

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.15 - phase: 0
(204 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
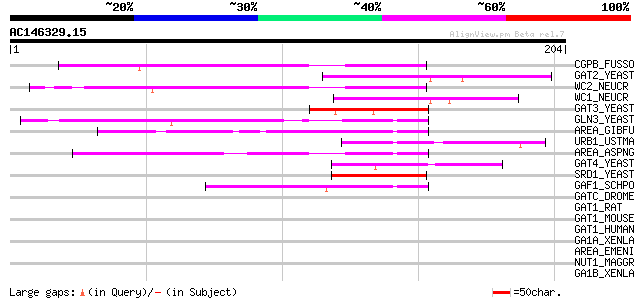
Score E
Sequences producing significant alignments: (bits) Value
CGPB_FUSSO (Q00858) Cutinase gene palindrome-binding protein (PBP) 60 3e-09
GAT2_YEAST (P40209) GAT2 protein 56 7e-08
WC2_NEUCR (P78714) White collar 2 protein (WC2) 55 9e-08
WC1_NEUCR (Q01371) White collar 1 protein (WC1) 55 9e-08
GAT3_YEAST (Q07928) GAT3 protein 52 7e-07
GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3 46 7e-05
AREA_GIBFU (P78688) Nitrogen regulatory protein areA 45 1e-04
URB1_USTMA (P40349) Siderophore biosynthesis regulatory protein ... 44 3e-04
AREA_ASPNG (O13412) Nitrogen regulatory protein areA 44 3e-04
GAT4_YEAST (P40569) GAT4 protein 44 3e-04
SRD1_YEAST (P09007) Pre-rRNA processing protein SRD1 (Suppressor... 43 4e-04
GAF1_SCHPO (Q10280) Transcription factor gaf1 (Gaf-1) 43 6e-04
GATC_DROME (P91623) GATA-binding factor-C (Transcription factor ... 42 0.001
GAT1_RAT (P43429) Erythroid transcription factor (GATA-1) (Eryf1... 40 0.003
GAT1_MOUSE (P17679) Erythroid transcription factor (GATA-1) (Ery... 40 0.003
GAT1_HUMAN (P15976) Erythroid transcription factor (GATA-1) (Ery... 40 0.003
GA1A_XENLA (P23767) GATA binding factor-1A (Transcription factor... 40 0.003
AREA_EMENI (P17429) Nitrogen regulatory protein areA 40 0.003
NUT1_MAGGR (Q01168) Nitrogen regulatory protein NUT1 40 0.004
GA1B_XENLA (P23768) GATA binding factor-1B (Transcription factor... 40 0.004
>CGPB_FUSSO (Q00858) Cutinase gene palindrome-binding protein (PBP)
Length = 457
Score = 60.5 bits (145), Expect = 3e-09
Identities = 38/137 (27%), Positives = 59/137 (42%), Gaps = 15/137 (10%)
Query: 19 LPKSNSSPTCEKTTVRRTRSKRPRLATF--SSHHSTMQLISSTSSFVGENMQDSVISNKG 76
L KS + E R S ++A + +SH T+++++ GE + N
Sbjct: 308 LNKSLTRENLEGAAGSRPDSLNDKMARYEGASHTETIEMLTGLRYIEGERSRGITTGNAS 367
Query: 77 ASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWR 136
+ K ++ +G+ KK KT + CT C +P+WR
Sbjct: 368 PTLIKGDAGIAIPLERDPRTGDKKKKLKTSEEYV-------------CTDCGTLDSPEWR 414
Query: 137 TGPEGPKTLCNACGVRY 153
GP GPKTLCNACG+R+
Sbjct: 415 KGPSGPKTLCNACGLRW 431
>GAT2_YEAST (P40209) GAT2 protein
Length = 560
Score = 55.8 bits (133), Expect = 7e-08
Identities = 34/96 (35%), Positives = 50/96 (51%), Gaps = 12/96 (12%)
Query: 116 NALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRY---------KSGRLCPEYRPA- 165
NA ++ C HC T+TP+WR GP G +TLCNACG+ Y KS L YR +
Sbjct: 464 NAKKIIEFCFHCGETETPEWRKGPYGTRTLCNACGLFYRKVTKKFGSKSSNLLLRYRRSI 523
Query: 166 --ASSTFSPDLHSNSHKKILEMRVMRRKDNKNSGIL 199
A+ PD + ++ I +M + D++ + IL
Sbjct: 524 DLANDRRIPDFITIPNRFIHDMDNDQTLDSEYNTIL 559
>WC2_NEUCR (P78714) White collar 2 protein (WC2)
Length = 530
Score = 55.5 bits (132), Expect = 9e-08
Identities = 40/147 (27%), Positives = 61/147 (41%), Gaps = 21/147 (14%)
Query: 8 PSSSVNKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHS-TMQLISSTSSFVGEN 66
P+SS+N L + N E R S R ++ + +H+ T+++++ GE
Sbjct: 371 PASSLN---IALTREN----LEGIAGSRPDSIREKMLRYEGNHADTIEMLTGLKYQEGER 423
Query: 67 MQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTH 126
N + K + +GE KK K + CT
Sbjct: 424 SHGITTGNASPTLIKGDAGIAIPLDRDPRTGEKKKKIKVAEEYV-------------CTD 470
Query: 127 CEATKTPQWRTGPEGPKTLCNACGVRY 153
C +P+WR GP GPKTLCNACG+R+
Sbjct: 471 CGTLDSPEWRKGPSGPKTLCNACGLRW 497
>WC1_NEUCR (Q01371) White collar 1 protein (WC1)
Length = 1167
Score = 55.5 bits (132), Expect = 9e-08
Identities = 27/72 (37%), Positives = 39/72 (53%), Gaps = 4/72 (5%)
Query: 120 LVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRY--KSGRLCP--EYRPAASSTFSPDLH 175
+VR C +C TP+WR GP G + LCN+CG+R+ ++GR+ P R + S +
Sbjct: 930 MVRDCANCHTRNTPEWRRGPSGNRDLCNSCGLRWAKQTGRVSPRTSSRGGNGDSMSKKSN 989
Query: 176 SNSHKKILEMRV 187
S SH L V
Sbjct: 990 SPSHSSPLHREV 1001
>GAT3_YEAST (Q07928) GAT3 protein
Length = 141
Score = 52.4 bits (124), Expect = 7e-07
Identities = 24/48 (50%), Positives = 30/48 (62%), Gaps = 4/48 (8%)
Query: 111 AALDHNAL---GLVRQCTHCEATKT-PQWRTGPEGPKTLCNACGVRYK 154
A + HN G+ R+C C KT PQWR GP+G TLCNACG+ Y+
Sbjct: 56 AVVQHNVQKRKGVTRRCPQCAVIKTSPQWREGPDGEVTLCNACGLFYR 103
>GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3
Length = 730
Score = 45.8 bits (107), Expect = 7e-05
Identities = 43/151 (28%), Positives = 65/151 (42%), Gaps = 24/151 (15%)
Query: 5 EQHPSSSVNKEDFVLPKSNSSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISS-TSSFV 63
+ H S ++ K L +NSS + T + F ++ L SS T++ V
Sbjct: 208 QSHHSFNIYK----LQNNNSSSSAMNITNNNNSNNSNIQHPFLKKSDSIGLSSSNTTNSV 263
Query: 64 GENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQ 123
+N +S+ + K S ++ SG++KK PL+ Q
Sbjct: 264 RKNSLIKPMSSTSLANFKRAASVSSSISNMEPSGQNKK------PLI------------Q 305
Query: 124 CTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
C +C+ KTP WR PEG TLCNACG+ K
Sbjct: 306 CFNCKTFKTPLWRRSPEG-NTLCNACGLFQK 335
>AREA_GIBFU (P78688) Nitrogen regulatory protein areA
Length = 971
Score = 45.1 bits (105), Expect = 1e-04
Identities = 38/122 (31%), Positives = 52/122 (42%), Gaps = 8/122 (6%)
Query: 33 VRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQ 92
+RR K PR A+ H Q + + ++MQ S + + F S +A +
Sbjct: 610 LRRQHPKLPRNASTPVHFGGQQ---NGFEQLAQSMQSSPAGDGNGTMSGF--SSVAPSRP 664
Query: 93 KLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVR 152
SS K T AA + N CT+C TP WR PEG + LCNACG+
Sbjct: 665 --SSPPMSKQGSTTNLQAAAGNGNDGNAPTTCTNCFTQTTPLWRRNPEG-QPLCNACGLF 721
Query: 153 YK 154
K
Sbjct: 722 LK 723
>URB1_USTMA (P40349) Siderophore biosynthesis regulatory protein
URBS1
Length = 950
Score = 43.9 bits (102), Expect = 3e-04
Identities = 26/77 (33%), Positives = 39/77 (49%), Gaps = 6/77 (7%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKI 182
+C++C T TP WR P+G T+CNACG+ KS +R A++ D +H+
Sbjct: 337 RCSNCGVTSTPLWRRAPDG-STICNACGLYIKSH---STHRSASNRLSGSDASPPTHEAK 392
Query: 183 LEMR--VMRRKDNKNSG 197
L R+D+ SG
Sbjct: 393 LAAAGPSCSREDDPKSG 409
Score = 42.0 bits (97), Expect = 0.001
Identities = 18/41 (43%), Positives = 27/41 (64%), Gaps = 2/41 (4%)
Query: 114 DHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
D + +G +R CT+C+ T TP WR +G +CNACG+ +K
Sbjct: 473 DKSVVGALR-CTNCQTTTTPLWRRDEDG-NNICNACGLYHK 511
>AREA_ASPNG (O13412) Nitrogen regulatory protein areA
Length = 882
Score = 43.9 bits (102), Expect = 3e-04
Identities = 33/131 (25%), Positives = 54/131 (41%), Gaps = 22/131 (16%)
Query: 24 SSPTCEKTTVRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFP 83
S+ + + R +R ++A +S +T QL+ + + + + G S+
Sbjct: 597 SAASVSEVRNREQDPRRQKIARTTSTPNTAQLLRQSMNANTSHTSPNTPPESGLSS---- 652
Query: 84 DSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPK 143
A + S G SK +++ P CT+C TP WR PEG +
Sbjct: 653 ----AVPSRPASPGGSKNGEQSSGPTT-------------CTNCFTQTTPLWRRNPEG-Q 694
Query: 144 TLCNACGVRYK 154
LCNACG+ K
Sbjct: 695 PLCNACGLFLK 705
>GAT4_YEAST (P40569) GAT4 protein
Length = 121
Score = 43.5 bits (101), Expect = 3e-04
Identities = 22/64 (34%), Positives = 32/64 (49%), Gaps = 3/64 (4%)
Query: 119 GLVRQCTHCEATKTP-QWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSN 177
G+ R C C KT QWR GP G LCNACG+ ++ +L + AA+ + +
Sbjct: 48 GITRTCGQCGEIKTSLQWREGPNGAACLCNACGLFFR--KLILRFGRAAAKRYMEQIKGT 105
Query: 178 SHKK 181
K+
Sbjct: 106 GTKR 109
>SRD1_YEAST (P09007) Pre-rRNA processing protein SRD1 (Suppressor of
rRNA processing defect protein 1)
Length = 221
Score = 43.1 bits (100), Expect = 4e-04
Identities = 15/35 (42%), Positives = 25/35 (70%)
Query: 119 GLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRY 153
G +++C+ C+ T T QWR+GP+ + LC+ CG+ Y
Sbjct: 163 GEIKECSKCKDTWTIQWRSGPDQNRELCSPCGLAY 197
>GAF1_SCHPO (Q10280) Transcription factor gaf1 (Gaf-1)
Length = 855
Score = 42.7 bits (99), Expect = 6e-04
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 10/91 (10%)
Query: 73 SNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDH---------NALGLVRQ 123
+N A+ + + + KK + S KN + AA D NA
Sbjct: 575 TNTAATPQAALPTTLDTKKDRSVSFNINKNAEKPTVSNAAEDKKGDANTRRANATNPTPT 634
Query: 124 CTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
CT+C+ TP WR P+G + LCNACG+ K
Sbjct: 635 CTNCQTRTTPLWRRSPDG-QPLCNACGLFMK 664
>GATC_DROME (P91623) GATA-binding factor-C (Transcription factor
GATA-C) (dGATA-C) (Grain protein)
Length = 486
Score = 41.6 bits (96), Expect = 0.001
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ YK +G+ P +P
Sbjct: 259 RECVNCGATSTPLWRRDGTG-HYLCNACGLYYKMNGQNRPLIKP 301
Score = 36.6 bits (83), Expect = 0.041
Identities = 21/60 (35%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Query: 96 SGESKKNKKTKAPL-LAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
+G+++ K K L L +L A C +C+ T T WR G + +CNACG+ YK
Sbjct: 292 NGQNRPLIKPKRRLTLQSLQSAAKRAGTSCANCKTTTTTLWRRNASG-EPVCNACGLYYK 350
>GAT1_RAT (P43429) Erythroid transcription factor (GATA-1) (Eryf1)
(GF-1) (NF-E1)
Length = 413
Score = 40.4 bits (93), Expect = 0.003
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ +K +G+ P RP
Sbjct: 202 RECVNCGATATPLWRRDRTG-HYLCNACGLYHKMNGQNRPLIRP 244
Score = 38.9 bits (89), Expect = 0.008
Identities = 16/32 (50%), Positives = 20/32 (62%), Gaps = 1/32 (3%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
QCT+C+ T T WR G +CNACG+ YK
Sbjct: 257 QCTNCQTTTTTLWRRNASG-DPVCNACGLYYK 287
>GAT1_MOUSE (P17679) Erythroid transcription factor (GATA-1) (Eryf1)
(GF-1) (NF-E1)
Length = 413
Score = 40.4 bits (93), Expect = 0.003
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ +K +G+ P RP
Sbjct: 202 RECVNCGATATPLWRRDRTG-HYLCNACGLYHKMNGQNRPLIRP 244
Score = 37.4 bits (85), Expect = 0.024
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
QCT+C+ T T WR G +CNACG+ +K
Sbjct: 257 QCTNCQTTTTTLWRRNASG-DPVCNACGLYFK 287
>GAT1_HUMAN (P15976) Erythroid transcription factor (GATA-1) (Eryf1)
(GF-1) (NF-E1)
Length = 413
Score = 40.4 bits (93), Expect = 0.003
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ +K +G+ P RP
Sbjct: 202 RECVNCGATATPLWRRDRTG-HYLCNACGLYHKMNGQNRPLIRP 244
Score = 38.9 bits (89), Expect = 0.008
Identities = 16/32 (50%), Positives = 20/32 (62%), Gaps = 1/32 (3%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
QCT+C+ T T WR G +CNACG+ YK
Sbjct: 257 QCTNCQTTTTTLWRRNASG-DPVCNACGLYYK 287
>GA1A_XENLA (P23767) GATA binding factor-1A (Transcription factor
xGATA-1A)
Length = 359
Score = 40.4 bits (93), Expect = 0.003
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ +K +G+ P RP
Sbjct: 176 RECVNCGATVTPLWRRDMSG-HYLCNACGLYHKMNGQNRPLIRP 218
Score = 35.0 bits (79), Expect = 0.12
Identities = 14/32 (43%), Positives = 19/32 (58%), Gaps = 1/32 (3%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
QC++C + T WR G +CNACG+ YK
Sbjct: 231 QCSNCHTSTTTLWRRNASG-DPVCNACGLYYK 261
>AREA_EMENI (P17429) Nitrogen regulatory protein areA
Length = 876
Score = 40.4 bits (93), Expect = 0.003
Identities = 35/120 (29%), Positives = 51/120 (42%), Gaps = 20/120 (16%)
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKL 94
R R + PR + ST +T+ + ++MQ+ S+ +T AA +
Sbjct: 603 RNRDQDPRRQKIARTSST----PNTAQLLRQSMQNQS-SHTSPNTPPESGLNSAAPSRPA 657
Query: 95 SSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
S G +K N + P CT+C TP WR PEG + LCNACG+ K
Sbjct: 658 SPGGTK-NGEQNGPTT-------------CTNCFTQTTPLWRRNPEG-QPLCNACGLFLK 702
>NUT1_MAGGR (Q01168) Nitrogen regulatory protein NUT1
Length = 956
Score = 40.0 bits (92), Expect = 0.004
Identities = 17/31 (54%), Positives = 20/31 (63%), Gaps = 1/31 (3%)
Query: 124 CTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
CT+C TP WR PEG + LCNACG+ K
Sbjct: 663 CTNCATQTTPLWRRNPEG-QPLCNACGLFLK 692
>GA1B_XENLA (P23768) GATA binding factor-1B (Transcription factor
xGATA-1B)
Length = 364
Score = 40.0 bits (92), Expect = 0.004
Identities = 21/44 (47%), Positives = 27/44 (60%), Gaps = 2/44 (4%)
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYK-SGRLCPEYRP 164
R+C +C AT TP WR G LCNACG+ +K +G+ P RP
Sbjct: 178 RECVNCGATVTPLWRRDLSG-HYLCNACGLYHKMNGQNRPLIRP 220
Score = 34.3 bits (77), Expect = 0.20
Identities = 14/32 (43%), Positives = 19/32 (58%), Gaps = 1/32 (3%)
Query: 123 QCTHCEATKTPQWRTGPEGPKTLCNACGVRYK 154
QC++C + T WR G +CNACG+ YK
Sbjct: 233 QCSNCHTSTTTLWRRNAGGDP-VCNACGLYYK 263
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.124 0.353
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,582,114
Number of Sequences: 164201
Number of extensions: 852215
Number of successful extensions: 2274
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 21
Number of HSP's successfully gapped in prelim test: 78
Number of HSP's that attempted gapping in prelim test: 2153
Number of HSP's gapped (non-prelim): 147
length of query: 204
length of database: 59,974,054
effective HSP length: 105
effective length of query: 99
effective length of database: 42,732,949
effective search space: 4230561951
effective search space used: 4230561951
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146329.15


