

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.8 + phase: 0
(268 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
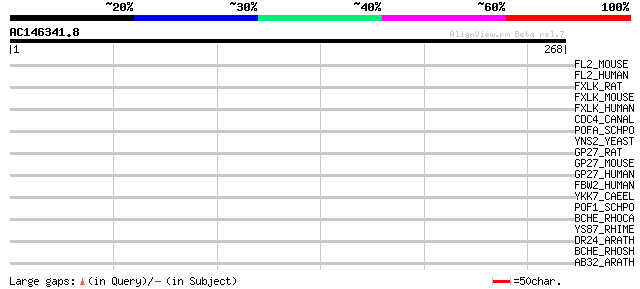
Score E
Sequences producing significant alignments: (bits) Value
FL2_MOUSE (Q8BH16) F-box/LRR-repeat protein 2 (F-box and leucine... 37 0.066
FL2_HUMAN (Q9UKC9) F-box/LRR-repeat protein 2 (F-box and leucine... 37 0.066
FXLK_RAT (Q9QZH7) F-box/LRR-repeat protein 20 (F-box and leucine... 36 0.11
FXLK_MOUSE (Q9CZV8) F-box/LRR-repeat protein 20 (F-box and leuci... 36 0.11
FXLK_HUMAN (Q96IG2) F-box/LRR-repeat protein 20 (F-box and leuci... 36 0.11
CDC4_CANAL (P53699) Cell division control protein 4 34 0.33
POFA_SCHPO (Q9P7W4) F-box/WD-repeat protein pof10 (Skp1-binding ... 33 0.73
YNS2_YEAST (P53877) Hypothetical 61.8 kDa Trp-Asp repeats contai... 31 2.8
GP27_RAT (Q9JJH3) Probable G protein-coupled receptor 27 (Super ... 31 2.8
GP27_MOUSE (O54897) Probable G protein-coupled receptor 27 (Supe... 31 2.8
GP27_HUMAN (Q9NS67) Probable G protein-coupled receptor 27 (Supe... 31 2.8
FBW2_HUMAN (Q9UKT8) F-box/WD-repeat protein 2 (F-box and WD-40 d... 31 3.6
YKK7_CAEEL (P34284) Hypothetical F-box/LRR-repeat protein C02F5.... 30 4.7
POF1_SCHPO (P87053) F-box/WD-repeat protein pof1 (Skp1-binding p... 30 6.2
BCHE_RHOCA (P26168) Magnesium-protoporphyrin IX monomethyl ester... 30 6.2
YS87_RHIME (Q92LX6) Hypothetical UPF0176 protein R02887 30 8.0
DR24_ARATH (O23317) Probable disease resistance protein At4g1461... 30 8.0
BCHE_RHOSH (Q9RFD3) Magnesium-protoporphyrin IX monomethyl ester... 30 8.0
AB32_ARATH (Q8L718) Inner membrane ALBINO3-like protein 2, chlor... 30 8.0
>FL2_MOUSE (Q8BH16) F-box/LRR-repeat protein 2 (F-box and
leucine-rich repeat protein 2)
Length = 423
Score = 36.6 bits (83), Expect = 0.066
Identities = 16/34 (47%), Positives = 23/34 (67%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
LP+EL+ +I S LD+ T+ R +SK+WN L D
Sbjct: 15 LPKELLLRIFSFLDIVTLCRCAQISKAWNILALD 48
>FL2_HUMAN (Q9UKC9) F-box/LRR-repeat protein 2 (F-box and
leucine-rich repeat protein 2) (F-box protein
FBL2/FBL3)
Length = 423
Score = 36.6 bits (83), Expect = 0.066
Identities = 16/34 (47%), Positives = 23/34 (67%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
LP+EL+ +I S LD+ T+ R +SK+WN L D
Sbjct: 15 LPKELLLRIFSFLDIVTLCRCAQISKAWNILALD 48
>FXLK_RAT (Q9QZH7) F-box/LRR-repeat protein 20 (F-box and
leucine-rich repeat protein 20) (F-box/LRR-repeat
protein 2-like)
Length = 276
Score = 35.8 bits (81), Expect = 0.11
Identities = 16/34 (47%), Positives = 23/34 (67%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
LP+EL+ +I S LDV T+ R +S++WN L D
Sbjct: 28 LPKELLLRIFSFLDVVTLCRCAQVSRAWNVLALD 61
>FXLK_MOUSE (Q9CZV8) F-box/LRR-repeat protein 20 (F-box and
leucine-rich repeat protein 20) (F-box/LRR-repeat
protein 2-like)
Length = 436
Score = 35.8 bits (81), Expect = 0.11
Identities = 16/34 (47%), Positives = 23/34 (67%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
LP+EL+ +I S LDV T+ R +S++WN L D
Sbjct: 28 LPKELLLRIFSFLDVVTLCRCAQVSRAWNVLALD 61
>FXLK_HUMAN (Q96IG2) F-box/LRR-repeat protein 20 (F-box and
leucine-rich repeat protein 20) (F-box/LRR-repeat
protein 2-like)
Length = 436
Score = 35.8 bits (81), Expect = 0.11
Identities = 16/34 (47%), Positives = 23/34 (67%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
LP+EL+ +I S LDV T+ R +S++WN L D
Sbjct: 28 LPKELLLRIFSFLDVVTLCRCAQVSRAWNVLALD 61
>CDC4_CANAL (P53699) Cell division control protein 4
Length = 684
Score = 34.3 bits (77), Expect = 0.33
Identities = 18/44 (40%), Positives = 27/44 (60%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKR 70
+P E+ +ILS LD KT+L + + K W +I++P K LKR
Sbjct: 218 VPFEVTMKILSYLDYKTLLSVAQVCKKWFDIINNPDTWIKLLKR 261
>POFA_SCHPO (Q9P7W4) F-box/WD-repeat protein pof10 (Skp1-binding
protein 2)
Length = 662
Score = 33.1 bits (74), Expect = 0.73
Identities = 15/54 (27%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Query: 8 SIKKKKKDKKNKKLTLSMF-LPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
++++ +D + ++ LP+E++ I S LD +++L C K W L+SD
Sbjct: 14 NLRRMNRDHSSNNTNRTVLNLPKEILIIIFSFLDPRSLLSAQCTCKYWKKLLSD 67
>YNS2_YEAST (P53877) Hypothetical 61.8 kDa Trp-Asp repeats
containing protein in NPR1-RPS3 intergenic region
Length = 555
Score = 31.2 bits (69), Expect = 2.8
Identities = 27/103 (26%), Positives = 43/103 (41%), Gaps = 14/103 (13%)
Query: 85 MRTIQSLP--------ITRLLKNPSITVSGDFCYDGGGL-----NDNCKVVISCNGLLCF 131
+RTIQ+L +T LL NP G+ ++G N+N + L
Sbjct: 379 IRTIQTLTTSQDSVGEVTNLLTNPYRLERGNLLFEGESKGKQPSNNNGHNFMKIPNLQRV 438
Query: 132 IFCSDNKEY-HNYWFRIWNPATGTRSKALGTNHDYNLQLRSLR 173
IF NK + H+ W++I P T +D+N L ++
Sbjct: 439 IFDGKNKGHLHDIWYQIGEPEAETDPNLALPLNDFNAYLEQVK 481
>GP27_RAT (Q9JJH3) Probable G protein-coupled receptor 27 (Super
conserved receptor expressed in brain 1)
Length = 377
Score = 31.2 bits (69), Expect = 2.8
Identities = 24/93 (25%), Positives = 39/93 (41%), Gaps = 5/93 (5%)
Query: 39 LDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMRTIQSLPITRLLK 98
L + T+ L+C+S + N L + ++ L R P +L C +R + LP L
Sbjct: 21 LRLATLSLLLCVSLAGNVLFALLIVRERSLHRAPYYLLLDLCLADGLRALACLPAVMLAA 80
Query: 99 NPSITVSGDFCYDGGGLNDNCKVVISCNGLLCF 131
+ +G G L CK++ L CF
Sbjct: 81 RRAAAAAGT---PPGAL--GCKLLAFLAALFCF 108
>GP27_MOUSE (O54897) Probable G protein-coupled receptor 27 (Super
conserved receptor expressed in brain 1)
Length = 379
Score = 31.2 bits (69), Expect = 2.8
Identities = 24/93 (25%), Positives = 39/93 (41%), Gaps = 5/93 (5%)
Query: 39 LDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMRTIQSLPITRLLK 98
L + T+ L+C+S + N L + ++ L R P +L C +R + LP L
Sbjct: 23 LRLATLSLLLCVSLAGNVLFALLIVRERSLHRAPYYLLLDLCLADGLRALACLPAVMLAA 82
Query: 99 NPSITVSGDFCYDGGGLNDNCKVVISCNGLLCF 131
+ +G G L CK++ L CF
Sbjct: 83 RRAAAAAGT---PPGAL--GCKLLAFLAALFCF 110
>GP27_HUMAN (Q9NS67) Probable G protein-coupled receptor 27 (Super
conserved receptor expressed in brain 1)
Length = 375
Score = 31.2 bits (69), Expect = 2.8
Identities = 24/93 (25%), Positives = 39/93 (41%), Gaps = 5/93 (5%)
Query: 39 LDVKTILRLICLSKSWNTLISDPTFVQKQLKRPPRLILKPPCWVYPMRTIQSLPITRLLK 98
L + T+ L+C+S + N L + ++ L R P +L C +R + LP L
Sbjct: 20 LKLATLSLLLCVSLAGNVLFALLIVRERSLHRAPYYLLLDLCLADGLRALACLPAVMLAA 79
Query: 99 NPSITVSGDFCYDGGGLNDNCKVVISCNGLLCF 131
+ +G G L CK++ L CF
Sbjct: 80 RRAAAAAG---APPGAL--GCKLLAFLAALFCF 107
>FBW2_HUMAN (Q9UKT8) F-box/WD-repeat protein 2 (F-box and WD-40
domain protein 2)
Length = 422
Score = 30.8 bits (68), Expect = 3.6
Identities = 17/38 (44%), Positives = 22/38 (57%)
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFV 64
LP EL +L LD +T+L +SK WN +IS T V
Sbjct: 60 LPLELSFYLLKWLDPQTLLTCCLVSKQWNKVISACTEV 97
>YKK7_CAEEL (P34284) Hypothetical F-box/LRR-repeat protein C02F5.7
in chromosome III
Length = 461
Score = 30.4 bits (67), Expect = 4.7
Identities = 12/38 (31%), Positives = 23/38 (59%)
Query: 23 LSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
++ LP+E++ ++ S LD K + R + +SW+ L D
Sbjct: 56 INRVLPKEVLLKVFSFLDTKALCRSAQVCRSWSILALD 93
>POF1_SCHPO (P87053) F-box/WD-repeat protein pof1 (Skp1-binding
protein 1)
Length = 605
Score = 30.0 bits (66), Expect = 6.2
Identities = 19/57 (33%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Query: 4 SKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISD 60
S L+S D + LS+ LP E+ +ILS LD +++ + +SK W L D
Sbjct: 91 SSLLSFASSTLDSLVRLDFLSL-LPVEISFRILSFLDARSLCQAAQVSKHWKELADD 146
>BCHE_RHOCA (P26168) Magnesium-protoporphyrin IX monomethyl ester
[oxidative] cyclase (EC 1.14.13.81) (Mg-protoporphyrin
IX monomethyl ester oxidative cyclase)
Length = 575
Score = 30.0 bits (66), Expect = 6.2
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 7/67 (10%)
Query: 141 HNYWFRIWNPATGTRSKAL-GTNHDYN-LQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLK 198
+N+ I P TR + L G +Y +R F++ G +RRRYLLGCLK
Sbjct: 401 YNFVTPIMKPKALTRGELLDGVMKNYRRFYMRKALFHYPWRGTG-----FRRRYLLGCLK 455
Query: 199 FTFGCGI 205
G+
Sbjct: 456 AFLKAGV 462
>YS87_RHIME (Q92LX6) Hypothetical UPF0176 protein R02887
Length = 315
Score = 29.6 bits (65), Expect = 8.0
Identities = 15/53 (28%), Positives = 26/53 (48%)
Query: 9 IKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDP 61
I K+ + L + + L +E++ + +D K ++ K WN LISDP
Sbjct: 86 IHKESRASAMPFLRMKVRLKKEIVTMGVENIDPKKVVGTYVEPKDWNALISDP 138
>DR24_ARATH (O23317) Probable disease resistance protein At4g14610
(pCol1)
Length = 719
Score = 29.6 bits (65), Expect = 8.0
Identities = 15/59 (25%), Positives = 31/59 (52%), Gaps = 1/59 (1%)
Query: 3 YSKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTIL-RLICLSKSWNTLISD 60
Y K++S+ K+ + + + + E L+AQ+ + T++ + L + WNTL+ D
Sbjct: 89 YGKMVSVMLKEVENLSSRGVFDVVTEENLVAQVEEMPIQSTVVGQETMLERVWNTLMKD 147
>BCHE_RHOSH (Q9RFD3) Magnesium-protoporphyrin IX monomethyl ester
[oxidative] cyclase (EC 1.14.13.81) (Mg-protoporphyrin
IX monomethyl ester oxidative cyclase)
Length = 611
Score = 29.6 bits (65), Expect = 8.0
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 7/67 (10%)
Query: 141 HNYWFRIWNPATGTRSKAL-GTNHDYN-LQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLK 198
+N+ I P TR + L G ++Y ++ F++ G +RRRYLLGCLK
Sbjct: 401 YNFVTPIMKPKALTRGELLDGVMNNYRRFYMKKALFHYPWRGTG-----FRRRYLLGCLK 455
Query: 199 FTFGCGI 205
G+
Sbjct: 456 AFLKAGV 462
>AB32_ARATH (Q8L718) Inner membrane ALBINO3-like protein 2,
chloroplast precursor (Ath5)
Length = 525
Score = 29.6 bits (65), Expect = 8.0
Identities = 21/60 (35%), Positives = 33/60 (55%), Gaps = 8/60 (13%)
Query: 9 IKKKKKDKKNKKLTLSMFLPEELIA---QILSLLDVKTILRLICLSKSWNTLISDPTFVQ 65
IK D K + L+L PEEL++ Q+LS D +T ++L+ L+ L DP +V+
Sbjct: 327 IKYVDSDSKEQTLSLQTLTPEELLSLSVQVLSKGDKETSIQLLRLA-----LEKDPGYVR 381
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,902,392
Number of Sequences: 164201
Number of extensions: 1443540
Number of successful extensions: 2816
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2805
Number of HSP's gapped (non-prelim): 19
length of query: 268
length of database: 59,974,054
effective HSP length: 108
effective length of query: 160
effective length of database: 42,240,346
effective search space: 6758455360
effective search space used: 6758455360
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146341.8


