

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.4 - phase: 0 /pseudo
(322 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
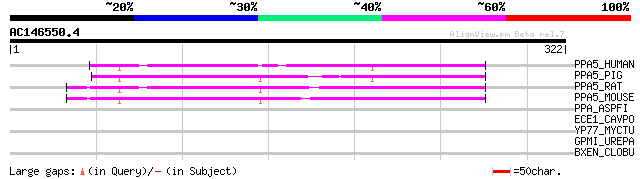
PPA5_HUMAN (P13686) Tartrate-resistant acid phosphatase type 5 p... 111 3e-24
PPA5_PIG (P09889) Tartrate-resistant acid phosphatase type 5 pre... 108 1e-23
PPA5_RAT (P29288) Tartrate-resistant acid phosphatase type 5 pre... 104 3e-22
PPA5_MOUSE (Q05117) Tartrate-resistant acid phosphatase type 5 p... 103 8e-22
PPA_ASPFI (Q12546) Acid phosphatase precursor (EC 3.1.3.2) (pH 6... 35 0.33
ECE1_CAVPO (P97739) Endothelin-converting enzyme 1 (EC 3.4.24.71... 32 2.8
YP77_MYCTU (Q50644) Hypothetical protein Rv2577/MT2654 31 4.7
GPMI_UREPA (Q9PQW1) 2,3-bisphosphoglycerate-independent phosphog... 31 4.7
BXEN_CLOBU (Q06366) Botulinum neurotoxin type E, nontoxic component 30 8.1
>PPA5_HUMAN (P13686) Tartrate-resistant acid phosphatase type 5
precursor (EC 3.1.3.2) (TR-AP) (Tartrate-resistant acid
ATPase) (TrATPase)
Length = 323
Score = 111 bits (277), Expect = 3e-24
Identities = 74/245 (30%), Positives = 121/245 (49%), Gaps = 25/245 (10%)
Query: 47 SLNFLVVGDWGRKGNY---------NQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEG 97
+L F+ VGDWG N N +A + I+G DF++S GDNFY G++
Sbjct: 25 ALRFVAVGDWGGVPNAPFHTAREMANAKEIARTVQILG----ADFILSLGDNFYFTGVQD 80
Query: 98 VDDPTFYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSFIL 156
++D F E+F ++++ SL+K+ WY + GNHD+ G+V AQ++ K RW F
Sbjct: 81 INDKRFQETFEDVFSDRSLRKVPWYVLAGNHDHLGNVSAQIAYSKISK--RWNFPSPFYR 138
Query: 157 DGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVK-----SK 211
+ F T + + D + + L ++ R L L+ +K ++
Sbjct: 139 ----LHFKIPQTNVSVAIFMLDTVTLCGNSDDFLSQQPERPRLTARTQLSWLKKQLAAAR 194
Query: 212 AKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERHGEA 271
+ +V GH+ + S+ HG T L +QL P+L + + AY+ GHDH LQ++ D G
Sbjct: 195 EDYVLVAGHYPVWSIAEHGPTHCLVKQLRPLLATYGVTAYLCGHDHNLQYLQDENGVGYV 254
Query: 272 ISSHG 276
+S G
Sbjct: 255 LSGAG 259
>PPA5_PIG (P09889) Tartrate-resistant acid phosphatase type 5
precursor (EC 3.1.3.2) (TR-AP) (Tartrate-resistant acid
ATPase) (TrATPase) (Uteroferrin) (UF)
Length = 340
Score = 108 bits (271), Expect = 1e-23
Identities = 78/245 (31%), Positives = 119/245 (47%), Gaps = 25/245 (10%)
Query: 48 LNFLVVGDWGRKGNY-----NQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPT 102
L F+ VGDWG N + A + L DF++S GDNFY G+ D
Sbjct: 34 LRFVAVGDWGGVPNAPFHTAREMANAKAIATTVKTLGADFILSLGDNFYFTGVHDAKDKR 93
Query: 103 FYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQK----DSRWVCLRSFILD 157
F E+F ++++ PSL+ + W+ + GNHD+ G+V AQ++ K S + LR I
Sbjct: 94 FQETFEDVFSDPSLRNVPWHVLAGNHDHLGNVSAQIAYSKISKRWNFPSPYYRLRFKIPR 153
Query: 158 GGI-VEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVK-----SK 211
+ V F +DT ++ D+ P E R L LA +K +K
Sbjct: 154 SNVSVAIFMLDTVTLCG--------NSDDFVSQQP-ERPRNLALARTQLAWIKKQLAAAK 204
Query: 212 AKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERHGEA 271
+ +V GH+ + S+ HG T L +QLLP+L ++ + AY+ GHDH LQ++ D G
Sbjct: 205 EDYVLVAGHYPVWSIAEHGPTHCLVKQLLPLLTTHKVTAYLCGHDHNLQYLQDENGLGFV 264
Query: 272 ISSHG 276
+S G
Sbjct: 265 LSGAG 269
>PPA5_RAT (P29288) Tartrate-resistant acid phosphatase type 5
precursor (EC 3.1.3.2) (TR-AP) (Tartrate-resistant acid
ATPase) (TrATPase)
Length = 327
Score = 104 bits (259), Expect = 3e-22
Identities = 80/259 (30%), Positives = 122/259 (46%), Gaps = 26/259 (10%)
Query: 34 LPRFEHHLKPQQQSLNFLVVGDWGRKGNY---------NQSLVAHQMGIVGDNLNIDFVI 84
LP H P +L F+ VGDWG N N +A + I+G DF++
Sbjct: 15 LPLLAHCTAPAS-TLRFVAVGDWGGVPNAPFHTAREMANAKEIARTVQIMG----ADFIM 69
Query: 85 STGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQ 143
S GDNFY G+ +D F E+F ++++ +L+ I WY + GNHD+ G+V AQ++
Sbjct: 70 SLGDNFYFTGVHDANDKRFQETFEDVFSDRALRNIPWYVLAGNHDHLGNVSAQIAYSKIS 129
Query: 144 K----DSRWVCLRSFILDGGI-VEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAE 198
K S + LR + I V F +DT + +PR+ A
Sbjct: 130 KRWNFPSPYYRLRFKVPRSNITVAIFMLDTVMLCGN-----SDDFVSQQPEMPRDLGVAR 184
Query: 199 L-LKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDH 257
L + L +K + +V GH+ I S+ HG T+ L + L P+L + + AY+ GHDH
Sbjct: 185 TQLSWLKKQLAAAKEDYVLVAGHYPIWSIAEHGPTRCLVKNLRPLLAAYGVTAYLCGHDH 244
Query: 258 CLQHIIDNERHGEAISSHG 276
LQ++ D G +S G
Sbjct: 245 NLQYLQDENGVGYVLSGAG 263
>PPA5_MOUSE (Q05117) Tartrate-resistant acid phosphatase type 5
precursor (EC 3.1.3.2) (TR-AP) (Tartrate-resistant acid
ATPase) (TrATPase)
Length = 327
Score = 103 bits (256), Expect = 8e-22
Identities = 78/255 (30%), Positives = 119/255 (46%), Gaps = 18/255 (7%)
Query: 34 LPRFEHHLKPQQQSLNFLVVGDWGRKGNY-----NQSLVAHQMGIVGDNLNIDFVISTGD 88
LP H P +L F+ VGDWG N + A ++ + DF++S GD
Sbjct: 15 LPLLTHGTAPTP-TLRFVAVGDWGGVPNAPFHTAREMANAKEIARTVQTMGADFIMSLGD 73
Query: 89 NFYKDGLEGVDDPTFYESFVNIYTAPSLQKI-WYSVLGNHDYRGDVDAQLSSILRQK--- 144
NFY G+ D F E+F ++++ +L+ I WY + GNHD+ G+V AQ++ K
Sbjct: 74 NFYFTGVHDASDKRFQETFEDVFSDRALRNIPWYVLAGNHDHLGNVSAQIAYSKISKRWN 133
Query: 145 -DSRWVCLRSFILDGGI-VEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAEL-LK 201
S + LR I I V F +DT + +PR+ A L
Sbjct: 134 FPSPYYRLRFKIPRTNITVAIFMLDTV-----MLCGNSDDFASQQPKMPRDLGVARTQLS 188
Query: 202 NVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQH 261
+ L +K + +V GH+ I S+ HG T+ L + L P+L + + AY+ GHDH LQ+
Sbjct: 189 WLKKQLAAAKEDYVLVAGHYPIWSIAEHGPTRCLVKNLRPLLATYGVTAYLCGHDHNLQY 248
Query: 262 IIDNERHGEAISSHG 276
+ D G +S G
Sbjct: 249 LQDENGVGYVLSGAG 263
>PPA_ASPFI (Q12546) Acid phosphatase precursor (EC 3.1.3.2) (pH
6-optimum acid phosphatase) (APase6)
Length = 614
Score = 34.7 bits (78), Expect = 0.33
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 1/60 (1%)
Query: 204 DLALV-KSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHI 262
DLA V +SK W IV+ H + S + + + +L +DAY++GH H + +
Sbjct: 447 DLAKVDRSKTPWVIVMSHRPMYSSAYSSYQLHVREAFEGLLLKYGVDAYLSGHIHWYERL 506
>ECE1_CAVPO (P97739) Endothelin-converting enzyme 1 (EC 3.4.24.71)
(ECE-1)
Length = 754
Score = 31.6 bits (70), Expect = 2.8
Identities = 21/110 (19%), Positives = 45/110 (40%), Gaps = 6/110 (5%)
Query: 168 TTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVG 227
T P + Y++ K G+LP Y K ++ + +VVGH +
Sbjct: 545 TPPMVNAYYSPTKNEIVFPAGILPAPFYTRSSPKALNFGGIG------VVVGHELTHAFD 598
Query: 228 HHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNERHGEAISSHGT 277
G + + L P K+++++A+ + ++ + +GE ++ T
Sbjct: 599 DQGREYDKDGNLRPWWKNSSVEAFKQQTECMVEQYSNYSVNGEPVNGRHT 648
>YP77_MYCTU (Q50644) Hypothetical protein Rv2577/MT2654
Length = 529
Score = 30.8 bits (68), Expect = 4.7
Identities = 15/50 (30%), Positives = 23/50 (46%), Gaps = 2/50 (4%)
Query: 210 SKAKWKIVVGHHTIKSVGHHGNTQEL--EQQLLPILKSNNIDAYINGHDH 257
S+ W +V H T S N +L Q+ LP+ +D + GH+H
Sbjct: 338 SEIDWVVVCMHQTAISTADDNNGADLGIRQEWLPLFDQYQVDLVVCGHEH 387
>GPMI_UREPA (Q9PQW1) 2,3-bisphosphoglycerate-independent
phosphoglycerate mutase (EC 5.4.2.1)
(Phosphoglyceromutase) (BPG-independent PGAM) (iPGM)
Length = 502
Score = 30.8 bits (68), Expect = 4.7
Identities = 36/154 (23%), Positives = 60/154 (38%), Gaps = 23/154 (14%)
Query: 153 SFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLALVKSKA 212
SF DGG+ +++ T I K TYD + P+ S +D
Sbjct: 329 SFFFDGGVNKYYSTKTQYLIPSQ----KVATYD---LCPQMSAHLITKTIIDHYFDHDV- 380
Query: 213 KWKIVVGHHTIKSVGHHGNTQELEQQLLPI----------LKSNNIDAYINGHDHCLQHI 262
+V + VGH GN Q+ + +L + + NN I G + +
Sbjct: 381 ---FIVNYANPDMVGHSGNMQQTIEAILSVDHEIRKLYDFFEKNNGVLMITGDHGNAEMM 437
Query: 263 IDNERHGEAISSHGTRK*SFTMTDKDSCLCRSQK 296
+D+++ + I+SH F +TD L +QK
Sbjct: 438 LDDQK--QVITSHSVNDVWFIITDNKVILNSTQK 469
>BXEN_CLOBU (Q06366) Botulinum neurotoxin type E, nontoxic component
Length = 1162
Score = 30.0 bits (66), Expect = 8.1
Identities = 19/65 (29%), Positives = 34/65 (52%), Gaps = 2/65 (3%)
Query: 62 YNQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWY 121
YN+ ++ ++ ++ + +N + + NFY DGL+G D FY +++ Y I Y
Sbjct: 357 YNKEII-NKPELIVNLINQNNTVLMKSNFYGDGLKGNVD-NFYSNYIIPYNLNYEHSINY 414
Query: 122 SVLGN 126
S L N
Sbjct: 415 SYLDN 419
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,583,060
Number of Sequences: 164201
Number of extensions: 1700784
Number of successful extensions: 4679
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 4661
Number of HSP's gapped (non-prelim): 12
length of query: 322
length of database: 59,974,054
effective HSP length: 110
effective length of query: 212
effective length of database: 41,911,944
effective search space: 8885332128
effective search space used: 8885332128
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146550.4


