

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146551.3 - phase: 1 /pseudo
(379 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
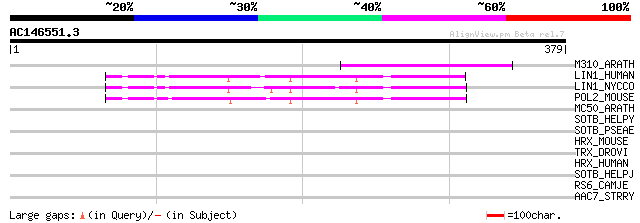
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 75 4e-13
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 55 2e-07
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 54 5e-07
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 46 2e-04
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 34 0.54
SOTB_HELPY (O25797) Probable sugar efflux transporter 32 2.7
SOTB_PSEAE (Q9HWR7) Probable sugar efflux transporter 31 4.5
HRX_MOUSE (P55200) Zinc finger protein HRX (ALL-1) (Fragment) 31 4.5
TRX_DROVI (Q24742) Trithorax protein 31 5.9
HRX_HUMAN (Q03164) Zinc finger protein HRX (ALL-1) (Trithorax-li... 31 5.9
SOTB_HELPJ (Q9ZK31) Probable sugar efflux transporter 30 7.7
RS6_CAMJE (Q9ZAH3) 30S ribosomal protein S6 30 7.7
AAC7_STRRY (P30180) Aminoglycoside N(3')-acetyltransferase VII (... 30 7.7
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 74.7 bits (182), Expect = 4e-13
Identities = 40/119 (33%), Positives = 63/119 (52%), Gaps = 2/119 (1%)
Query: 227 SLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKA-EGGLGVRR 285
+LPVY +S F+ + + S F+W E RKI+W+ W +C SK +GGLG R
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 286 MGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEGGRIR-EGGRLSSAWWKMVCHIRE 343
+G FN +LL K +R++ + L R+L+++Y + G S W+ + H RE
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSRYFPHSSMMECSVGTRPSYAWRSIIHGRE 120
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 55.5 bits (132), Expect = 2e-07
Identities = 59/254 (23%), Positives = 108/254 (42%), Gaps = 22/254 (8%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP L + E VL A+ +G EV+++ FADD ++ E ++
Sbjct: 661 PLSPLLPNIVLE---VLARAIRQEKEIKGIQLGK-EEVKLS--LFADDMIVYLENPIVSA 714
Query: 126 RTMMAVLLLFEEISGLKVNFHKS---MLTGVNISDSWLVEASALMNCCRGNFTFVYLGVP 182
+ ++ ++ F ++SG K+N KS + T ++S ++ + YLG+
Sbjct: 715 QNLLKLISNFSKVSGYKINVQKSQAFLYTNNRQTESQIMSELPFTIASK---RIKYLGIQ 771
Query: 183 IGGDPRKL--SFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSF--FKA 238
+ D + L +KP+++ I + W N S GR+ ++K+ + +Y + K
Sbjct: 772 LTRDVKDLFKENYKPLLNEIKEDTNKWKNIPCSWVGRINIVKMAILPKVIYRFNAIPIKL 831
Query: 239 PAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWC 298
P + +E F W N+K A I T+ GG+ + + + + K
Sbjct: 832 PMTFFTELEKTTLKFIW-----NQKRAHIAKSTLSQKNKAGGITLPDFKLYYKATVTKTA 886
Query: 299 WRMLTERD-ELWYR 311
W RD + W R
Sbjct: 887 WYWYQNRDIDQWNR 900
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 54.3 bits (129), Expect = 5e-07
Identities = 65/262 (24%), Positives = 113/262 (42%), Gaps = 36/262 (13%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP LF + E VL A+ +G+ E++++ FADD ++ E +
Sbjct: 661 PLSPLLFNIVME---VLAIAIREEKAIKGIHIGS-EEIKLS--LFADDMIVYLENTRDST 714
Query: 126 RTMMAVLLLFEEISGLKVNFHKS---MLTGVNISDSWLVEASALMNCCRGNFTFV----- 177
++ V+ + +SG K+N HKS + T N ++ + ++ FT V
Sbjct: 715 TKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQAEKTVKDSIP--------FTVVPKKMK 766
Query: 178 YLGVPIGGDPRKL--SFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSF 235
YLGV + D + L ++ + I ++ W N S GR+ ++K +S LP +F
Sbjct: 767 YLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVK--MSILPKAIYNF 824
Query: 236 ----FKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNV 291
KAP +E I F W N+K I + GG+ + + +
Sbjct: 825 NAIPIKAPLSYFKDLEKIILHFIW-----NQKKPQIAKTLLSNKNKAGGITLPDLRLYYK 879
Query: 292 SLLGKWCWRMLTERD-ELWYRV 312
S++ K W R+ ++W R+
Sbjct: 880 SIVIKTAWYWHKNREVDVWNRI 901
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 45.8 bits (107), Expect = 2e-04
Identities = 58/254 (22%), Positives = 105/254 (40%), Gaps = 20/254 (7%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PLSP+LF + E VL A+ +G EV+++ L ADD ++ +
Sbjct: 688 PLSPYLFNIVLE---VLARAIRQQKEIKGIQIGK-EEVKISLL--ADDMIVYISDPKNST 741
Query: 126 RTMMAVLLLFEEISGLKVNFHKSM--LTGVNISDSWLVEASALMNCCRGNFTFVYLGVPI 183
R ++ ++ F E+ G K+N +KSM L N + + + N YLGV +
Sbjct: 742 RELLNLINSFGEVVGYKINSNKSMAFLYTKNKQAEKEIRETTPFSIVTNNIK--YLGVTL 799
Query: 184 GGDPRKL--SFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSF--FKAP 239
+ + L +K + I L W + S GR+ ++K+ + +Y + K P
Sbjct: 800 TKEVKDLYDKNFKSLKKEIKEDLRRWKDLPCSWIGRINIVKMAILPKAIYRFNAIPIKIP 859
Query: 240 AGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCW 299
+ +E F W N K I + + GG+ + + + +++ K W
Sbjct: 860 TQFFNELEGAICKFVW-----NNKKPRIAKSLLKDKRTSGGITMPDLKLYYRAIVIKTAW 914
Query: 300 RMLTERD-ELWYRV 312
+R + W R+
Sbjct: 915 YWYRDRQVDQWNRI 928
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 34.3 bits (77), Expect = 0.54
Identities = 20/49 (40%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDT 114
PLSP+LF+L E + L V N S R+ HL FADDT
Sbjct: 33 PLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSP-RINHLLFADDT 80
>SOTB_HELPY (O25797) Probable sugar efflux transporter
Length = 391
Score = 32.0 bits (71), Expect = 2.7
Identities = 30/128 (23%), Positives = 58/128 (44%), Gaps = 8/128 (6%)
Query: 128 MMAVLLLFEEISGLKVNFHKSMLTGVNISDS----WLVEASALMNCCRGNFTFVYLGVPI 183
+ A+ +L +S L NF +L+ + I+ + W + AS ++ N LG+
Sbjct: 85 LFALFILSHILSALAWNFWVLLLSRMGIAFAHSIFWSITASLVIRVAPRNKKQQALGLLA 144
Query: 184 GGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGII 243
G + P + RI+G++ W + IGG L+ +++ L + S AG +
Sbjct: 145 LGSSLAMILGLP-LGRIIGQILDWRSTFGVIGGVATLIALLMWKLLPHLPS---RNAGTL 200
Query: 244 SSIESIFK 251
+S+ + K
Sbjct: 201 ASVPVLMK 208
>SOTB_PSEAE (Q9HWR7) Probable sugar efflux transporter
Length = 396
Score = 31.2 bits (69), Expect = 4.5
Identities = 30/113 (26%), Positives = 50/113 (43%), Gaps = 9/113 (7%)
Query: 126 RTMMAVLLLF---EEISGLKVNFHKSMLTGVNISDS----WLVEASALMNCCRGNFTFVY 178
+ ++ V LLF +SGL +F ML+ + I+ + W + AS +
Sbjct: 79 KLLVGVFLLFIASHVLSGLAWSFQVLMLSRIGIAFAHAVFWAITASLAVRVAPPGQQAKA 138
Query: 179 LGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLS-SLPV 230
LG+ G + P + R+VG W ++I G VL + L+ SLP+
Sbjct: 139 LGLLATGTTLAMVLGIP-LGRVVGEALGWRTTFMAIAGLSVLTLLYLARSLPL 190
>HRX_MOUSE (P55200) Zinc finger protein HRX (ALL-1) (Fragment)
Length = 3866
Score = 31.2 bits (69), Expect = 4.5
Identities = 19/78 (24%), Positives = 34/78 (43%), Gaps = 7/78 (8%)
Query: 108 LQFADDT-------LIIGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWL 160
L + DD+ L IG+ W +V + +FE+ G N H +++ G + +
Sbjct: 1779 LMYGDDSANDAGRLLYIGQNEWTHVNCALWSAEVFEDDDGSLKNVHMAVIRGKQLRCEFC 1838
Query: 161 VEASALMNCCRGNFTFVY 178
+ A + CC + T Y
Sbjct: 1839 QKPGATVGCCLTSCTSNY 1856
>TRX_DROVI (Q24742) Trithorax protein
Length = 3828
Score = 30.8 bits (68), Expect = 5.9
Identities = 16/55 (29%), Positives = 24/55 (43%)
Query: 115 LIIGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNC 169
L G CW+++ M +FEEI G N H ++ G I + A + C
Sbjct: 1729 LYCGHDCWVHINCAMWSAEVFEEIDGSLQNVHSAVARGRMIKCTVCGNRGATVGC 1783
>HRX_HUMAN (Q03164) Zinc finger protein HRX (ALL-1) (Trithorax-like
protein)
Length = 3969
Score = 30.8 bits (68), Expect = 5.9
Identities = 19/78 (24%), Positives = 34/78 (43%), Gaps = 7/78 (8%)
Query: 108 LQFADDT-------LIIGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWL 160
L + DD+ L IG+ W +V + +FE+ G N H +++ G + +
Sbjct: 1877 LTYGDDSANDAGRLLYIGQNEWTHVNCALWSAEVFEDDDGSLKNVHMAVIRGKQLRCEFC 1936
Query: 161 VEASALMNCCRGNFTFVY 178
+ A + CC + T Y
Sbjct: 1937 QKPGATVGCCLTSCTSNY 1954
>SOTB_HELPJ (Q9ZK31) Probable sugar efflux transporter
Length = 391
Score = 30.4 bits (67), Expect = 7.7
Identities = 29/128 (22%), Positives = 57/128 (43%), Gaps = 8/128 (6%)
Query: 128 MMAVLLLFEEISGLKVNFHKSMLTGVNISDS----WLVEASALMNCCRGNFTFVYLGVPI 183
+ A+ + +S L NF +L+ + I+ + W + AS ++ N LG+
Sbjct: 85 LFALFIFSHILSALAWNFWVLLLSRMGIAFAHSIFWSITASLVIRVAPRNKKQQALGLLA 144
Query: 184 GGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGII 243
G + P + RI+G++ W + IGG L+ +++ L + S AG +
Sbjct: 145 LGSSLAMILGLP-LGRIIGQILDWRSTFGVIGGVATLIMLLMWKLLPHLPS---RNAGTL 200
Query: 244 SSIESIFK 251
+S+ + K
Sbjct: 201 ASVPILMK 208
>RS6_CAMJE (Q9ZAH3) 30S ribosomal protein S6
Length = 125
Score = 30.4 bits (67), Expect = 7.7
Identities = 16/63 (25%), Positives = 32/63 (50%), Gaps = 4/63 (6%)
Query: 189 KLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIES 248
KL F K V+ + + + ++ +G R + K+ YF+ +FKAP +I+ +E
Sbjct: 22 KLEFVKEVLTKNSAEIET----VVPMGTRKLAYKIKKYERGTYFVIYFKAPTNLIAELER 77
Query: 249 IFK 251
+ +
Sbjct: 78 VLR 80
>AAC7_STRRY (P30180) Aminoglycoside N(3')-acetyltransferase VII (EC
2.3.1.81) (ACC(3)-VII) (Aminocyclitol
3-N-acetyltransferase type VII)
Length = 288
Score = 30.4 bits (67), Expect = 7.7
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 4/69 (5%)
Query: 56 SD*AWSSPRGPLSPFLFLLAAEGFTVLMNAVVG--TDMYHSFGVGNGSEVRVTHLQFADD 113
+D W P GP SP L+A G +L+ A + T ++H+ + + R + +
Sbjct: 145 ADHPWDDPHGPDSPLARLVAMGGRVLLLGAPLEALTLLHHAEALADAPGKR--FVDYEQP 202
Query: 114 TLIIGEKCW 122
L+ GE+ W
Sbjct: 203 ILVDGERVW 211
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.146 0.494
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 46,404,558
Number of Sequences: 164201
Number of extensions: 2004295
Number of successful extensions: 4810
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 4800
Number of HSP's gapped (non-prelim): 17
length of query: 379
length of database: 59,974,054
effective HSP length: 112
effective length of query: 267
effective length of database: 41,583,542
effective search space: 11102805714
effective search space used: 11102805714
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146551.3


