

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.2 + phase: 0
(404 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
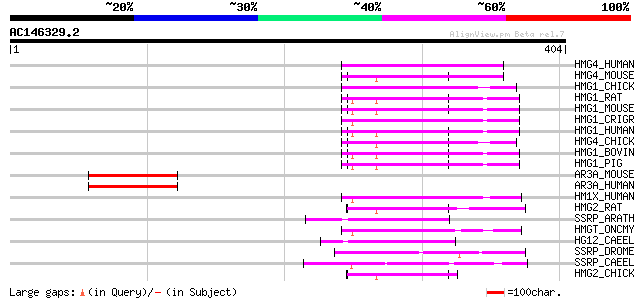
Score E
Sequences producing significant alignments: (bits) Value
HMG4_HUMAN (O15347) High mobility group protein 4 (HMG-4) (High ... 67 8e-11
HMG4_MOUSE (O54879) High mobility group protein 4 (HMG-4) (High ... 64 5e-10
HMG1_CHICK (P36194) High mobility group protein 1 (HMG-1) 64 7e-10
HMG1_RAT (P63159) High mobility group protein 1 (HMG-1) (Amphote... 64 9e-10
HMG1_MOUSE (P63158) High mobility group protein 1 (HMG-1) 64 9e-10
HMG1_CRIGR (P07156) High mobility group protein 1 (HMG-1) (Fragm... 64 9e-10
HMG1_HUMAN (P09429) High mobility group protein 1 (HMG-1) 63 1e-09
HMG4_CHICK (P40618) High mobility group protein 4 (HMG-4) (High ... 62 3e-09
HMG1_BOVIN (P10103) High mobility group protein 1 (HMG-1) 62 3e-09
HMG1_PIG (P12682) High mobility group protein 1 (HMG-1) 61 4e-09
AR3A_MOUSE (Q62431) AT-rich interactive domain-containing protei... 60 8e-09
AR3A_HUMAN (Q99856) AT-rich interactive domain-containing protei... 60 8e-09
HM1X_HUMAN (Q9UGV6) High mobility group protein 1-like 10 (HMG-1... 60 1e-08
HMG2_RAT (P52925) High mobility group protein 2 (HMG-2) 59 2e-08
SSRP_ARATH (Q05153) Structure-specific recognition protein 1 hom... 59 2e-08
HMGT_ONCMY (P07746) High mobility group-T protein (HMG-T) (HMG-T... 59 2e-08
HG12_CAEEL (Q09390) High mobility group protein 1.2 59 2e-08
SSRP_DROME (Q05344) Single-strand recognition protein (SSRP) (Ch... 57 8e-08
SSRP_CAEEL (P41848) Probable structure-specific recognition prot... 57 8e-08
HMG2_CHICK (P26584) High mobility group protein 2 (HMG-2) 57 8e-08
>HMG4_HUMAN (O15347) High mobility group protein 4 (HMG-4) (High
mobility group protein 2a) (HMG-2a)
Length = 199
Score = 67.0 bits (162), Expect = 8e-11
Identities = 38/119 (31%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L +SEK
Sbjct: 81 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLNDSEKQP 140
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E D + E + + ++
Sbjct: 141 YITKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEDEEQEEEEEEEEEEEDE 199
Score = 43.1 bits (100), Expect = 0.001
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EK+ + +
Sbjct: 2 KGDPKKPKGKTSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTVSGKEKSKFDE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKVRYDREMKDY 77
>HMG4_MOUSE (O54879) High mobility group protein 4 (HMG-4) (High
mobility group protein 2a) (HMG-2a)
Length = 199
Score = 64.3 bits (155), Expect = 5e-10
Identities = 36/119 (30%), Positives = 63/119 (52%), Gaps = 1/119 (0%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L ++EK
Sbjct: 81 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLSDNEKQP 140
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQ 359
Y KA K KE+Y ++ Y+ K K D A R+++ E + + E + + ++
Sbjct: 141 YVTKAAKLKEKYEKDVADYKSKGKFDGAKGPAKVARKKVEEEEEEEEEEEEEEEEEEDE 199
Score = 44.3 bits (103), Expect = 6e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EK+ + +
Sbjct: 2 KGDPKKPKGKMSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTMSSKEKSKFDE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKVRYDREMKDY 77
>HMG1_CHICK (P36194) High mobility group protein 1 (HMG-1)
Length = 200
Score = 63.9 bits (154), Expect = 7e-10
Identities = 40/129 (31%), Positives = 63/129 (48%), Gaps = 9/129 (6%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK SG+ F +E P++K + G ++++ +GE+WN L + EK
Sbjct: 80 KGGKKKKDPNAPKRPPSGFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLSDGEKQP 139
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y +KA K KE+Y ++ Y+ K K D A ++ E E D + ++
Sbjct: 140 YNNKAAKLKEKYEKDVADYKSKGKFDGAKGAATKAARKKVE--------EEDEEEEEDEE 191
Query: 361 SLDGSDDYE 369
D DD E
Sbjct: 192 EEDEDDDDE 200
Score = 41.6 bits (96), Expect = 0.004
Identities = 27/76 (35%), Positives = 34/76 (44%), Gaps = 4/76 (5%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EKA + +
Sbjct: 2 KGDPKKPKGKMSAYAFFVQTCREEHKK-NPEVPVNFAEFSKKCSERWKTMSSKEKAKFDE 60
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 61 MAKADKVRYDREMKDY 76
>HMG1_RAT (P63159) High mobility group protein 1 (HMG-1)
(Amphoterin) (Heparin-binding protein p30)
Length = 214
Score = 63.5 bits (153), Expect = 9e-10
Identities = 40/133 (30%), Positives = 64/133 (48%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 81 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + + E D E
Sbjct: 141 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--DDEEDEEDEEE 198
Query: 359 QSSLDGSDDYEDD 371
+ + D+ EDD
Sbjct: 199 EEEEEDEDEEEDD 211
Score = 45.4 bits (106), Expect = 3e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK ++D
Sbjct: 2 KGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKARYEREMKTY 77
>HMG1_MOUSE (P63158) High mobility group protein 1 (HMG-1)
Length = 214
Score = 63.5 bits (153), Expect = 9e-10
Identities = 40/133 (30%), Positives = 64/133 (48%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 81 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + + E D E
Sbjct: 141 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--DDEEDEEDEEE 198
Query: 359 QSSLDGSDDYEDD 371
+ + D+ EDD
Sbjct: 199 EEEEEDEDEEEDD 211
Score = 45.4 bits (106), Expect = 3e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK ++D
Sbjct: 2 KGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKARYEREMKTY 77
>HMG1_CRIGR (P07156) High mobility group protein 1 (HMG-1)
(Fragment)
Length = 180
Score = 63.5 bits (153), Expect = 9e-10
Identities = 40/133 (30%), Positives = 64/133 (48%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 47 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 106
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + + E D E
Sbjct: 107 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--DDEEDEEDEEE 164
Query: 359 QSSLDGSDDYEDD 371
+ + D+ EDD
Sbjct: 165 EEEEEDEDEEEDD 177
Score = 33.5 bits (75), Expect = 0.99
Identities = 15/39 (38%), Positives = 20/39 (50%)
Query: 281 EISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYY 319
E S+ E W + EK ++D A DK RY EM+ Y
Sbjct: 5 EFSKKCSERWKTMSAKEKGKFEDMAKADKARYEREMKTY 43
>HMG1_HUMAN (P09429) High mobility group protein 1 (HMG-1)
Length = 214
Score = 63.2 bits (152), Expect = 1e-09
Identities = 40/133 (30%), Positives = 63/133 (47%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 81 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + E D E
Sbjct: 141 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--EDEEDEEDEEE 198
Query: 359 QSSLDGSDDYEDD 371
+ + D+ EDD
Sbjct: 199 EEDEEDEDEEEDD 211
Score = 45.4 bits (106), Expect = 3e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK ++D
Sbjct: 2 KGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKARYEREMKTY 77
>HMG4_CHICK (P40618) High mobility group protein 4 (HMG-4) (High
mobility group protein 2a) (HMG-2a)
Length = 201
Score = 61.6 bits (148), Expect = 3e-09
Identities = 39/129 (30%), Positives = 62/129 (47%), Gaps = 9/129 (6%)
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAV 300
K KK+DP PK S + F +E P++K + G ++++ +GE+WN L + EK
Sbjct: 81 KGGKKKKDPNAPKRPPSAFFLFCSEFRPKIKSTNPGISIGDVAKKLGEMWNNLSDGEKQP 140
Query: 301 YQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQS 360
Y +KA K KE+Y ++ Y+ K K D A ++ E E D + ++
Sbjct: 141 YNNKAAKLKEKYEKDVADYKSKGKFDGAKGAATKAARKKVE--------EEDEEEEEDEE 192
Query: 361 SLDGSDDYE 369
D DD E
Sbjct: 193 EEDEDDDDE 201
Score = 45.4 bits (106), Expect = 3e-04
Identities = 27/76 (35%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP PK S Y FF E+H + P E S+ E W + EKA + +
Sbjct: 2 KGDPKKPKGKMSAYAFFVQTCREEHKKKNPEVPVNFAEFSKKCSERWKTMSSKEKAKFDE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKVRYDREMKDY 77
>HMG1_BOVIN (P10103) High mobility group protein 1 (HMG-1)
Length = 214
Score = 61.6 bits (148), Expect = 3e-09
Identities = 39/133 (29%), Positives = 63/133 (47%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 81 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + E D E
Sbjct: 141 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--EDEEDEEDEEE 198
Query: 359 QSSLDGSDDYEDD 371
+ + ++ EDD
Sbjct: 199 EEDEEDEEEEEDD 211
Score = 45.4 bits (106), Expect = 3e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK ++D
Sbjct: 2 KGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKARYEREMKTY 77
>HMG1_PIG (P12682) High mobility group protein 1 (HMG-1)
Length = 214
Score = 61.2 bits (147), Expect = 4e-09
Identities = 39/133 (29%), Positives = 63/133 (47%), Gaps = 5/133 (3%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F +E P++K H G ++++ +GE+WN +K
Sbjct: 81 KGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + E D E
Sbjct: 141 HPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE--EDEEDEEDEEE 198
Query: 359 QSSLDGSDDYEDD 371
+ + ++ EDD
Sbjct: 199 EEDEEDEEEEEDD 211
Score = 45.4 bits (106), Expect = 3e-04
Identities = 26/76 (34%), Positives = 34/76 (44%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK ++D
Sbjct: 2 KGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKADKARYEREMKTY 77
>AR3A_MOUSE (Q62431) AT-rich interactive domain-containing protein
3A (ARID domain-containing protein 3A) (Dead ringer
like-1 protein) (B-cell regulator of IgH transcription)
(Bright)
Length = 601
Score = 60.5 bits (145), Expect = 8e-09
Identities = 27/65 (41%), Positives = 41/65 (62%)
Query: 58 IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSL 117
IP++ + LDL L+V VT +GG ++I + W+E+T N P++ T+A+F LR Y
Sbjct: 267 IPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGLNLPTSITSAAFTLRTQYMKY 326
Query: 118 LYHYE 122
LY YE
Sbjct: 327 LYPYE 331
>AR3A_HUMAN (Q99856) AT-rich interactive domain-containing protein
3A (ARID domain-containing protein 3A) (B-cell regulator
of IgH transcription) (Bright) (E2F binding protein 1)
Length = 593
Score = 60.5 bits (145), Expect = 8e-09
Identities = 27/65 (41%), Positives = 41/65 (62%)
Query: 58 IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSL 117
IP++ + LDL L+V VT +GG ++I + W+E+T N P++ T+A+F LR Y
Sbjct: 262 IPIMAKQVLDLFMLYVLVTEKGGLVEVINKKLWREITKGLNLPTSITSAAFTLRTQYMKY 321
Query: 118 LYHYE 122
LY YE
Sbjct: 322 LYPYE 326
>HM1X_HUMAN (Q9UGV6) High mobility group protein 1-like 10
(HMG-1L10)
Length = 211
Score = 59.7 bits (143), Expect = 1e-08
Identities = 38/134 (28%), Positives = 64/134 (47%), Gaps = 7/134 (5%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KK+ DP PK S + F + P++K H G ++++ +GE+WN +K
Sbjct: 82 KGETKKKFKDPNAPKRTPSAFFLFCSAYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDK 141
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA K KE+Y ++ YR K K D V + + + + E + + E
Sbjct: 142 QPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEE----EDEEDEEDE 197
Query: 359 QSSLDGSDDYEDDK 372
+ + +D EDD+
Sbjct: 198 EEEDEEDEDEEDDE 211
Score = 37.7 bits (86), Expect = 0.053
Identities = 24/78 (30%), Positives = 32/78 (40%), Gaps = 3/78 (3%)
Query: 245 MKKRDPAHPKPNRSGYNFF---FAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
M K DP + S + FF E H + P E S+ E W + EK +
Sbjct: 1 MSKGDPKKLRGKMSSHAFFGQTCREAHKKKHPDASVNLSEFSKKCSERWKTMSAKEKGKF 60
Query: 302 QDKAVKDKERYITEMEYY 319
+D A DK Y EM+ Y
Sbjct: 61 EDMAKADKAHYEREMKTY 78
>HMG2_RAT (P52925) High mobility group protein 2 (HMG-2)
Length = 209
Score = 59.3 bits (142), Expect = 2e-08
Identities = 36/131 (27%), Positives = 64/131 (48%), Gaps = 9/131 (6%)
Query: 246 KKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAVYQDK 304
KK+DP PK S + F +E P++K H G + ++ +GE+W++ +K Y+ K
Sbjct: 87 KKKDPNAPKRPPSAFFLFCSEHRPKIKSEHPGLSIGDTAKKLGEMWSEQSAKDKQPYEQK 146
Query: 305 AVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQSSLDG 364
A K KE+Y ++ YR K K++ + ++ P T + E+ D
Sbjct: 147 AAKLKEKYEKDIAAYRAKGKSE--------VGKKGPGRPTGSKKKNEPEDEEEEEEEEDD 198
Query: 365 SDDYEDDKAKE 375
D+ E+D+ +E
Sbjct: 199 EDEEEEDEDEE 209
Score = 45.8 bits (107), Expect = 2e-04
Identities = 26/76 (34%), Positives = 35/76 (45%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y FF E+H + P E S+ E W + EK+ ++D
Sbjct: 2 KGDPNKPRGKMSSYAFFVQTCREEHKKKHPDSSVNFAEFSKKCSERWKTMSAKEKSKFED 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 LAKSDKARYDREMKNY 77
>SSRP_ARATH (Q05153) Structure-specific recognition protein 1
homolog (HMG protein)
Length = 646
Score = 58.9 bits (141), Expect = 2e-08
Identities = 33/106 (31%), Positives = 53/106 (49%), Gaps = 5/106 (4%)
Query: 216 ASHHSVPANNNNVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLH 275
+S +P V A G +R++ KK K+DP PK SG+ FF + +K H
Sbjct: 529 SSSKGLPPKRKTVAADEGSSKRKKPKK----KKDPNAPKRAMSGFMFFSQMERDNIKKEH 584
Query: 276 RG-KDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYR 320
G E+ + +G+ W ++ +K Y+ KA DK+RY E+ Y+
Sbjct: 585 PGIAFGEVGKVLGDKWRQMSADDKEPYEAKAQVDKQRYKDEISDYK 630
>HMGT_ONCMY (P07746) High mobility group-T protein (HMG-T) (HMG-T1)
(HMG-1)
Length = 204
Score = 58.9 bits (141), Expect = 2e-08
Identities = 42/134 (31%), Positives = 63/134 (46%), Gaps = 13/134 (9%)
Query: 242 KSEMKKR--DPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
K E KKR DP PK S + F A+ P++K G ++++ +GE WN L +K
Sbjct: 81 KGEKKKRFKDPNAPKRPSSAFFIFCADFRPQVKGETPGLSIGDVAKKLGEKWNNLTAEDK 140
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA + KE+Y ++ YR K K + ++P + P D D D +
Sbjct: 141 VPYEKKASRLKEKYEKDITAYRNKGK----VPVSMPAKAAAPAKDDD------DDDDDDD 190
Query: 359 QSSLDGSDDYEDDK 372
D DD EDD+
Sbjct: 191 DDEDDDDDDDEDDE 204
Score = 42.7 bits (99), Expect = 0.002
Identities = 24/75 (32%), Positives = 33/75 (44%), Gaps = 3/75 (4%)
Query: 248 RDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDK 304
+DP P+ S Y +F E+H + P E S+ E W + EK ++D
Sbjct: 3 KDPRKPRGKMSSYAYFVQTRREEHKKKHPEASVNFSEFSKKCSERWKTMSAKEKGKFEDL 62
Query: 305 AVKDKERYITEMEYY 319
A DK RY EM Y
Sbjct: 63 AKLDKVRYEREMRSY 77
>HG12_CAEEL (Q09390) High mobility group protein 1.2
Length = 235
Score = 58.9 bits (141), Expect = 2e-08
Identities = 30/99 (30%), Positives = 57/99 (57%), Gaps = 4/99 (4%)
Query: 227 NVTASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRG-KDREISRT 285
+V A G R+RK++ K+DP PK S + F+ ++ P ++ H K ++++
Sbjct: 112 SVAAYGGEDAMRKRKRA---KKDPHAPKRALSAFFFYSQDKRPEIQAGHPDWKVGQVAQE 168
Query: 286 IGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLK 324
+G++W +P+ K +Y+ KA DK+RY EM Y+ +++
Sbjct: 169 LGKMWKLVPQETKDMYEQKAQADKDRYADEMRNYKAEMQ 207
Score = 37.0 bits (84), Expect = 0.090
Identities = 24/74 (32%), Positives = 34/74 (45%), Gaps = 5/74 (6%)
Query: 248 RDPAHP--KPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQ 302
RD P + S Y FF E+H + P + EIS+ E W + + EK +
Sbjct: 38 RDMGKPPVRGKTSPYGFFVKMCYEEHKKKYPNENVQVTEISKKCSEKWKTMVDDEKRRFY 97
Query: 303 DKAVKDKERYITEM 316
+ A KD ERY E+
Sbjct: 98 ELAQKDAERYQAEV 111
>SSRP_DROME (Q05344) Single-strand recognition protein (SSRP)
(Chorion-factor 5)
Length = 723
Score = 57.0 bits (136), Expect = 8e-08
Identities = 41/143 (28%), Positives = 66/143 (45%), Gaps = 7/143 (4%)
Query: 237 RRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRG-KDREISRTIGELWNKLPE 295
+ R KK KK+D PK + + + + +K + G K EI++ GE+W +L +
Sbjct: 539 KERTKKPSKKKKDSGKPKRATTAFMLWLNDTRESIKRENPGIKVTEIAKKGGEMWKELKD 598
Query: 296 SEKAVYQDKAVKDKERYITEMEYYREKLKND---EVISDAVPLRQRLPEPDTDMLNAEAD 352
K ++D A KDK+RY EM Y+ + D E + R+ P P + N
Sbjct: 599 KSK--WEDAAAKDKQRYHDEMRNYKPEAGGDSDNEKGGKSSKKRKTEPSP-SKKANTSGS 655
Query: 353 SLQTPEQSSLDGSDDYEDDKAKE 375
++ E S D S +D+K E
Sbjct: 656 GFKSKEYISDDDSTSSDDEKDNE 678
>SSRP_CAEEL (P41848) Probable structure-specific recognition protein
1 (SSRP1) (Recombination signal sequence recognition
protein)
Length = 697
Score = 57.0 bits (136), Expect = 8e-08
Identities = 43/167 (25%), Positives = 75/167 (44%), Gaps = 13/167 (7%)
Query: 215 PASHHSVPANNNNVTASVGVHRRRRRKKSEMKK----RDPAHPKPNRSGYNFFFAEQHPR 270
P S VP+ ++ +R+K E KK +DP PK S Y +F
Sbjct: 514 PDSEQDVPSKRRKGEPKEKREKKEKREKKEGKKGKKDKDPNAPKRATSAYMQWFLASRNE 573
Query: 271 LKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVIS 330
LK ++++ G W + +K +++KA +DK RY EM+ YR KN S
Sbjct: 574 LKE-DGDSVADVAKKGGAKWKTMSSDDKKKWEEKAEEDKSRYEKEMKEYR---KNGPPSS 629
Query: 331 DAVPLRQRLPEPDTDMLNAEADSLQTPEQSSLDGSDDYEDDKAKEKD 377
+ P + + + +++A S + + SDD +D++ K+K+
Sbjct: 630 SSKPSSSKTSKKSSGPSSSKAIS-----KEYISDSDDSDDEEPKKKE 671
>HMG2_CHICK (P26584) High mobility group protein 2 (HMG-2)
Length = 206
Score = 57.0 bits (136), Expect = 8e-08
Identities = 29/82 (35%), Positives = 47/82 (56%), Gaps = 1/82 (1%)
Query: 246 KKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEKAVYQDK 304
KK+DP PK S + F +E P++K H G + ++ +GE+W++ +K Y+ K
Sbjct: 87 KKKDPNAPKRPPSAFFLFCSEHRPKIKNDHPGLSIGDTAKKLGEMWSEQLAKDKQPYEQK 146
Query: 305 AVKDKERYITEMEYYREKLKND 326
A K KE+Y ++ YR K K+D
Sbjct: 147 AAKLKEKYEKDIAAYRAKSKSD 168
Score = 43.9 bits (102), Expect = 7e-04
Identities = 25/76 (32%), Positives = 34/76 (43%), Gaps = 3/76 (3%)
Query: 247 KRDPAHPKPNRSGYNFFFA---EQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQD 303
K DP P+ S Y +F E+H + P E SR E W + EK +++
Sbjct: 2 KGDPNKPRGKMSSYAYFVQTCREEHKKKHPDSSVNFAEFSRKCSERWKTMSSKEKGKFEE 61
Query: 304 KAVKDKERYITEMEYY 319
A DK RY EM+ Y
Sbjct: 62 MAKGDKARYDREMKNY 77
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,468,744
Number of Sequences: 164201
Number of extensions: 2306914
Number of successful extensions: 5990
Number of sequences better than 10.0: 161
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 5780
Number of HSP's gapped (non-prelim): 236
length of query: 404
length of database: 59,974,054
effective HSP length: 113
effective length of query: 291
effective length of database: 41,419,341
effective search space: 12053028231
effective search space used: 12053028231
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC146329.2


