

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.6 + phase: 0 /pseudo
(799 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
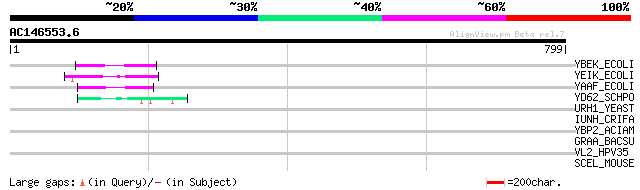
YBEK_ECOLI (P41409) Hypothetical protein ybeK 59 4e-08
YEIK_ECOLI (P33022) Hypothetical protein yeiK 55 1e-06
YAAF_ECOLI (P22564) Hypothetical protein yaaF 50 2e-05
YD62_SCHPO (Q10314) Hypothetical protein C17G8.02 in chromosome I 48 1e-04
URH1_YEAST (Q04179) Uridine nucleosidase (EC 3.2.2.3) (Uridine r... 45 0.001
IUNH_CRIFA (Q27546) Inosine-uridine preferring nucleoside hydrol... 42 0.005
YBP2_ACIAM (P32986) Hypothetical protein in bps2 5'region (ORF2)... 34 1.3
GRAA_BACSU (P07868) Spore germination protein A1 33 3.9
VL2_HPV35 (P27234) Minor capsid protein L2 32 5.1
SCEL_MOUSE (Q9EQG3) Sciellin 32 8.7
>YBEK_ECOLI (P41409) Hypothetical protein ybeK
Length = 311
Score = 59.3 bits (142), Expect = 4e-08
Identities = 37/117 (31%), Positives = 53/117 (44%), Gaps = 26/117 (22%)
Query: 95 SAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCG 154
SA P+T++ TG TN+A+ L ++P L + + IMGG +
Sbjct: 115 SAEPVTIVSTGPQTNVALLLNSHPELHSKIARIVIMGGAMGLG----------------- 157
Query: 155 DTGNVLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEFFDEF 211
N P AE+NI+ DP AA V SGIP+ + LD T+ I E + F
Sbjct: 158 ---------NWTPAAEFNIYVDPEAAEIVFQSGIPVVMAGLDVTHKAQIHVEDTERF 205
>YEIK_ECOLI (P33022) Hypothetical protein yeiK
Length = 313
Score = 54.7 bits (130), Expect = 1e-06
Identities = 40/145 (27%), Positives = 63/145 (42%), Gaps = 36/145 (24%)
Query: 80 PLRQPTTQQV--------LIDKISA--GPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYI 129
P+ +P T+Q +ID + A G ITL+ G +N+A+ + P + + + +
Sbjct: 90 PVFEPLTRQAESTHAVKYIIDTLMASDGDITLVPVGPLSNIAVAMRMQPAILPKIREIVL 149
Query: 130 MGGVIRSKTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAEYNIFGDPFAAYKVIHSGIP 189
MGG TGN P AE+NIF DP AA V SG+P
Sbjct: 150 MGGAY--------------------GTGNF------TPSAEFNIFADPEAARVVFTSGVP 183
Query: 190 ITLVPLDATNTIPISEEFFDEFEKS 214
+ ++ LD TN + + E++
Sbjct: 184 LVMMGLDLTNQTVCTPDVIARMERA 208
>YAAF_ECOLI (P22564) Hypothetical protein yaaF
Length = 304
Score = 50.4 bits (119), Expect = 2e-05
Identities = 33/110 (30%), Positives = 45/110 (40%), Gaps = 26/110 (23%)
Query: 98 PITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTG 157
P+TL+ G TN+A+ L P K + + IMGG C
Sbjct: 117 PVTLVAIGPLTNIALLLSQCPECKPYIRRLVIMGGSAGRGNC------------------ 158
Query: 158 NVLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATNTIPISEEF 207
P AE+NI DP AA V SGI I + LD TN ++ ++
Sbjct: 159 --------TPNAEFNIAADPEAAACVFRSGIEIVMCGLDVTNQAILTPDY 200
>YD62_SCHPO (Q10314) Hypothetical protein C17G8.02 in chromosome I
Length = 330
Score = 47.8 bits (112), Expect = 1e-04
Identities = 46/187 (24%), Positives = 68/187 (35%), Gaps = 56/187 (29%)
Query: 98 PITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTG 157
P+TL+ TG TN+A+ L P + N+E MGG G
Sbjct: 132 PVTLVATGPLTNIALLLATYPSVTDNIERFIFMGG--------------------STGIG 171
Query: 158 NVLTNYNANPYAEYNIFGDPFAAYKVIHSGI---PITLVPLDATNTI------------- 201
N+ + AE+N++ DP AA V+ + + +VPLD T+ +
Sbjct: 172 NITSQ------AEFNVYADPEAARLVLETKSLIGKLFMVPLDVTHKVLLDANIIQLLRQH 225
Query: 202 --PISEEFFDEFEKSQDTYEAQYCFKSLKMAHDT-----------WFDNQFYTSYFMWDS 248
P S + Q TYE Y ++ HD W Y + + DS
Sbjct: 226 SNPFSSTLVELMTVFQQTYENVYGIRNGVPVHDVCAVALALWPSLWTSRSMYVTVSL-DS 284
Query: 249 FTSGVAV 255
T G V
Sbjct: 285 LTLGRTV 291
>URH1_YEAST (Q04179) Uridine nucleosidase (EC 3.2.2.3) (Uridine
ribohydrolase)
Length = 378
Score = 44.7 bits (104), Expect = 0.001
Identities = 41/146 (28%), Positives = 59/146 (40%), Gaps = 23/146 (15%)
Query: 97 GPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDT 156
G I+ + TGA T LA P+LKK+V+++ IMGG + C N S+
Sbjct: 161 GEISFVSTGALTTLATVFRCKPYLKKSVKYISIMGGGLHGLGNCNPNLSAE--------- 211
Query: 157 GNVLTNYNANPYAEYNIFGDPFAAYKVI-------HSGI---PITLVPLDATNTIPISEE 206
N +P A IF DP K I H I + + + N + E
Sbjct: 212 ----FNVWIDPDAANYIFRDPDVKDKCIVVPLNLTHKAIATYKVNEMIYNEKNNSKLREL 267
Query: 207 FFDEFEKSQDTYEAQYCFKSLKMAHD 232
F + F+ TY+ F+S HD
Sbjct: 268 FLELFQFFAHTYKDMQGFESGPPIHD 293
>IUNH_CRIFA (Q27546) Inosine-uridine preferring nucleoside hydrolase
(EC 3.2.2.1) (IU-nucleoside hydrolase) (IU-NH) (Purine
nucleosidase)
Length = 314
Score = 42.4 bits (98), Expect = 0.005
Identities = 41/161 (25%), Positives = 63/161 (38%), Gaps = 45/161 (27%)
Query: 99 ITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRSKTCCTKNASSSCIPSKCGDTGN 158
ITL+ TG TN+A+ P + V+ V +MGG
Sbjct: 120 ITLVPTGGLTNIAMAARLEPRIVDRVKEVVLMGGGYHEG--------------------- 158
Query: 159 VLTNYNANPYAEYNIFGDPFAAYKVIHSGIPITLVPLDATN----TIPISEEFFDEFEKS 214
NA AE+NI DP AA+ V + +T+V LD T+ T PI + K
Sbjct: 159 -----NATSVAEFNIIIDPEAAHIVFNESWQVTMVGLDLTHQALATPPILQRV-----KE 208
Query: 215 QDTYEAQYCFKSLKMAHDTWFDNQFYTSYFMWDSFTSGVAV 255
DT A++ + + +YT + + + + AV
Sbjct: 209 VDTNPARFMLEIM----------DYYTKIYQSNRYMAAAAV 239
>YBP2_ACIAM (P32986) Hypothetical protein in bps2 5'region (ORF2)
(Fragment)
Length = 171
Score = 34.3 bits (77), Expect = 1.3
Identities = 24/105 (22%), Positives = 43/105 (40%), Gaps = 27/105 (25%)
Query: 77 KYTPLRQPTTQQVL-IDKISAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIR 135
K +P ++ ++ + K G + ++ TNLA+ + +P + K ++ V+IMGG
Sbjct: 93 KISPEKEHAIDAIIRLSKEYEGELEILAVSPLTNLALAYLKDPTIVKRIKKVWIMGG--- 149
Query: 136 SKTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAEYNIFGDPFAA 180
+ N P AE+N + DP AA
Sbjct: 150 -----------------------AFSRGNTTPIAEFNFWVDPEAA 171
>GRAA_BACSU (P07868) Spore germination protein A1
Length = 482
Score = 32.7 bits (73), Expect = 3.9
Identities = 22/66 (33%), Positives = 31/66 (46%), Gaps = 6/66 (9%)
Query: 65 GIRKAF-LPQGKRKYTPLRQPTTQQVLIDKISAGPITLIVTGAHTNLAIFLMNNPHLKKN 123
G+ KA+ L GK+K L +PTT +K+ GP V TNLA+ H K
Sbjct: 106 GLDKAYILTTGKKKTRSLTEPTT-----EKVVRGPKVAFVEDIDTNLALIRQRTSHPKLI 160
Query: 124 VEHVYI 129
+ + I
Sbjct: 161 TKKIMI 166
>VL2_HPV35 (P27234) Minor capsid protein L2
Length = 469
Score = 32.3 bits (72), Expect = 5.1
Identities = 29/101 (28%), Positives = 40/101 (38%), Gaps = 9/101 (8%)
Query: 10 GVGGEGGILPNGTILPNVGGYLPIIEQGMTTAGYCRYRQAIPVGFGGRLDID-TNLGIRK 68
G GG G +P GT P +P I +T ++IP+ G LD +L
Sbjct: 65 GTGGRSGYVPLGTTPPTAATNIP-IRPPVTV-------ESIPLDTIGPLDSSIVSLVEET 116
Query: 69 AFLPQGKRKYTPLRQPTTQQVLIDKISAGPITLIVTGAHTN 109
+F+ G TP PTT + P L VT T+
Sbjct: 117 SFIESGAPVVTPRVPPTTGFTITTSTDTTPAILDVTSISTH 157
>SCEL_MOUSE (Q9EQG3) Sciellin
Length = 653
Score = 31.6 bits (70), Expect = 8.7
Identities = 23/94 (24%), Positives = 38/94 (39%)
Query: 77 KYTPLRQPTTQQVLIDKISAGPITLIVTGAHTNLAIFLMNNPHLKKNVEHVYIMGGVIRS 136
K TP ++Q L + I+ P T+ T H +L F+ NP + N + + + IR
Sbjct: 389 KVTPSANRSSQHSLDELINTSPQTIKTTARHQDLDKFIKVNPDVLTNNQRNHDVDSTIRG 448
Query: 137 KTCCTKNASSSCIPSKCGDTGNVLTNYNANPYAE 170
T+ S + + + L N N E
Sbjct: 449 NPTGTRCEQSEELDNLIKVKPSALRNTNGGQEVE 482
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 97,338,395
Number of Sequences: 164201
Number of extensions: 4388306
Number of successful extensions: 8956
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 8937
Number of HSP's gapped (non-prelim): 16
length of query: 799
length of database: 59,974,054
effective HSP length: 118
effective length of query: 681
effective length of database: 40,598,336
effective search space: 27647466816
effective search space used: 27647466816
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC146553.6


