

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.3 - phase: 0
(670 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
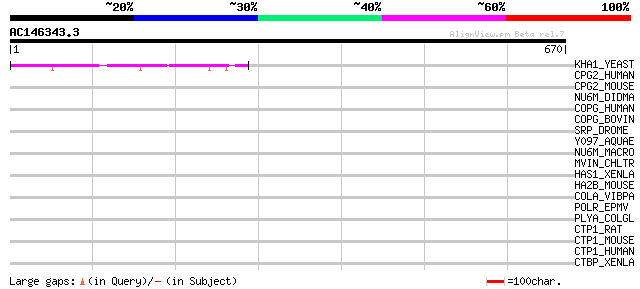
KHA1_YEAST (P40309) K(+)/H(+) antiporter 1 61 1e-08
CPG2_HUMAN (Q9UBF2) Coatomer gamma-2 subunit (Gamma-2 coat prote... 37 0.13
CPG2_MOUSE (Q9QXK3) Coatomer gamma-2 subunit (Gamma-2 coat prote... 36 0.29
NU6M_DIDMA (P41315) NADH-ubiquinone oxidoreductase chain 6 (EC 1... 33 1.9
COPG_HUMAN (Q9Y678) Coatomer gamma subunit (Gamma-coat protein) ... 33 2.4
COPG_BOVIN (P53620) Coatomer gamma subunit (Gamma-coat protein) ... 33 2.4
SRP_DROME (P52172) Box A-binding factor (ABF) (Serpent protein) ... 33 3.2
Y097_AQUAE (O66504) Hypothetical protein AQ_097 32 5.4
NU6M_MACRO (P92670) NADH-ubiquinone oxidoreductase chain 6 (EC 1... 32 7.1
MVIN_CHLTR (Q46378) Virulence factor mviN homolog 32 7.1
HAS1_XENLA (P13563) Hyaluronan synthase 1 (EC 2.4.1.212) (Hyalur... 32 7.1
HA2B_MOUSE (P14434) H-2 class II histocompatibility antigen, A-B... 32 7.1
COLA_VIBPA (Q56696) Microbial collagenase precursor (EC 3.4.24.3... 32 7.1
POLR_EPMV (P20126) RNA replicase polyprotein (EC 2.7.7.48) 31 9.2
PLYA_COLGL (Q00374) Pectin lyase precursor (EC 4.2.2.10) 31 9.2
CTP1_RAT (Q9Z2F5) C-terminal binding protein 1 (EC 1.1.1.-) (CtB... 31 9.2
CTP1_MOUSE (O88712) C-terminal binding protein 1 (EC 1.1.1.-) (C... 31 9.2
CTP1_HUMAN (Q13363) C-terminal binding protein 1 (EC 1.1.1.-) (C... 31 9.2
CTBP_XENLA (Q9YHU0) C-terminal binding protein (CtBP) 31 9.2
>KHA1_YEAST (P40309) K(+)/H(+) antiporter 1
Length = 873
Score = 60.8 bits (146), Expect = 1e-08
Identities = 57/301 (18%), Positives = 125/301 (40%), Gaps = 30/301 (9%)
Query: 1 MDFTMITRTGRKAWTIAFCSFVIPMFFG-LVVCYRFQEFWKLEMGNFEAKN---LPVIVI 56
+D I + +KA I + +P FG L+ F + G K + I +
Sbjct: 105 VDIAFIKKHLKKALVIGIVTLAVPFGFGCLLAIPLFHTYANKTEGERHIKFSVFMVFIAV 164
Query: 57 GQSGCYFAVIASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDN 116
S F V+ +L++L ++ G + L+ ++ D I+ + S+
Sbjct: 165 SISVTAFPVLCRILNELRLIKDRAGIVVLAAGIINDIMGWILLALSIILSSA-------- 216
Query: 117 GDGKGTLKAFLNVFYYLCFMVVTPLVLRPILKWFVKKTPE---GRPMKKVYMYIVFIIAL 173
+G ++ + + F++ L+ +L+W + +T E +P M I+FI+ +
Sbjct: 217 -EGSPVNTVYILLITFAWFLIYF-FPLKYLLRWVLIRTHELDRSKPSPLATMCILFIMFI 274
Query: 174 AVGMLGLLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDL 233
+ ++ + G I GL+VP ++ +++E + PI+ + +DL
Sbjct: 275 SAYFTDIIGVHPIFGAF-IAGLVVPRDDHYVVKLTERMEDIPNIVFIPIYFAVAGLNVDL 333
Query: 234 SVYVKS---DYIYVWLGIIVAVHLFKMLVT---IGICWYCNMPMADGLCLALMLSCKGVV 287
++ + Y++ +GI + + +T G+ W + +++SCKG+V
Sbjct: 334 TLLNEGRDWGYVFATIGIAIFTKIISGTLTAKLTGLFW------REATAAGVLMSCKGIV 387
Query: 288 D 288
+
Sbjct: 388 E 388
>CPG2_HUMAN (Q9UBF2) Coatomer gamma-2 subunit (Gamma-2 coat protein)
(Gamma-2 COP)
Length = 871
Score = 37.4 bits (85), Expect = 0.13
Identities = 34/129 (26%), Positives = 54/129 (41%), Gaps = 14/129 (10%)
Query: 447 FMHDDICYLALDKLASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILV-- 504
++ +D+ D + ++ F W E G +ET +TK LE A ++ +
Sbjct: 748 YVLEDLEVTVSDHIQKVLKPNFAAAWEEVGDTFEKEETFALSSTKTLEEAVNNIITFLGM 807
Query: 505 ---NRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKND 561
R P N N +S +A IF GG D L + R D + V T+++
Sbjct: 808 QPCERSDKVPENKNSHSLYLAGIFRGGYD----LLVRSRLALADGVTMQV-----TVRSK 858
Query: 562 EFTSWDVML 570
E T DV+L
Sbjct: 859 ERTPVDVIL 867
>CPG2_MOUSE (Q9QXK3) Coatomer gamma-2 subunit (Gamma-2 coat protein)
(Gamma-2 COP)
Length = 871
Score = 36.2 bits (82), Expect = 0.29
Identities = 33/129 (25%), Positives = 54/129 (41%), Gaps = 14/129 (10%)
Query: 447 FMHDDICYLALDKLASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILV-- 504
++ +D+ D + I+ F W E G +ET +TK LE A ++ +
Sbjct: 748 YVLEDLEVTVSDHIQKILKPNFAAAWEEVGDAFEKEETFALSSTKTLEEAVNNIITFLGM 807
Query: 505 ---NRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKND 561
R P N N +S +A ++ GG D L + R D + V T+++
Sbjct: 808 QPCERSDKVPENKNSHSLYLAGVYRGGYD----LLVRSRLALADGVTMQV-----TVRSK 858
Query: 562 EFTSWDVML 570
E T DV+L
Sbjct: 859 ERTPVDVIL 867
>NU6M_DIDMA (P41315) NADH-ubiquinone oxidoreductase chain 6 (EC
1.6.5.3)
Length = 168
Score = 33.5 bits (75), Expect = 1.9
Identities = 27/110 (24%), Positives = 56/110 (50%), Gaps = 9/110 (8%)
Query: 160 MKKVYMYIVFIIALAVGMLGLLTKQS-VLGGICIVGLIVPEGPPLGTEMIKQLE-LFCSW 217
MK + +YI+ ++ L +G + +K S + GG+ L+V G LG M+ LE +F
Sbjct: 1 MKMMTIYIISLL-LMIGFVAFASKPSPIYGGL---SLVVSGG--LGCGMVVSLEDVFLGL 54
Query: 218 FLFPIFVTSCAMKIDLSVYVKSD-YIYVWLGIIVAVHLFKMLVTIGICWY 266
+F +++ + + + ++ Y W+G +VA + ++ + + WY
Sbjct: 55 VVFLVYLGGMLVVFGYTTAMATEEYPETWVGNVVAFIMLLFVLLLQVGWY 104
>COPG_HUMAN (Q9Y678) Coatomer gamma subunit (Gamma-coat protein)
(Gamma-COP)
Length = 874
Score = 33.1 bits (74), Expect = 2.4
Identities = 22/89 (24%), Positives = 37/89 (40%), Gaps = 5/89 (5%)
Query: 447 FMHDDICYLALDKLASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILV-- 504
++ +D+ D + ++ L F W E G +ET K LE A ++ +
Sbjct: 751 YVLEDLEVTVADHIQKVMKLNFEAAWDEVGDEFEKEETFTLSTIKTLEEAVGNIVKFLGM 810
Query: 505 ---NRGHSSPFNHNENSKQIAMIFLGGSD 530
R P N N ++ +A +F GG D
Sbjct: 811 HPCERSDKVPDNKNTHTLLLAGVFRGGHD 839
>COPG_BOVIN (P53620) Coatomer gamma subunit (Gamma-coat protein)
(Gamma-COP)
Length = 874
Score = 33.1 bits (74), Expect = 2.4
Identities = 30/129 (23%), Positives = 53/129 (40%), Gaps = 14/129 (10%)
Query: 447 FMHDDICYLALDKLASIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILV-- 504
++ +D+ D + ++ L F W E G +ET K LE A ++ +
Sbjct: 751 YVLEDLEVTIADHIQKVMKLNFEAAWDEVGDEFQKEETFTLSTIKTLEEAVGNIVKFLGM 810
Query: 505 ---NRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAKRTIKEDTYHLVVYHLVSTIKND 561
R P N N ++ +A +F GG D + + R + DT + V T ++
Sbjct: 811 HPCERSDKVPDNKNTHTLLLAGVFRGGHD----ILVRSRLLLLDTVTMQV-----TARSS 861
Query: 562 EFTSWDVML 570
E D++L
Sbjct: 862 EELPVDIVL 870
>SRP_DROME (P52172) Box A-binding factor (ABF) (Serpent protein)
(GATA-binding factor-B) (Transcription factor GATA-B)
(dGATA-B)
Length = 779
Score = 32.7 bits (73), Expect = 3.2
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 6/65 (9%)
Query: 67 ASLLSDLEILNSELGRLALSTAMVMDSFNSIVTGIGTAFISSIKTDSHDNGDGKGTLKAF 126
AS + LE+L + +A +TA V +S+ G G+ SH G G GTL AF
Sbjct: 89 ASGSAPLELLGATAAAVAAATAAVNGHNSSLEDGYGSP------RSSHSGGGGGGTLPAF 142
Query: 127 LNVFY 131
+ Y
Sbjct: 143 QRIAY 147
>Y097_AQUAE (O66504) Hypothetical protein AQ_097
Length = 407
Score = 32.0 bits (71), Expect = 5.4
Identities = 34/144 (23%), Positives = 59/144 (40%), Gaps = 8/144 (5%)
Query: 126 FLNVFYYLCFMVVTPLVLRPILKWFVKKTP------EGRPMKKVYMYIVFIIALAVGMLG 179
F + YL F + +P + R L W V TP G + +Y V +GM G
Sbjct: 15 FTSFILYLLFFL-SPFLERKRL-WTVAVTPLASIIGSGYLVSAPLLYYVLGKYAILGMAG 72
Query: 180 LLTKQSVLGGICIVGLIVPEGPPLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKS 239
++ ++GG +I E G+E K L F + A I ++ Y++
Sbjct: 73 IVILAYLIGGAIRYNIIKAEPILYGSESSKLKALLLDVERFSNLSLAFAYMISIAFYLRL 132
Query: 240 DYIYVWLGIIVAVHLFKMLVTIGI 263
+V+ G +++ L+T G+
Sbjct: 133 LSSFVFSGFFERNEVYERLLTTGL 156
>NU6M_MACRO (P92670) NADH-ubiquinone oxidoreductase chain 6 (EC
1.6.5.3)
Length = 167
Score = 31.6 bits (70), Expect = 7.1
Identities = 28/108 (25%), Positives = 50/108 (45%), Gaps = 8/108 (7%)
Query: 162 KVYMYIVFIIALAVGMLGLLTKQS-VLGGICIVGLIVPEGPPLGTEMIKQLE-LFCSWFL 219
K+ + +F I L G + +K S V GG+ L+V G LG ++ LE +F +
Sbjct: 2 KMMVVFLFSILLVFGFVAFASKPSPVYGGL---SLVVSGG--LGCAIVVSLEDVFLGLIV 56
Query: 220 FPIFVTSCAMKIDLSVYVKSD-YIYVWLGIIVAVHLFKMLVTIGICWY 266
F I++ + + + ++ Y W+G VA+ + V + WY
Sbjct: 57 FLIYLGGMLVVFGYTAAMATEEYPESWVGNTVALSMLLFTVIVESAWY 104
>MVIN_CHLTR (Q46378) Virulence factor mviN homolog
Length = 536
Score = 31.6 bits (70), Expect = 7.1
Identities = 24/89 (26%), Positives = 42/89 (46%), Gaps = 9/89 (10%)
Query: 254 LFKMLVTIGICWYCNM---PMADGLCLALMLSCKGV---VDFCNNVFLHDSMLLSSEALS 307
LF +++ +G+C +C+ + D L L ++L G+ + N+ LH S L+
Sbjct: 102 LFTLIIELGLCVWCSCVTGSLFDTLLLTIILLPSGIFLMMYTVNSTLLHCEKKFFSVGLA 161
Query: 308 VLSINVLVIGTLARIGVKYLYDPSRKYAG 336
+NV IGT + + YDP + G
Sbjct: 162 PSVVNVSWIGT---VFLARNYDPRNRIFG 187
>HAS1_XENLA (P13563) Hyaluronan synthase 1 (EC 2.4.1.212)
(Hyaluronate synthase 1) (Hyaluronic acid synthase 1)
(HA synthase 1) (XHAS1) (DG42 protein)
Length = 588
Score = 31.6 bits (70), Expect = 7.1
Identities = 26/113 (23%), Positives = 53/113 (46%), Gaps = 19/113 (16%)
Query: 218 FLFPIFVTSCAMKIDLSVYVKSDYIYVWLGIIVAVHLFKMLVTIGICWYCNMPMADGLCL 277
F+FP F+T+ +++ +Y + + VWL ++ + + + +I CW G +
Sbjct: 412 FIFPFFITATVIRL---IYAGTIWNVVWL--LLCIQIMSLFKSIYACW------LRGNFI 460
Query: 278 ALMLSCKGVVDFCNNVFLHDSMLLSSEALSVLSINVLVIGTLARIGVKYLYDP 330
L++S + L+ + LL S+ ++L++N GT R + Y P
Sbjct: 461 MLLMSLYSM--------LYMTGLLPSKYFALLTLNKTGWGTSGRKKIVGNYMP 505
>HA2B_MOUSE (P14434) H-2 class II histocompatibility antigen, A-B
alpha chain precursor (IAalpha)
Length = 256
Score = 31.6 bits (70), Expect = 7.1
Identities = 25/93 (26%), Positives = 43/93 (45%), Gaps = 14/93 (15%)
Query: 424 DLFEHDNAGTASVSTYTAISPV-----RFMHDDICYLALDKLASIIILPFHLRWSEDGSV 478
D E D+ GT +S Y + + F D++ Y+ LDK ++ +LP E G +
Sbjct: 25 DDIEADHVGTYGISVYQSPGDIGQYTFEFDGDELFYVDLDKKETVWMLP------EFGQL 78
Query: 479 ESADETTRSLNTKVLERAPCSVAILVNRGHSSP 511
S D N V++ ++ +L R +S+P
Sbjct: 79 ASFDPQGGLQNIAVVKH---NLGVLTKRSNSTP 108
>COLA_VIBPA (Q56696) Microbial collagenase precursor (EC 3.4.24.3)
(prtVp)
Length = 816
Score = 31.6 bits (70), Expect = 7.1
Identities = 17/37 (45%), Positives = 21/37 (55%)
Query: 632 ASWTEYPELGVLGDLLASPDTNTKASILVVQQQVMPK 668
A++ + L G LLASPD TK L V QQVM +
Sbjct: 240 ANFIVFNALRETGRLLASPDQETKRKALAVMQQVMQR 276
>POLR_EPMV (P20126) RNA replicase polyprotein (EC 2.7.7.48)
Length = 1839
Score = 31.2 bits (69), Expect = 9.2
Identities = 26/115 (22%), Positives = 54/115 (46%), Gaps = 10/115 (8%)
Query: 276 CLALMLSCKGVVDFCNNVFLHDSML----LSSEALSVLSINVLVIGTLARIGVKYLYDPS 331
CL + + V F N+ L + + S+AL +I++ + + +Y+P
Sbjct: 1178 CLVALTRSRTGVYFVGNLHLASNSFGTNYMFSQALCQGTIDLNNVFPHIMPHLPKMYEPI 1237
Query: 332 RKYAGYQKRNILSLKPNSELKIVSCILKPSHI---IPIKNVLDICSPTSSNPLVI 383
R + L+ +P + +++S + KP+H+ IP + LD+ SNP+++
Sbjct: 1238 RSRSNRFVSGSLNFRPTTNSRLLSSLTKPTHLPPHIPTNHSLDV---LVSNPVLL 1289
>PLYA_COLGL (Q00374) Pectin lyase precursor (EC 4.2.2.10)
Length = 380
Score = 31.2 bits (69), Expect = 9.2
Identities = 28/113 (24%), Positives = 50/113 (43%), Gaps = 14/113 (12%)
Query: 557 TIKNDEF---TSWDVMLDDELLKGVKGVYGSVDNVTYEKVEVENTSDTTEFISDIAIQH- 612
TI N+E TSW D+ GV + GS D VT+ K + +TS + I+ ++ H
Sbjct: 211 TISNNEIDGSTSWSATCDNHHYWGVY-LTGSNDMVTFSKNYIHHTSGRSPKIAGNSLVHI 269
Query: 613 --DFIIVGRRNGIKSP-------QTQALASWTEYPELGVLGDLLASPDTNTKA 656
++ + +++ + + + G+ G + +SPD NT A
Sbjct: 270 SGNYFYANSGHAMEADAGAKVVLEGNVFQNVVAAMQSGLAGKVFSSPDANTNA 322
>CTP1_RAT (Q9Z2F5) C-terminal binding protein 1 (EC 1.1.1.-) (CtBP1)
(C-terminal binding protein 3) (CtBP3) (50 kDa
BFA-dependent ADP-ribosylation substrate) (BARS-50)
Length = 430
Score = 31.2 bits (69), Expect = 9.2
Identities = 15/53 (28%), Positives = 25/53 (46%)
Query: 462 SIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH 514
++I P +SE S+E +E R + + R P S+ VN+ H + H
Sbjct: 298 NLICTPHAAWYSEQASIEMREEAAREIRRAITGRIPDSLKNCVNKDHLTAATH 350
>CTP1_MOUSE (O88712) C-terminal binding protein 1 (EC 1.1.1.-)
(CtBP1)
Length = 440
Score = 31.2 bits (69), Expect = 9.2
Identities = 15/53 (28%), Positives = 25/53 (46%)
Query: 462 SIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH 514
++I P +SE S+E +E R + + R P S+ VN+ H + H
Sbjct: 309 NLICTPHAAWYSEQASIEMREEAAREIRRAITGRIPDSLKNCVNKDHLTAATH 361
>CTP1_HUMAN (Q13363) C-terminal binding protein 1 (EC 1.1.1.-)
(CtBP1)
Length = 440
Score = 31.2 bits (69), Expect = 9.2
Identities = 15/53 (28%), Positives = 25/53 (46%)
Query: 462 SIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH 514
++I P +SE S+E +E R + + R P S+ VN+ H + H
Sbjct: 309 NLICTPHAAWYSEQASIEMREEAAREIRRAITGRIPDSLKNCVNKDHLTAATH 361
>CTBP_XENLA (Q9YHU0) C-terminal binding protein (CtBP)
Length = 440
Score = 31.2 bits (69), Expect = 9.2
Identities = 15/53 (28%), Positives = 25/53 (46%)
Query: 462 SIIILPFHLRWSEDGSVESADETTRSLNTKVLERAPCSVAILVNRGHSSPFNH 514
++I P +SE S+E +E R + + R P S+ VN+ H + H
Sbjct: 309 NLICTPHAAWYSEQASIEMREEAAREIRRAITGRIPDSLKNCVNKDHLTAATH 361
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 75,830,564
Number of Sequences: 164201
Number of extensions: 3175310
Number of successful extensions: 9678
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 9667
Number of HSP's gapped (non-prelim): 22
length of query: 670
length of database: 59,974,054
effective HSP length: 117
effective length of query: 553
effective length of database: 40,762,537
effective search space: 22541682961
effective search space used: 22541682961
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146343.3


