 45 3e-04
YDJX_ECOLI (P76219) Hypothetical UPF0043 protein ydjX 44 5e-04
Y703_HAEIN (P44832) Hypothetical protein HI0703 41 0.004
Y703_HAESO (P36684) Hypothetical protein HI0703 homolog 38 0.037
MVIN_BUCAP (Q8K9L3) Virulence factor mviN homolog 38 0.037
LCTP_STRIN (O33654) L-lactate permease 36 0.11
DEDA_ECOLI (P09548) DedA protein (DSG-1 protein) 36 0.14
SFRB_HUMAN (Q05519) Splicing factor arginine/serine-rich 11 (Arg... 33 0.91
YQJA_ECOLI (P42614) Hypothetical protein yqjA 32 1.6
YDH0_PHOPR (Q6LIN8) Hypothetical UPF0324 membrane protein PBPRB0970 32 1.6
RODA_HELPY (P56098) Rod shape-determining protein rodA 32 2.0
YCF4_MESVI (Q9MUN8) Photosystem I assembly protein ycf4 32 2.7
SA34_CAEEL (Q10936) Serpentine receptor class alpha 34 (Sra-34 p... 31 3.5
DGTP_SALTY (P40733) Deoxyguanosinetriphosphate triphosphohydrola... 31 3.5
DGTP_SALTI (Q8Z9B1) Deoxyguanosinetriphosphate triphosphohydrola... 31 4.5
DGTP_ESCFE (Q59435) Deoxyguanosinetriphosphate triphosphohydrola... 31 4.5
DGTP_ECOLI (P15723) Deoxyguanosinetriphosphate triphosphohydrola... 31 4.5
>Y305_SYNY3 (Q55909) Hypothetical UPF0043 protein slr0305
Length = 209
Score = 74.3 bits (181), Expect = 4e-13
Identities = 56/188 (29%), Positives = 87/188 (45%), Gaps = 12/188 (6%)
Query: 119 LTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHK 178
+ +L +VA + LP + AGV G G + + A +G + F++G +
Sbjct: 21 IAFMLLYTVAT--VVFLPGSILTLGAGVVFGVILGSIYVFIGATLGATAAFLVG---RYL 75
Query: 179 IEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYGPYII 237
GW+ K K+ + V L R+SP FP+ + NY TNV Y+I
Sbjct: 76 ARGWVAKKIAGNQKFKAIDEAVGKEGLKIVILTRLSPVFPFNLLNYAYGITNVSLKDYVI 135
Query: 238 GSLVGMVPEIFVAIYTGIL---IKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITV 294
GSL GM+P + +Y G L + TL A+N Q+ Q + +GF TV T+ +T
Sbjct: 136 GSL-GMIPGTIMYVYIGSLAGSLATLGTATN--QANPTLQWTIRIVGFIATVAVTIYVTK 192
Query: 295 YAKRRLKE 302
A++ L E
Sbjct: 193 IARKALNE 200
>YK68_YEAST (P36164) Hypothetical 38.3 kDa protein in PRP16-SRP40
intergenic region
Length = 337
Score = 51.6 bits (122), Expect = 2e-06
Identities = 57/248 (22%), Positives = 114/248 (44%), Gaps = 25/248 (10%)
Query: 73 WVKMVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVLTILLFASVAIFPT 132
W +++++ L + + + +L V I +V+ N E S + ++L VA P
Sbjct: 93 WQQIIIVLLGIMLMIMGILLLVFHNAILHKVVVTSNDLREKMSTHFILMVLIFFVAFPPM 152
Query: 133 I---LLPSTPSMWVAGVTLGYGF-GFLLIITAAAIGVSLPFII-GSIFHHKIEGWLE--- 184
I LL +T G+ G F G++ + + G F++ +I H + E +
Sbjct: 153 IGYSLLSTT-----TGLIYGVSFEGWVTLALGSVTGSIASFVVFKTILHSRAEKLVHLNR 207
Query: 185 KYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFN-YCAVATNVKYGPYIIGSLVGM 243
++ ASIL+ + + +AL+R+ PFPY + N A + + I +++
Sbjct: 208 RFEALASILQENNS------YWILALLRLCPFPYSLTNGAIAGVYGISVRNFSIANII-T 260
Query: 244 VPEIFVAIYTGILIKTLADA-SNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKE 302
P++F+ ++ G +K+LA++ S + II+ L + +T ++ K+R E
Sbjct: 261 TPKLFIYLFIGSRVKSLAESESTGSRVFDLVSIIITLL---ILSLTAWLLYFKTKKRYLE 317
Query: 303 LQEEDKML 310
LQ D+ +
Sbjct: 318 LQNRDRQV 325
>YQED_BACSU (P54449) Hypothetical protein yqeD
Length = 208
Score = 45.8 bits (107), Expect = 1e-04
Identities = 34/137 (24%), Positives = 57/137 (40%), Gaps = 7/137 (5%)
Query: 122 LLFASVAIFPTILLPSTPSMWVA---GVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHK 178
+LF+ + I + P P +A G G G + +T + +G L F + +
Sbjct: 37 VLFSMLLIAADVFFPIVPFALIAALNGAVFGTANGIWITLTGSMLGTILLFFLA---RYS 93
Query: 179 IEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYGPYII 237
W K + ++S A + F AV L R+ P P +V N + V++ +
Sbjct: 94 FRDWARKKVQAYPAIQSYEASFNKNAFTAVLLGRLIPVIPSLVMNVICGLSQVRWHVFFF 153
Query: 238 GSLVGMVPEIFVAIYTG 254
SL+G +P I V G
Sbjct: 154 ASLIGKIPNIVVVTIAG 170
>YTXB_BACSU (P06568) Hypothetical UPF0043 protein ytxB (ORF-213)
Length = 213
Score = 44.7 bits (104), Expect = 3e-04
Identities = 34/129 (26%), Positives = 56/129 (43%), Gaps = 8/129 (6%)
Query: 118 VLTILLFASVAIF-PTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFH 176
V L+F ++I P +L P + G+ G G L + + ++ F +F
Sbjct: 43 VFAPLMFIGISIVRPLVLFPVSVISIAGGLAFGPLLGTLYTLFGSMCASAVSFFAAGLFS 102
Query: 177 HKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYI 236
K G Y + +I K +F+ F L+RI P + +Y A +NVK PY
Sbjct: 103 AKKNG---HYERLEAIQKQMEDNGFFYIF----LLRILPINFDFVSYAAGLSNVKALPYF 155
Query: 237 IGSLVGMVP 245
+ VG++P
Sbjct: 156 AATAVGIIP 164
>YDJX_ECOLI (P76219) Hypothetical UPF0043 protein ydjX
Length = 236
Score = 43.9 bits (102), Expect = 5e-04
Identities = 43/186 (23%), Positives = 80/186 (42%), Gaps = 15/186 (8%)
Query: 119 LTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHK 178
L ILLF + +LLP + + G+ G G LL + AA + S F++
Sbjct: 50 LYILLFIIATL---LLLPGSILVIAGGIVFGPLLGTLLSLIAATLASSCSFLLARWLGRD 106
Query: 179 IEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYGPYII 237
+ L KY ++ ++ G + + L R+ P FPY + NY T + + PY +
Sbjct: 107 L---LLKYVGHSNTFQAIEKGIARNGIDFLILTRLIPLFPYNIQNYAYGLTTIAFWPYTL 163
Query: 238 GSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAK 297
S + +P GI+I T+ + + ++ + I+ L + + +YA+
Sbjct: 164 ISALTTLP--------GIVIYTVMASDLANEGITLRFILQLCLAGLALFILVQLAKLYAR 215
Query: 298 RRLKEL 303
+ +L
Sbjct: 216 HKHVDL 221
>Y703_HAEIN (P44832) Hypothetical protein HI0703
Length = 192
Score = 40.8 bits (94), Expect = 0.004
Identities = 42/190 (22%), Positives = 80/190 (42%), Gaps = 22/190 (11%)
Query: 107 INWETETFSPPVLTILLFASVAIFP----TILLPSTPSMWVAGVTLGYGFGFLLIITAAA 162
+ W F+ L+ + F FP +L+P + S + V F F I A+
Sbjct: 12 MEWSKHRFAVFWLSFVSFIEAIFFPIPPDVMLIPMSMSKPKSAVK----FAFYTAI-ASV 66
Query: 163 IGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALI--RISPFPYMV 220
IG + + IG +E ++++ A K+ +WF Q+ + + SP PY V
Sbjct: 67 IGGIIGYAIGFYATDWVENIVQQWGYAAHWAKAV---SWFEQWGVLVVFVAGFSPIPYKV 123
Query: 221 FNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNAL 280
F CA + + P++I + V + +L+ LA E+ + ++ I +
Sbjct: 124 FTLCAGVMQMAFFPFVITAFVSRLARF-------LLVAKLAAWGGEKFAAKLRKSI-EII 175
Query: 281 GFCLTVVTTV 290
G+ + V+ +
Sbjct: 176 GWSVVVLAVI 185
>Y703_HAESO (P36684) Hypothetical protein HI0703 homolog
Length = 191
Score = 37.7 bits (86), Expect = 0.037
Identities = 39/194 (20%), Positives = 83/194 (42%), Gaps = 22/194 (11%)
Query: 107 INWETETFSPPVLTILLFASVAIFP----TILLPSTPSMWVAGVTLGYGFGFLLIITAAA 162
+ W F+ LT + F FP +L+P M + F F + A+A
Sbjct: 12 MQWANHRFATFWLTFVSFIEAIFFPIPPDVMLIP----MSINKPKCATKFAFYAAM-ASA 66
Query: 163 IGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFR--AVALIRISPFPYMV 220
IG ++ + +G I+ +++++ + A +WF ++ V + SP PY +
Sbjct: 67 IGGAIGYGLGYYAFDFIQSYIQQWGYQQHW---ETALSWFKEWGIWVVFVAGFSPIPYKI 123
Query: 221 FNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNAL 280
F CA + + P+++ + + + +L+ LA S ++ + +Q I +
Sbjct: 124 FTICAGVMQMAFLPFLLTAFISRIARF-------LLVTHLAAWSGKKFAAKLRQSI-EFI 175
Query: 281 GFCLTVVTTVIITV 294
G+ + ++ V+ V
Sbjct: 176 GWSVVIIAIVVYLV 189
>MVIN_BUCAP (Q8K9L3) Virulence factor mviN homolog
Length = 514
Score = 37.7 bits (86), Expect = 0.037
Identities = 44/187 (23%), Positives = 84/187 (44%), Gaps = 17/187 (9%)
Query: 129 IFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKIEGWLEKYPK 188
+FP IL S S+ + + Y + F+ ++++ + +S+ I+ S F + E P
Sbjct: 134 MFPYILFISLSSL-CSSILNSYNYFFIPSLSSSLLNISI--IVFSFF---FSDYFE--PS 185
Query: 189 KASILKSAGAGNWF---HQFRAVALIRISPFPYMVF-NYCAVATNVKYGPYIIGSLVGMV 244
S+ S G +F +QF + I++ FP + F N + K GP +GS +
Sbjct: 186 IISLAWSVMIGGFFQLFYQFPHLYKIKMLVFPKINFKNIGLIKVLKKMGPSTLGSCANQI 245
Query: 245 PEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQ 304
IF I++ +L + + A ++I +G ++T++ T ++ Q
Sbjct: 246 SLIFNTIFSSLL-----HSGSISWIYYADRLIEFPIGIIGVSLSTILFTSFSCSYSNNTQ 300
Query: 305 EEDKMLL 311
E K+LL
Sbjct: 301 SEYKILL 307
>LCTP_STRIN (O33654) L-lactate permease
Length = 474
Score = 36.2 bits (82), Expect = 0.11
Identities = 22/65 (33%), Positives = 34/65 (51%), Gaps = 7/65 (10%)
Query: 118 VLTILLFASVAIFPTILLPST-----PSMWVAGVTLGYGFGFLL--IITAAAIGVSLPFI 170
++++ LF + P +L+ T P V G+TL G F L I+ + +G LP I
Sbjct: 180 IVSLQLFILIVAIPFVLVSLTGEGKRPIKGVFGITLASGLAFALPQILVSNYVGAELPSI 239
Query: 171 IGSIF 175
IGS+F
Sbjct: 240 IGSLF 244
>DEDA_ECOLI (P09548) DedA protein (DSG-1 protein)
Length = 219
Score = 35.8 bits (81), Expect = 0.14
Identities = 33/177 (18%), Positives = 70/177 (38%), Gaps = 7/177 (3%)
Query: 118 VLTILLFASVAIFPTILLPSTPSMWVAGV-------TLGYGFGFLLIITAAAIGVSLPFI 170
+L ++LF + T LP ++VAG L +L++ AA +G ++ +
Sbjct: 32 ILFLILFCETGLVVTPFLPGDSLLFVAGALASLETNDLNVHMMVVLMLIAAIVGDAVNYT 91
Query: 171 IGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVATNV 230
IG +F K+ + S L H + + L R P + A ++
Sbjct: 92 IGRLFGEKLFSNPNSKIFRRSYLDKTHQFYEKHGGKTIILARFVPIVRTFAPFVAGMGHM 151
Query: 231 KYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVV 287
Y + +++G + + + Y G T+ + + L I+++ L + ++
Sbjct: 152 SYRHFAAYNVIGALLWVLLFTYAGYFFGTIPMVQDNLKLLIVGIIVVSILPGVIEII 208
>SFRB_HUMAN (Q05519) Splicing factor arginine/serine-rich 11
(Arginine-rich 54 kDa nuclear protein) (p54)
Length = 484
Score = 33.1 bits (74), Expect = 0.91
Identities = 18/55 (32%), Positives = 27/55 (48%), Gaps = 9/55 (16%)
Query: 9 GGGGGGGGWKKEVVSDVTITIEADDD---------NKGDYIKLIPGSDECLPLTA 54
GGGGGGGG EV+ ++ A + K D ++L P D LP+++
Sbjct: 22 GGGGGGGGGGTEVIQVTNVSPSASSEQMRTLFGFLGKIDELRLFPPDDSPLPVSS 76
>YQJA_ECOLI (P42614) Hypothetical protein yqjA
Length = 220
Score = 32.3 bits (72), Expect = 1.6
Identities = 21/77 (27%), Positives = 35/77 (45%), Gaps = 7/77 (9%)
Query: 118 VLTILLFASVAIFPTILLPSTPSMWVAGV-----TLGYGFGFLLIITAAAIGVSLPFIIG 172
VL ++LF + P LP + + GV +GY LL+ AA++G + +I G
Sbjct: 31 VLFVILFLENGLLPAAFLPGDSLLVLVGVLIAKGAMGYPQTILLLTVAASLGCWVSYIQG 90
Query: 173 SIFHH--KIEGWLEKYP 187
+ ++ WL P
Sbjct: 91 RWLGNTRTVQNWLSHLP 107
>YDH0_PHOPR (Q6LIN8) Hypothetical UPF0324 membrane protein PBPRB0970
Length = 326
Score = 32.3 bits (72), Expect = 1.6
Identities = 27/71 (38%), Positives = 32/71 (45%), Gaps = 3/71 (4%)
Query: 115 SPPVLTILLF--ASVAIFPTIL-LPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFII 171
S PV IL F AS PT L + + +A +G GFG L AA I+
Sbjct: 40 SSPVALILGFTLASFGFVPTKLNIAAATKKLLAYSIIGLGFGIHLDQAIAASTQGFGLIV 99
Query: 172 GSIFHHKIEGW 182
GSIF I GW
Sbjct: 100 GSIFFTLIFGW 110
>RODA_HELPY (P56098) Rod shape-determining protein rodA
Length = 381
Score = 32.0 bits (71), Expect = 2.0
Identities = 37/146 (25%), Positives = 55/146 (37%), Gaps = 35/146 (23%)
Query: 70 VWYWVKMVLLFLS--LGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVLTILLFASV 127
V+YW ++LL L +G+ + +W+ VIP I+ + P + ILL +
Sbjct: 69 VFYWACVILLALVDFMGYSKLGAQRWL--------VIPFISITLQPSEPVKIAILLLLAH 120
Query: 128 AI-----------------------FPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIG 164
I P L+ P + A + L GFG LLI+
Sbjct: 121 LIKINPPPFKGYDWGMFLKLSFYICLPAALILKQPDLGTALIVLIMGFGILLIV-GLRTR 179
Query: 165 VSLPFIIGSIFHHKIE-GWLEKYPKK 189
V LP I + I +L Y KK
Sbjct: 180 VWLPLFIALLVASPIAYHFLHDYQKK 205
>YCF4_MESVI (Q9MUN8) Photosystem I assembly protein ycf4
Length = 187
Score = 31.6 bits (70), Expect = 2.7
Identities = 21/79 (26%), Positives = 39/79 (48%), Gaps = 6/79 (7%)
Query: 57 MVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSP 116
++ E + SRR ++W +VLL S GFL V + + + +++P ++ + F P
Sbjct: 11 ILRESVVGSRRISNYWWASVVLLGAS-GFLIVGISSY-----LQYDLVPFLSAKNIVFVP 64
Query: 117 PVLTILLFASVAIFPTILL 135
L + + S I +I L
Sbjct: 65 QGLVMCFYGSAGILLSIYL 83
>SA34_CAEEL (Q10936) Serpentine receptor class alpha 34 (Sra-34
protein)
Length = 283
Score = 31.2 bits (69), Expect = 3.5
Identities = 24/80 (30%), Positives = 38/80 (47%), Gaps = 8/80 (10%)
Query: 54 AVEMVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKW---VGPYLIDKEVIPIINW- 109
AV+M ++ S+ S V WY V ++ L AV+K +GP L+ V+ W
Sbjct: 154 AVQMSKQESINSSLVVTWYLVSQIIFLGLYSILIYAVIKLKDVIGPLLVTNVVL----WC 209
Query: 110 ETETFSPPVLTILLFASVAI 129
T T++ L I++ S I
Sbjct: 210 YTYTYACMFLPIIILTSTKI 229
>DGTP_SALTY (P40733) Deoxyguanosinetriphosphate triphosphohydrolase
(EC 3.1.5.1) (dGTPase) (dGTP triphosphohydrolase)
Length = 504
Score = 31.2 bits (69), Expect = 3.5
Identities = 23/90 (25%), Positives = 40/90 (43%), Gaps = 5/90 (5%)
Query: 169 FIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVAT 228
F + ++HH W + +K S+ + GN + + RA L R + + F Y V T
Sbjct: 285 FSVEQLYHHLYHAWC--HHEKDSLFELV-VGNAWEKSRANTLSRSTEDQF--FMYLRVNT 339
Query: 229 NVKYGPYIIGSLVGMVPEIFVAIYTGILIK 258
K PY + +P+IF + L++
Sbjct: 340 LNKLVPYAAQRFIDNLPQIFAGTFNQALLE 369
>DGTP_SALTI (Q8Z9B1) Deoxyguanosinetriphosphate triphosphohydrolase
(EC 3.1.5.1) (dGTPase) (dGTP triphosphohydrolase)
Length = 504
Score = 30.8 bits (68), Expect = 4.5
Identities = 23/90 (25%), Positives = 40/90 (43%), Gaps = 5/90 (5%)
Query: 169 FIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVAT 228
F + ++HH W + +K S+ + GN + + RA L R + + F Y V T
Sbjct: 285 FSVEQLYHHLYHAW--GHHEKDSLFELV-VGNAWEKSRANTLSRSTEDQF--FMYLRVNT 339
Query: 229 NVKYGPYIIGSLVGMVPEIFVAIYTGILIK 258
K PY + +P+IF + L++
Sbjct: 340 LNKLVPYAAQRFIDNLPQIFAGTFNQALLE 369
>DGTP_ESCFE (Q59435) Deoxyguanosinetriphosphate triphosphohydrolase
(EC 3.1.5.1) (dGTPase) (dGTP triphosphohydrolase)
Length = 504
Score = 30.8 bits (68), Expect = 4.5
Identities = 23/97 (23%), Positives = 42/97 (42%), Gaps = 5/97 (5%)
Query: 169 FIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVAT 228
F + ++HH E W + +K S+ W + R+ +L R + + F Y V T
Sbjct: 285 FTVEQLYHHLHEAWGQH--EKGSLFSLVVENAW-EKSRSNSLSRSTEDQF--FMYLRVNT 339
Query: 229 NVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASN 265
K PY + +P IF + L++ ++ S+
Sbjct: 340 LNKLVPYAAQRFIDNLPAIFAGTFNHALLEDASECSD 376
>DGTP_ECOLI (P15723) Deoxyguanosinetriphosphate triphosphohydrolase
(EC 3.1.5.1) (dGTPase) (dGTP triphosphohydrolase)
Length = 504
Score = 30.8 bits (68), Expect = 4.5
Identities = 23/97 (23%), Positives = 42/97 (42%), Gaps = 5/97 (5%)
Query: 169 FIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVAT 228
F + ++HH E W + +K S+ W + R+ +L R + + F Y V T
Sbjct: 285 FTVEQLYHHLHEAWGQH--EKGSLFSLVVENAW-EKSRSNSLSRSTEDQF--FMYLRVNT 339
Query: 229 NVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASN 265
K PY + +P IF + L++ ++ S+
Sbjct: 340 LNKLVPYAAQRFIDNLPAIFAGTFNHALLEDASECSD 376
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,235,813
Number of Sequences: 164201
Number of extensions: 1641825
Number of successful extensions: 7550
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 7534
Number of HSP's gapped (non-prelim): 32
length of query: 312
length of database: 59,974,054
effective HSP length: 110
effective length of query: 202
effective length of database: 41,911,944
effective search space: 8466212688
effective search space used: 8466212688
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146564.14


