

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146333.2 - phase: 0
(615 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
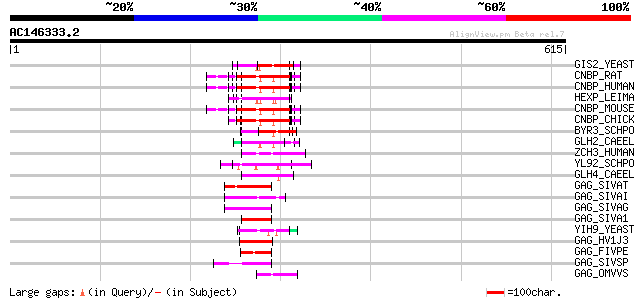
Score E
Sequences producing significant alignments: (bits) Value
GIS2_YEAST (P53849) Zinc-finger protein GIS2 65 4e-10
CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP) (... 62 6e-09
CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)... 62 6e-09
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 61 8e-09
CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)... 60 2e-08
CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP) 60 2e-08
BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog 58 8e-08
GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-... 55 5e-07
ZCH3_HUMAN (Q9NUD5) Zinc finger CCHC domain containing protein 3 54 2e-06
YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I 54 2e-06
GLH4_CAEEL (O76743) ATP-dependent RNA helicase glh-4 (EC 3.6.1.-... 53 3e-06
GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17; ... 52 4e-06
GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17; ... 52 4e-06
GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17; ... 50 2e-05
GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17; ... 50 2e-05
YIH9_YEAST (P40507) Hypothetical 41.6 kDa protein in SDS3-THS1 i... 50 2e-05
GAG_HV1J3 (P12494) Gag polyprotein [Contains: Core protein p17 (... 48 7e-05
GAG_FIVPE (P16087) Gag polyprotein [Contains: Core protein p15; ... 48 7e-05
GAG_SIVSP (P19504) Gag polyprotein [Contains: Core protein p17; ... 48 9e-05
GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16; ... 48 9e-05
>GIS2_YEAST (P53849) Zinc-finger protein GIS2
Length = 153
Score = 65.5 bits (158), Expect = 4e-10
Identities = 30/74 (40%), Positives = 42/74 (56%), Gaps = 6/74 (8%)
Query: 253 AEIVCFNCGEKGHKSNACPE----EIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISS 308
+E +C+NC + GH C E K+C CG+ GHV ++C T CFNCN GHIS
Sbjct: 21 SERLCYNCNKPGHVQTDCTMPRTVEFKQCYNCGETGHVRSEC--TVQRCFNCNQTGHISR 78
Query: 309 QCTQPKRAPTTGRV 322
+C +PK+ +V
Sbjct: 79 ECPEPKKTSRFSKV 92
Score = 57.4 bits (137), Expect = 1e-07
Identities = 27/69 (39%), Positives = 38/69 (54%), Gaps = 6/69 (8%)
Query: 247 PKKKDA-AEIVCFNCGEKGHKSNACPEEIK----KCVRCGKKGHVVADCNRTDIVCFNCN 301
PKK +++ C+ CG H + C +E KC CG+ GH+ DC + D +C+NCN
Sbjct: 83 PKKTSRFSKVSCYKCGGPNHMAKDCMKEDGISGLKCYTCGQAGHMSRDC-QNDRLCYNCN 141
Query: 302 GEGHISSQC 310
GHIS C
Sbjct: 142 ETGHISKDC 150
Score = 49.3 bits (116), Expect = 3e-05
Identities = 18/40 (45%), Positives = 27/40 (67%), Gaps = 1/40 (2%)
Query: 275 KKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPK 314
K C CGK GH+ DC+ ++ +C+NCN GH+ + CT P+
Sbjct: 4 KACYVCGKIGHLAEDCD-SERLCYNCNKPGHVQTDCTMPR 42
>CNBP_RAT (P62634) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 61.6 bits (148), Expect = 6e-09
Identities = 32/82 (39%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 248 KKKDAAEIVCFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNG 302
K D E C+NCG GH + C E ++ C CGK GH+ DC+ D C++C
Sbjct: 65 KDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGE 124
Query: 303 EGHISSQCTQPK--RAPTTGRV 322
GHI CT+ K R TG V
Sbjct: 125 FGHIQKDCTKVKCYRCGETGHV 146
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 252 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 310
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 114 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
Query: 311 T 311
T
Sbjct: 172 T 172
Score = 58.5 bits (140), Expect = 5e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 243 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 301
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 86 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 141
Query: 302 GEGHISSQCTQ 312
GH++ C++
Sbjct: 142 ETGHVAINCSK 152
Score = 56.6 bits (135), Expect = 2e-07
Identities = 26/87 (29%), Positives = 35/87 (39%), Gaps = 28/87 (32%)
Query: 257 CFNCGEKGHKSNACPEEIKK----------------------------CVRCGKKGHVVA 288
CF CG GH + CP + C RCG+ GH+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 289 DCNRTDIVCFNCNGEGHISSQCTQPKR 315
DC+ + C+NC GHI+ C +PKR
Sbjct: 66 DCDLQEDACYNCGRGGHIAKDCKEPKR 92
Score = 56.2 bits (134), Expect = 2e-07
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 13/96 (13%)
Query: 219 KGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCV 278
+G+G +SR + +D+G Q + + +C+ CGE GH + C + C
Sbjct: 25 RGRGMRSRGRG-GFTSDRGFQFV--------SSSLPDICYRCGESGHLAKDCDLQEDACY 75
Query: 279 RCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 310
CG+ GH+ DC + C+NC GH++ C
Sbjct: 76 NCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 43.9 bits (102), Expect = 0.001
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 129 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
>CNBP_HUMAN (P62633) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 177
Score = 61.6 bits (148), Expect = 6e-09
Identities = 32/82 (39%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 248 KKKDAAEIVCFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNG 302
K D E C+NCG GH + C E ++ C CGK GH+ DC+ D C++C
Sbjct: 65 KDCDLQEDACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGE 124
Query: 303 EGHISSQCTQPK--RAPTTGRV 322
GHI CT+ K R TG V
Sbjct: 125 FGHIQKDCTKVKCYRCGETGHV 146
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 252 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 310
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 114 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
Query: 311 T 311
T
Sbjct: 172 T 172
Score = 58.5 bits (140), Expect = 5e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 243 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 301
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 86 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 141
Query: 302 GEGHISSQCTQ 312
GH++ C++
Sbjct: 142 ETGHVAINCSK 152
Score = 56.6 bits (135), Expect = 2e-07
Identities = 26/87 (29%), Positives = 35/87 (39%), Gaps = 28/87 (32%)
Query: 257 CFNCGEKGHKSNACPEEIKK----------------------------CVRCGKKGHVVA 288
CF CG GH + CP + C RCG+ GH+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 289 DCNRTDIVCFNCNGEGHISSQCTQPKR 315
DC+ + C+NC GHI+ C +PKR
Sbjct: 66 DCDLQEDACYNCGRGGHIAKDCKEPKR 92
Score = 56.2 bits (134), Expect = 2e-07
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 13/96 (13%)
Query: 219 KGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKKCV 278
+G+G +SR + +D+G Q + + +C+ CGE GH + C + C
Sbjct: 25 RGRGMRSRGRG-GFTSDRGFQFV--------SSSLPDICYRCGESGHLAKDCDLQEDACY 75
Query: 279 RCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 310
CG+ GH+ DC + C+NC GH++ C
Sbjct: 76 NCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 111
Score = 43.9 bits (102), Expect = 0.001
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 129 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 171
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 61.2 bits (147), Expect = 8e-09
Identities = 28/76 (36%), Positives = 35/76 (45%), Gaps = 14/76 (18%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIKK-------CVRCGKKGHVVADCNRT-------D 294
K D CF CGE+GH S CP E + C RCG+ GH+ DC +
Sbjct: 37 KGDERSTTCFRCGEEGHMSRECPNEARSGAAGAMTCFRCGEAGHMSRDCPNSAKPGAAKG 96
Query: 295 IVCFNCNGEGHISSQC 310
C+ C EGH+S C
Sbjct: 97 FECYKCGQEGHLSRDC 112
Score = 57.8 bits (138), Expect = 8e-08
Identities = 28/82 (34%), Positives = 41/82 (49%), Gaps = 16/82 (19%)
Query: 243 DVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKK-------CVRCGKKGHVVADCNRT-- 293
DV+RP+ + + C NCG++GH + CPE K C RCG++GH+ +C
Sbjct: 6 DVKRPRTESSTS--CRNCGKEGHYARECPEADSKGDERSTTCFRCGEEGHMSRECPNEAR 63
Query: 294 -----DIVCFNCNGEGHISSQC 310
+ CF C GH+S C
Sbjct: 64 SGAAGAMTCFRCGEAGHMSRDC 85
Score = 53.1 bits (126), Expect = 2e-06
Identities = 24/75 (32%), Positives = 35/75 (46%), Gaps = 14/75 (18%)
Query: 252 AAEIVCFNCGEKGHKSNACPE--------EIKKCVRCGKKGHVVADC------NRTDIVC 297
A + C+ CG+ GH S CP +KC +CG+ GH+ +C D C
Sbjct: 165 AGDRTCYKCGDAGHISRDCPNGQGGYSGAGDRKCYKCGESGHMSRECPSAGSTGSGDRAC 224
Query: 298 FNCNGEGHISSQCTQ 312
+ C GHIS +C +
Sbjct: 225 YKCGKPGHISRECPE 239
Score = 53.1 bits (126), Expect = 2e-06
Identities = 25/76 (32%), Positives = 32/76 (41%), Gaps = 17/76 (22%)
Query: 252 AAEIVCFNCGEKGHKSNACPEE------IKKCVRCGKKGHVVADCNRT-----------D 294
A + C+ CGE GH S CP + C +CGK GH+ +C D
Sbjct: 193 AGDRKCYKCGESGHMSRECPSAGSTGSGDRACYKCGKPGHISRECPEAGGSYGGSRGGGD 252
Query: 295 IVCFNCNGEGHISSQC 310
C+ C GHIS C
Sbjct: 253 RTCYKCGEAGHISRDC 268
Score = 45.4 bits (106), Expect = 4e-04
Identities = 23/85 (27%), Positives = 30/85 (35%), Gaps = 31/85 (36%)
Query: 257 CFNCGEKGHKSNACPEE-----------------------IKKCVRCGKKGHVVADC--- 290
C+ CG++GH S CP + C +CG GH+ DC
Sbjct: 99 CYKCGQEGHLSRDCPSSQGGSRGGYGQKRGRSGAQGGYSGDRTCYKCGDAGHISRDCPNG 158
Query: 291 -----NRTDIVCFNCNGEGHISSQC 310
D C+ C GHIS C
Sbjct: 159 QGGYSGAGDRTCYKCGDAGHISRDC 183
>CNBP_MOUSE (P53996) Cellular nucleic acid binding protein (CNBP)
(Zinc finger protein 9)
Length = 178
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 252 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 310
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 115 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 172
Query: 311 T 311
T
Sbjct: 173 T 173
Score = 58.9 bits (141), Expect = 4e-08
Identities = 29/73 (39%), Positives = 38/73 (51%), Gaps = 7/73 (9%)
Query: 257 CFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNGEGHISSQCT 311
C+NCG GH + C E ++ C CGK GH+ DC+ D C++C GHI CT
Sbjct: 75 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCT 134
Query: 312 QPK--RAPTTGRV 322
+ K R TG V
Sbjct: 135 KVKCYRCGETGHV 147
Score = 58.5 bits (140), Expect = 5e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 243 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 301
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 87 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 142
Query: 302 GEGHISSQCTQ 312
GH++ C++
Sbjct: 143 ETGHVAINCSK 153
Score = 55.5 bits (132), Expect = 4e-07
Identities = 28/97 (28%), Positives = 47/97 (47%), Gaps = 14/97 (14%)
Query: 219 KGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACP-EEIKKC 277
+G+G +SR + +D+G Q + + +C+ CGE GH + C +E + C
Sbjct: 25 RGRGMRSRGRG-GFTSDRGFQFV--------SSSLPDICYRCGESGHLAKDCDLQEDEAC 75
Query: 278 VRCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 310
CG+ GH+ DC + C+NC GH++ C
Sbjct: 76 YNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 112
Score = 54.7 bits (130), Expect = 7e-07
Identities = 27/88 (30%), Positives = 36/88 (40%), Gaps = 29/88 (32%)
Query: 257 CFNCGEKGHKSNACPEEIKK----------------------------CVRCGKKGHVVA 288
CF CG GH + CP + C RCG+ GH+
Sbjct: 6 CFKCGRSGHWARECPTGGGRGRGMRSRGRGGFTSDRGFQFVSSSLPDICYRCGESGHLAK 65
Query: 289 DCN-RTDIVCFNCNGEGHISSQCTQPKR 315
DC+ + D C+NC GHI+ C +PKR
Sbjct: 66 DCDLQEDEACYNCGRGGHIAKDCKEPKR 93
Score = 43.9 bits (102), Expect = 0.001
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 130 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 172
>CNBP_CHICK (O42395) Cellular nucleic acid binding protein (CNBP)
Length = 172
Score = 60.1 bits (144), Expect = 2e-08
Identities = 25/61 (40%), Positives = 38/61 (61%), Gaps = 3/61 (4%)
Query: 252 AAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 310
A E C++CGE GH C + KC RCG+ GHV +C++T ++ C+ C GH++ +C
Sbjct: 109 ADEQKCYSCGEFGHIQKDCTKV--KCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 166
Query: 311 T 311
T
Sbjct: 167 T 167
Score = 59.7 bits (143), Expect = 2e-08
Identities = 28/82 (34%), Positives = 37/82 (44%), Gaps = 23/82 (28%)
Query: 257 CFNCGEKGHKSNACPEEIKK----------------------CVRCGKKGHVVADCN-RT 293
CF CG GH + CP I + C RCG+ GH+ DC+ +
Sbjct: 6 CFKCGRTGHWARECPTGIGRGRGMRSRGRAGFQFMSSSLPDICYRCGESGHLAKDCDLQE 65
Query: 294 DIVCFNCNGEGHISSQCTQPKR 315
D C+NC GHI+ C +PKR
Sbjct: 66 DKACYNCGRGGHIAKDCKEPKR 87
Score = 58.9 bits (141), Expect = 4e-08
Identities = 29/73 (39%), Positives = 38/73 (51%), Gaps = 7/73 (9%)
Query: 257 CFNCGEKGHKSNACPEEIKK----CVRCGKKGHVVADCNRTD-IVCFNCNGEGHISSQCT 311
C+NCG GH + C E ++ C CGK GH+ DC+ D C++C GHI CT
Sbjct: 69 CYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDCT 128
Query: 312 QPK--RAPTTGRV 322
+ K R TG V
Sbjct: 129 KVKCYRCGETGHV 141
Score = 58.5 bits (140), Expect = 5e-08
Identities = 25/71 (35%), Positives = 41/71 (57%), Gaps = 5/71 (7%)
Query: 243 DVRRPKKKDAAEIVCFNCGEKGHKSNACPE-EIKKCVRCGKKGHVVADCNRTDIVCFNCN 301
D + PK++ E C+NCG+ GH + C + +KC CG+ GH+ DC T + C+ C
Sbjct: 81 DCKEPKRE--REQCCYNCGKPGHLARDCDHADEQKCYSCGEFGHIQKDC--TKVKCYRCG 136
Query: 302 GEGHISSQCTQ 312
GH++ C++
Sbjct: 137 ETGHVAINCSK 147
Score = 56.6 bits (135), Expect = 2e-07
Identities = 22/60 (36%), Positives = 32/60 (52%), Gaps = 5/60 (8%)
Query: 256 VCFNCGEKGHKSNACP-EEIKKCVRCGKKGHVVADC----NRTDIVCFNCNGEGHISSQC 310
+C+ CGE GH + C +E K C CG+ GH+ DC + C+NC GH++ C
Sbjct: 47 ICYRCGESGHLAKDCDLQEDKACYNCGRGGHIAKDCKEPKREREQCCYNCGKPGHLARDC 106
Score = 43.9 bits (102), Expect = 0.001
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 1/43 (2%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
+KD ++ C+ CGE GH + C + + C RCG+ GH+ +C
Sbjct: 124 QKDCTKVKCYRCGETGHVAINCSKTSEVNCYRCGESGHLAREC 166
>BYR3_SCHPO (P36627) Cellular nucleic acid binding protein homolog
Length = 179
Score = 57.8 bits (138), Expect = 8e-08
Identities = 24/62 (38%), Positives = 34/62 (54%), Gaps = 7/62 (11%)
Query: 256 VCFNCGEKGHKSNAC--PEEIKKCVRCGKKGHVVADC-----NRTDIVCFNCNGEGHISS 308
+C+NC + GHK++ C P++ K C CG GH+V DC R C+ C GHI+
Sbjct: 37 ICYNCNQTGHKASECTEPQQEKTCYACGTAGHLVRDCPSSPNPRQGAECYKCGRVGHIAR 96
Query: 309 QC 310
C
Sbjct: 97 DC 98
Score = 53.5 bits (127), Expect = 2e-06
Identities = 23/59 (38%), Positives = 32/59 (53%), Gaps = 3/59 (5%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDI--VCFNCNGEGHISSQCTQP 313
C+ CG GH++ C +K C CGK GH +C + +C+ CN GHI+ CT P
Sbjct: 118 CYACGSYGHQARDCTMGVK-CYSCGKIGHRSFECQQASDGQLCYKCNQPGHIAVNCTSP 175
Score = 49.3 bits (116), Expect = 3e-05
Identities = 18/43 (41%), Positives = 28/43 (64%), Gaps = 1/43 (2%)
Query: 276 KCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPT 318
+C CG+ GH +C + I C+NCN GH +S+CT+P++ T
Sbjct: 18 RCYNCGENGHQARECTKGSI-CYNCNQTGHKASECTEPQQEKT 59
Score = 36.6 bits (83), Expect = 0.20
Identities = 12/38 (31%), Positives = 24/38 (62%), Gaps = 2/38 (5%)
Query: 255 IVCFNCGEKGHKSNACPE--EIKKCVRCGKKGHVVADC 290
+ C++CG+ GH+S C + + + C +C + GH+ +C
Sbjct: 135 VKCYSCGKIGHRSFECQQASDGQLCYKCNQPGHIAVNC 172
>GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-)
(Germline helicase-2)
Length = 974
Score = 55.1 bits (131), Expect = 5e-07
Identities = 31/101 (30%), Positives = 39/101 (37%), Gaps = 42/101 (41%)
Query: 257 CFNCGEKGHKSNACPEEIKK-----CVRCGKKGHVVADC--------------------- 290
CFNC + GH+SN CPE K+ C C + GH DC
Sbjct: 373 CFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSRDCPEERKPREGRNGFTSGFGGGN 432
Query: 291 ----------------NRTDIVCFNCNGEGHISSQCTQPKR 315
R + CFNC GEGH S++C +P R
Sbjct: 433 DGGFGGGNAEGFGNNEERGPMKCFNCKGEGHRSAECPEPPR 473
Score = 54.3 bits (129), Expect = 9e-07
Identities = 30/104 (28%), Positives = 39/104 (36%), Gaps = 37/104 (35%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIK--------------------------------- 275
KK+ VC+NC + GH S CPEE K
Sbjct: 390 KKEREPRVCYNCQQPGHNSRDCPEERKPREGRNGFTSGFGGGNDGGFGGGNAEGFGNNEE 449
Query: 276 ----KCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKR 315
KC C +GH A+C CFNC +GH S++C P +
Sbjct: 450 RGPMKCFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPNPAK 493
Score = 47.4 bits (111), Expect = 1e-04
Identities = 25/65 (38%), Positives = 31/65 (47%), Gaps = 16/65 (24%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRA 316
CFNC + GH+SN CPE K+ R VC+NC GH S C + +R
Sbjct: 259 CFNCQQPGHRSNDCPEPKKE---------------REPRVCYNCQQPGHNSRDCPE-ERK 302
Query: 317 PTTGR 321
P GR
Sbjct: 303 PREGR 307
Score = 45.1 bits (105), Expect = 6e-04
Identities = 19/48 (39%), Positives = 25/48 (51%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEG 304
CFNC +GH+S CPE + C CG++GH +C GEG
Sbjct: 455 CFNCKGEGHRSAECPEPPRGCFNCGEQGHRSNECPNPAKPREGAEGEG 502
Score = 35.8 bits (81), Expect = 0.34
Identities = 17/36 (47%), Positives = 21/36 (58%), Gaps = 2/36 (5%)
Query: 249 KKDAAEIVCFNCGEKGHKSNACPEEIKKCVRCGKKG 284
KK+ VC+NC + GH S CPEE K R G+ G
Sbjct: 276 KKEREPRVCYNCQQPGHNSRDCPEERKP--REGRNG 309
>ZCH3_HUMAN (Q9NUD5) Zinc finger CCHC domain containing protein 3
Length = 404
Score = 53.5 bits (127), Expect = 2e-06
Identities = 24/71 (33%), Positives = 37/71 (51%), Gaps = 3/71 (4%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRA 316
CF CG + H S +C ++ +C RCG++GH+ C R IVC C GH +QC +
Sbjct: 336 CFKCGSRTHMSGSCTQD--RCFRCGEEGHLSPYC-RKGIVCNLCGKRGHAFAQCPKAVHN 392
Query: 317 PTTGRVFALTG 327
++ + G
Sbjct: 393 SVAAQLTGVAG 403
Score = 37.4 bits (85), Expect = 0.12
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 2/36 (5%)
Query: 275 KKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQC 310
K C +CG + H+ C T CF C EGH+S C
Sbjct: 334 KTCFKCGSRTHMSGSC--TQDRCFRCGEEGHLSPYC 367
>YL92_SCHPO (Q9HFF2) Hypothetical protein C683.02c in chromosome I
Length = 218
Score = 53.5 bits (127), Expect = 2e-06
Identities = 31/97 (31%), Positives = 43/97 (43%), Gaps = 18/97 (18%)
Query: 234 ADKGKQKMVDVRRPKKKDA-----------AEIVCFNCGEKGHKSNACPE---EIKKCVR 279
A G K D R+ KK+ + CF C ++GH CPE + C R
Sbjct: 45 ASFGSSKRYDERQKKKRSEYRRLRRINQRNRDKFCFACRQQGHIVQDCPEAKDNVSICFR 104
Query: 280 CGKKGHVVADCNRTDIV----CFNCNGEGHISSQCTQ 312
CG K H + C++ + CF C+ GH+S QC Q
Sbjct: 105 CGSKEHSLNACSKKGPLKFAKCFICHENGHLSGQCEQ 141
Score = 51.2 bits (121), Expect = 8e-06
Identities = 31/100 (31%), Positives = 43/100 (43%), Gaps = 13/100 (13%)
Query: 247 PKKKDAAEIVCFNCGEKGHKSNAC----PEEIKKCVRCGKKGHVVADCNRTDI------- 295
P+ KD I CF CG K H NAC P + KC C + GH+ C +
Sbjct: 93 PEAKDNVSI-CFRCGSKEHSLNACSKKGPLKFAKCFICHENGHLSGQCEQNPKGLYPKGG 151
Query: 296 VCFNCNGEGHISSQCTQPKRAPTT-GRVFALTGTQTENED 334
C C+ H++ C Q + + G V + GT +ED
Sbjct: 152 CCKFCSSVHHLAKDCDQVNKDDVSFGHVVGVAGTTGADED 191
>GLH4_CAEEL (O76743) ATP-dependent RNA helicase glh-4 (EC 3.6.1.-)
(Germline helicase-4)
Length = 1156
Score = 52.8 bits (125), Expect = 3e-06
Identities = 25/64 (39%), Positives = 36/64 (56%), Gaps = 6/64 (9%)
Query: 257 CFNCGEKGHKSNACPE-EIKK--CVRCGKKGHVVADCNRTDIV---CFNCNGEGHISSQC 310
C NCGE+GH S C + ++ + C C + GH +DC++ + C NC EGH + C
Sbjct: 572 CHNCGEEGHISKECDKPKVPRFPCRNCEQLGHFASDCDQPRVPRGPCRNCGIEGHFAVDC 631
Query: 311 TQPK 314
QPK
Sbjct: 632 DQPK 635
Score = 43.5 bits (101), Expect = 0.002
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 6/60 (10%)
Query: 257 CFNCGEKGHKSNACPEEIKK---CVRCGKKGHVVADCNRTDI---VCFNCNGEGHISSQC 310
C NC + GH ++ C + C CG +GH DC++ + C NC EGH + C
Sbjct: 595 CRNCEQLGHFASDCDQPRVPRGPCRNCGIEGHFAVDCDQPKVPRGPCRNCGQEGHFAKDC 654
Score = 41.2 bits (95), Expect = 0.008
Identities = 16/46 (34%), Positives = 26/46 (55%), Gaps = 3/46 (6%)
Query: 272 EEIKKCVRCGKKGHVVADCNRTDI---VCFNCNGEGHISSQCTQPK 314
E + C CG++GH+ +C++ + C NC GH +S C QP+
Sbjct: 567 ERPRGCHNCGEEGHISKECDKPKVPRFPCRNCEQLGHFASDCDQPR 612
Score = 33.1 bits (74), Expect = 2.2
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 6/40 (15%)
Query: 257 CFNCGEKGHKSNACPEE------IKKCVRCGKKGHVVADC 290
C NCG++GH + C E + C RC ++GH +C
Sbjct: 641 CRNCGQEGHFAKDCQNERVRMEPTEPCRRCAEEGHWGYEC 680
>GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 519
Score = 52.4 bits (124), Expect = 4e-06
Identities = 24/53 (45%), Positives = 32/53 (60%), Gaps = 2/53 (3%)
Query: 239 QKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
Q MV PK++ + C+NCG+ GH CPE K KC++CGK GH+ DC
Sbjct: 382 QNMVQQGGPKRQ-RPPLRCYNCGKFGHMQRQCPEPRKTKCLKCGKLGHLAKDC 433
>GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 513
Score = 52.4 bits (124), Expect = 4e-06
Identities = 29/69 (42%), Positives = 34/69 (49%), Gaps = 6/69 (8%)
Query: 239 QKMVDVRRPKKKDAAEIVCFNCGEKGHKSNAC--PEEIKKCVRCGKKGHVVADCNRTDIV 296
Q MV V KK + CFNCG+ GH C P +I KC +CGK GH+ DC
Sbjct: 374 QNMVQVGPQKKGPRGPLKCFNCGKFGHMQRECKAPRQI-KCFKCGKIGHMAKDCKNGQA- 431
Query: 297 CFNCNGEGH 305
N G GH
Sbjct: 432 --NFLGYGH 438
Score = 37.4 bits (85), Expect = 0.12
Identities = 15/36 (41%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Query: 276 KCVRCGKKGHVVADCNRT-DIVCFNCNGEGHISSQC 310
KC CGK GH+ +C I CF C GH++ C
Sbjct: 391 KCFNCGKFGHMQRECKAPRQIKCFKCGKIGHMAKDC 426
>GAG_SIVAG (P27978) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 521
Score = 50.1 bits (118), Expect = 2e-05
Identities = 20/53 (37%), Positives = 30/53 (55%), Gaps = 1/53 (1%)
Query: 239 QKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
Q M+ + + + C+NCG+ GH CPE K +C++CGK GH+ DC
Sbjct: 386 QNMMQQGGQRGRPRPPVKCYNCGKFGHMQRQCPEPRKMRCLKCGKPGHLAKDC 438
>GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 520
Score = 50.1 bits (118), Expect = 2e-05
Identities = 19/35 (54%), Positives = 24/35 (68%), Gaps = 1/35 (2%)
Query: 257 CFNCGEKGHKSNACPEEIK-KCVRCGKKGHVVADC 290
C+NCG+ GH CPE K KC++CGK GH+ DC
Sbjct: 400 CYNCGKFGHMQRQCPEPRKIKCLKCGKPGHLAKDC 434
Score = 35.4 bits (80), Expect = 0.44
Identities = 15/41 (36%), Positives = 18/41 (43%), Gaps = 1/41 (2%)
Query: 271 PEEIKKCVRCGKKGHVVADC-NRTDIVCFNCNGEGHISSQC 310
P KC CGK GH+ C I C C GH++ C
Sbjct: 394 PRPPPKCYNCGKFGHMQRQCPEPRKIKCLKCGKPGHLAKDC 434
>YIH9_YEAST (P40507) Hypothetical 41.6 kDa protein in SDS3-THS1
intergenic region
Length = 360
Score = 49.7 bits (117), Expect = 2e-05
Identities = 25/59 (42%), Positives = 29/59 (48%), Gaps = 4/59 (6%)
Query: 253 AEIVCFNCGEKGHKSNACPEEIKKCVRCG-KKGHVVADCNRTDIVCFNCNGEGHISSQC 310
AE C NC ++GH CP I C CG H C + I+C NCN GH SQC
Sbjct: 72 AEPKCNNCSQRGHLKRNCPHVI--CTYCGFMDDHYSQHCPKA-IICTNCNANGHYKSQC 127
Score = 46.2 bits (108), Expect = 3e-04
Identities = 26/88 (29%), Positives = 34/88 (38%), Gaps = 23/88 (26%)
Query: 255 IVCFNCGEKGHKSNACPEEIKK--CVRCGKKGH--------------VVADCNRTD---- 294
I+C NC GH + CP + KK C C K H D N+ D
Sbjct: 112 IICTNCNANGHYKSQCPHKWKKVFCTLCNSKRHSRERCPSIWRSYLLKTKDANQGDFDFQ 171
Query: 295 -IVCFNCNGEGHISSQCTQPK--RAPTT 319
+ C+NC GH C + + R P T
Sbjct: 172 TVFCYNCGNAGHFGDDCAERRSSRVPNT 199
>GAG_HV1J3 (P12494) Gag polyprotein [Contains: Core protein p17
(Matrix protein); Core protein p24 (Core antigen); Core
protein p2; Core protein p7 (Nucleocapsid protein); Core
protein p1; Core protein p6]
Length = 499
Score = 48.1 bits (113), Expect = 7e-05
Identities = 20/38 (52%), Positives = 26/38 (67%), Gaps = 1/38 (2%)
Query: 255 IVCFNCGEKGHKSNACPEEIKK-CVRCGKKGHVVADCN 291
I CFNCG++GH + C KK C +CGK+GH + DCN
Sbjct: 389 IKCFNCGKEGHLARNCRAPRKKGCWKCGKEGHQMKDCN 426
Score = 37.0 bits (84), Expect = 0.15
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Query: 273 EIKKCVRCGKKGHVVADCNRTDIV-CFNCNGEGHISSQCTQPKRAPTTGRVF 323
+I KC CGK+GH+ +C C+ C EGH C + ++A G+++
Sbjct: 387 KIIKCFNCGKEGHLARNCRAPRKKGCWKCGKEGHQMKDCNE-RQANFLGKIW 437
Score = 33.1 bits (74), Expect = 2.2
Identities = 18/63 (28%), Positives = 28/63 (43%), Gaps = 6/63 (9%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCG----KKGHVVADCNRTDIVCFNCNGEGHISSQCTQ 312
C G GHK+ E + + ++G+ R I CFNC EGH++ C
Sbjct: 349 CQGVGGPGHKARVLAEAMSQVTNSTTIMMQRGNFRNQ--RKIIKCFNCGKEGHLARNCRA 406
Query: 313 PKR 315
P++
Sbjct: 407 PRK 409
>GAG_FIVPE (P16087) Gag polyprotein [Contains: Core protein p15;
Major core protein p24; Nucleic acid binding protein
p10]
Length = 450
Score = 48.1 bits (113), Expect = 7e-05
Identities = 19/35 (54%), Positives = 24/35 (68%), Gaps = 1/35 (2%)
Query: 256 VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADC 290
VCFNC + GH + C E+KKC +CGK GH+ A C
Sbjct: 376 VCFNCKKPGHLARQC-REVKKCNKCGKPGHLAAKC 409
Score = 35.4 bits (80), Expect = 0.44
Identities = 19/65 (29%), Positives = 27/65 (41%), Gaps = 8/65 (12%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRA 316
C G G+K E + K KG + VCFNC GH++ QC + K+
Sbjct: 345 CQEIGSPGYKMQLLAEALTKVQVVQSKG--------SGPVCFNCKKPGHLARQCREVKKC 396
Query: 317 PTTGR 321
G+
Sbjct: 397 NKCGK 401
>GAG_SIVSP (P19504) Gag polyprotein [Contains: Core protein p17;
Core protein p24]
Length = 507
Score = 47.8 bits (112), Expect = 9e-05
Identities = 22/65 (33%), Positives = 36/65 (54%), Gaps = 14/65 (21%)
Query: 227 PKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEIKK-CVRCGKKGH 285
P P++A KG++K++ C+NCG++GH + C ++ C +CGK GH
Sbjct: 377 PLPFAAVQQKGQRKIIK-------------CWNCGKEGHSARQCRAPRRQGCWKCGKAGH 423
Query: 286 VVADC 290
V+A C
Sbjct: 424 VMAKC 428
Score = 34.7 bits (78), Expect = 0.76
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 2/61 (3%)
Query: 257 CFNCGEKGHKSNACPEEIKKCVRCGKK--GHVVADCNRTDIVCFNCNGEGHISSQCTQPK 314
C G G K+ E +K + G V R I C+NC EGH + QC P+
Sbjct: 352 CQGVGGPGQKARLMAEALKDALTQGPLPFAAVQQKGQRKIIKCWNCGKEGHSARQCRAPR 411
Query: 315 R 315
R
Sbjct: 412 R 412
Score = 34.3 bits (77), Expect = 0.99
Identities = 14/41 (34%), Positives = 21/41 (51%), Gaps = 1/41 (2%)
Query: 273 EIKKCVRCGKKGHVVADCNRTDIV-CFNCNGEGHISSQCTQ 312
+I KC CGK+GH C C+ C GH+ ++C +
Sbjct: 390 KIIKCWNCGKEGHSARQCRAPRRQGCWKCGKAGHVMAKCPE 430
>GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 446
Score = 47.8 bits (112), Expect = 9e-05
Identities = 19/46 (41%), Positives = 26/46 (56%), Gaps = 1/46 (2%)
Query: 274 IKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPTT 319
++KC CGK GH+ C R I+C +C GH+ C Q K PT+
Sbjct: 383 VQKCYYCGKPGHLARQC-RQGIICHHCGKRGHMQKDCRQKKGNPTS 427
Score = 43.9 bits (102), Expect = 0.001
Identities = 25/81 (30%), Positives = 40/81 (48%), Gaps = 12/81 (14%)
Query: 239 QKMVDVRRPKKKDAAEIV--CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIV 296
Q + RP +K+ + V C+ CG+ GH + C + I C CGK+GH+ DC +
Sbjct: 366 QLLAQALRPPRKEGKQGVQKCYYCGKPGHLARQCRQGI-ICHHCGKRGHMQKDCRQK--- 421
Query: 297 CFNCNGEGHISSQCTQPKRAP 317
+G+ +SQ +R P
Sbjct: 422 ------KGNPTSQQGNSRRGP 436
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 70,941,755
Number of Sequences: 164201
Number of extensions: 3058422
Number of successful extensions: 8450
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 51
Number of HSP's successfully gapped in prelim test: 74
Number of HSP's that attempted gapping in prelim test: 7874
Number of HSP's gapped (non-prelim): 409
length of query: 615
length of database: 59,974,054
effective HSP length: 116
effective length of query: 499
effective length of database: 40,926,738
effective search space: 20422442262
effective search space used: 20422442262
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC146333.2


