

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146559.3 + phase: 0
(130 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
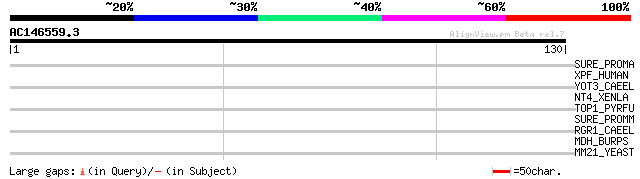
Score E
Sequences producing significant alignments: (bits) Value
SURE_PROMA (Q7VAV8) Acid phosphatase surE (EC 3.1.3.2) 30 0.71
XPF_HUMAN (Q92889) DNA repair endonuclease XPF (EC 3.1.-.-) (DNA... 29 2.1
YOT3_CAEEL (P34649) Hypothetical protein ZK632.3 in chromosome III 28 2.7
NT4_XENLA (P24727) Neurotrophin-4 precursor (NT-4) 28 3.5
TOP1_PYRFU (O73954) DNA topoisomerase I (EC 5.99.1.2) (Omega-pro... 28 4.6
SURE_PROMM (Q7V8I0) Acid phosphatase surE (EC 3.1.3.2) 28 4.6
RGR1_CAEEL (Q03570) Mediator complex subunit rgr-1 28 4.6
MDH_BURPS (P80536) Malate dehydrogenase (EC 1.1.1.37) 28 4.6
MM21_YEAST (P38632) DNA repair protein MMS21 27 7.9
>SURE_PROMA (Q7VAV8) Acid phosphatase surE (EC 3.1.3.2)
Length = 262
Score = 30.4 bits (67), Expect = 0.71
Identities = 18/41 (43%), Positives = 23/41 (55%), Gaps = 6/41 (14%)
Query: 32 KTFRSASDSDAVWNQFLPSDISSIISHSPSLTNIPTKKDLY 72
K SA D W PSD+S I ++SPSLT P + DL+
Sbjct: 216 KDLESAGDGPKEW----PSDVSQIETNSPSLT--PIQPDLF 250
>XPF_HUMAN (Q92889) DNA repair endonuclease XPF (EC 3.1.-.-) (DNA
excision repair protein ERCC-4) (DNA-repair protein
complementing XP-F cell) (Xeroderma pigmentosum group F
complementing protein)
Length = 905
Score = 28.9 bits (63), Expect = 2.1
Identities = 21/79 (26%), Positives = 33/79 (41%), Gaps = 3/79 (3%)
Query: 19 TTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNIPTKKDLYLALSDR 78
T +A L ++ KTF S ++ VW +F D S I S P K+ S +
Sbjct: 414 TLGAEAFLLRLYRKTFEKDSKAEEVWMKFRKEDSSKRIRKSHKRPKDPQNKE---RASTK 470
Query: 79 PVIIDHDKKSIQLERKSGK 97
+ K+ + L + GK
Sbjct: 471 ERTLKKKKRKLTLTQMVGK 489
>YOT3_CAEEL (P34649) Hypothetical protein ZK632.3 in chromosome III
Length = 510
Score = 28.5 bits (62), Expect = 2.7
Identities = 15/42 (35%), Positives = 24/42 (56%), Gaps = 2/42 (4%)
Query: 46 QFLPSDISSIISHSPSL--TNIPTKKDLYLALSDRPVIIDHD 85
QFL DI +II+ + N+PT L+ ++D ++ DHD
Sbjct: 420 QFLTRDIQNIITFFTRIGTPNLPTYVQLFNLITDLDMVEDHD 461
>NT4_XENLA (P24727) Neurotrophin-4 precursor (NT-4)
Length = 236
Score = 28.1 bits (61), Expect = 3.5
Identities = 37/127 (29%), Positives = 53/127 (41%), Gaps = 22/127 (17%)
Query: 11 CIAAIVSRTTPLDAGRLSVFSKTFRSA---SDSDAVWNQFLPSDISSIIS---------H 58
C A SRTT LD G + R + + S + N F P D+SS S +
Sbjct: 17 CAAPFQSRTTDLDYGPDKTSEASDRQSVPNNFSHVLQNGFFP-DLSSTYSSMAGKDWNLY 75
Query: 59 SPSLT---NIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSAR-SLAI----- 109
SP +T P+ L + V + K+ +L+R SG LS R L++
Sbjct: 76 SPRVTLSSEEPSGPPLLFLSEETVVHPEPANKTSRLKRASGSDSVSLSRRGELSVCDSVN 135
Query: 110 VWGDDRR 116
VW D+R
Sbjct: 136 VWVTDKR 142
>TOP1_PYRFU (O73954) DNA topoisomerase I (EC 5.99.1.2)
(Omega-protein) (Relaxing enzyme) (Untwisting enzyme)
(Swivelase) [Contains: Endonuclease PI-PfuI (EC 3.1.-.-)
(Pfu topA intein)]
Length = 1060
Score = 27.7 bits (60), Expect = 4.6
Identities = 12/41 (29%), Positives = 22/41 (53%)
Query: 67 TKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSL 107
+++ LY+ S PV D D++S R G Y++ +S+
Sbjct: 618 SRRKLYVETSQVPVFTDFDERSYDFPRILGGDIYIIGIKSI 658
>SURE_PROMM (Q7V8I0) Acid phosphatase surE (EC 3.1.3.2)
Length = 269
Score = 27.7 bits (60), Expect = 4.6
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 6/41 (14%)
Query: 32 KTFRSASDSDAVWNQFLPSDISSIISHSPSLTNIPTKKDLY 72
K +A D W PSD++ I ++SPSLT P + DL+
Sbjct: 216 KDLETAGDGPRDW----PSDVAQIETNSPSLT--PIQPDLF 250
>RGR1_CAEEL (Q03570) Mediator complex subunit rgr-1
Length = 1374
Score = 27.7 bits (60), Expect = 4.6
Identities = 25/89 (28%), Positives = 39/89 (43%), Gaps = 10/89 (11%)
Query: 7 LPEGCIAAI--VSRTTPLDAGRLSVFS----KTFRSASDSDAVWNQFLPSDISSII---- 56
L EGC A+ VSR P AG +FS +T +D A+ P +
Sbjct: 867 LLEGCKIAVRDVSRYRPRCAGLFQLFSNIDSETTAIMNDEIAIPQSDNPQTAGPTMWTPE 926
Query: 57 SHSPSLTNIPTKKDLYLALSDRPVIIDHD 85
SL P + D +A++ +P+++ HD
Sbjct: 927 QFMDSLDERPEEIDPRMAITSQPILMSHD 955
>MDH_BURPS (P80536) Malate dehydrogenase (EC 1.1.1.37)
Length = 326
Score = 27.7 bits (60), Expect = 4.6
Identities = 23/82 (28%), Positives = 37/82 (45%), Gaps = 19/82 (23%)
Query: 42 AVWNQFLPSDISSIISHSPSLTNIPTKKDLYLALSDRPVI----------IDHDKKSIQL 91
A N+ D+ ++ +P+ TN Y+A+ P + +DH++ QL
Sbjct: 114 AALNEVASRDVKVLVVGNPANTNA------YIAMKSAPDLPKKNFTAMLRLDHNRALSQL 167
Query: 92 ERKSGKKCYMLSARSLAIVWGD 113
KSGK + S LA VWG+
Sbjct: 168 AAKSGKP--VASIEKLA-VWGN 186
>MM21_YEAST (P38632) DNA repair protein MMS21
Length = 267
Score = 26.9 bits (58), Expect = 7.9
Identities = 15/39 (38%), Positives = 23/39 (58%), Gaps = 5/39 (12%)
Query: 73 LALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVW 111
+AL+D P+ KS+ L KSGK + L AR L+ ++
Sbjct: 1 MALNDNPI-----PKSVPLHPKSGKYFHNLHARDLSNIY 34
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.132 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,427,534
Number of Sequences: 164201
Number of extensions: 517957
Number of successful extensions: 1268
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1266
Number of HSP's gapped (non-prelim): 9
length of query: 130
length of database: 59,974,054
effective HSP length: 106
effective length of query: 24
effective length of database: 42,568,748
effective search space: 1021649952
effective search space used: 1021649952
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146559.3


