

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.11 + phase: 0
(327 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
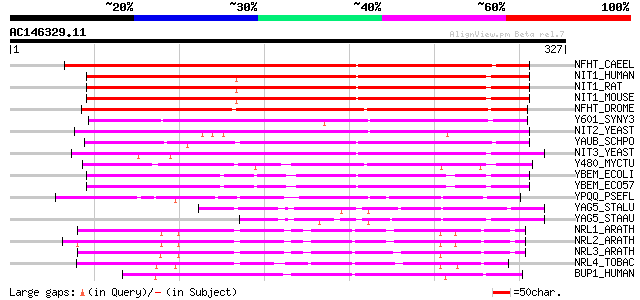
Sequences producing significant alignments: (bits) Value
NFHT_CAEEL (O76463) Nitrilase and fragile histidine triad fusion... 260 3e-69
NIT1_HUMAN (Q86X76) Nitrilase homolog 1 (EC 3.5.-.-) 257 3e-68
NIT1_RAT (Q7TQ94) Nitrilase homolog 1 (EC 3.5.-.-) 253 4e-67
NIT1_MOUSE (Q8VDK1) Nitrilase homolog 1 (EC 3.5.-.-) 253 4e-67
NFHT_DROME (O76464) Nitrilase and fragile histidine triad fusion... 215 1e-55
Y601_SYNY3 (P55175) Hypothetical UPF0012 protein sll0601 (EC 3.5... 182 1e-45
NIT2_YEAST (P47016) Probable hydrolase NIT2 (EC 3.5.-.-) 179 1e-44
YAUB_SCHPO (Q10166) Hypothetical UPF0012 protein C26A3.11 in chr... 165 1e-40
NIT3_YEAST (P49954) Probable hydrolase NIT3 (EC 3.5.-.-) 147 4e-35
Y480_MYCTU (Q11146) Hypothetical UPF0012 protein Rv0480c/MT0498 ... 112 1e-24
YBEM_ECOLI (P39874) Hypothetical UPF0012 protein ybeM (EC 3.5.-.-) 108 2e-23
YBEM_ECO57 (P58054) Hypothetical UPF0012 protein ybeM (EC 3.5.-.-) 106 9e-23
YPQQ_PSEFL (P55176) Hypothetical UPF0012 protein in pqqF 5'regio... 86 1e-16
YAG5_STALU (P55178) Hypothetical UPF0012 protein in agr operon (... 84 5e-16
YAG5_STAAU (P55177) Hypothetical UPF0012 protein in agr operon (... 80 5e-15
NRL1_ARATH (P32961) Nitrilase 1 (EC 3.5.5.1) 80 5e-15
NRL2_ARATH (P32962) Nitrilase 2 (EC 3.5.5.1) 79 2e-14
NRL3_ARATH (P46010) Nitrilase 3 (EC 3.5.5.1) 78 3e-14
NRL4_TOBAC (Q42965) Nitrilase 4 (EC 3.5.5.1) 74 7e-13
BUP1_HUMAN (Q9UBR1) Beta-ureidopropionase (EC 3.5.1.6) (Beta-ala... 70 6e-12
>NFHT_CAEEL (O76463) Nitrilase and fragile histidine triad fusion
protein NitFhit [Includes:
Bis(5'-adenosyl)-triphosphatase (EC 3.6.1.29)
(Diadenosine 5',5'''-P1,P3-triphosphate hydrolase)
(Dinucleosidetriphosphatase) (AP3A hydrolase) (AP3AASE);
Nitrilas
Length = 440
Score = 260 bits (665), Expect = 3e-69
Identities = 130/276 (47%), Positives = 180/276 (65%), Gaps = 5/276 (1%)
Query: 33 TTGESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKD 92
T TMAT +A QMTS DL NF +++ A +++ PE F F+G
Sbjct: 4 TVFRRTMATGRHFIAVCQMTSDNDLEKNFQAAKNMIERAGEKKCEMVFLPECFDFIGLNK 63
Query: 93 GDSVSIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKG-SDPRHLFNTHVVVDDTGKIQT 151
+ + +A D M++Y LAR+ +IWLSLGG K SD H +NTH+++D G +
Sbjct: 64 NEQIDLAMATDCEYMEKYRELARKHNIWLSLGGLHHKDPSDAAHPWNTHLIIDSDGVTRA 123
Query: 152 TYRKIHLFDVDVPGGRVYKESNFTESGKDIVA-VDSPIGRLGLSVCYDLRFPELYQLLRF 210
Y K+HLFD+++PG ES F+++G +++ VD+PIGRLGLS+CYD+RFPEL L
Sbjct: 124 EYNKLHLFDLEIPGKVRLMESEFSKAGTEMIPPVDTPIGRLGLSICYDVRFPEL-SLWNR 182
Query: 211 QHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIID 270
+ GAQ+L P+AFT TG AHWE LLRARAIENQCYV+AAAQ G HN KR+SYG ++++D
Sbjct: 183 KRGAQLLSFPSAFTLNTGLAHWETLLRARAIENQCYVVAAAQTGAHNPKRQSYGHSMVVD 242
Query: 271 PWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPI 306
PWG VV + +R+ + A+IDLS VD++RE P+
Sbjct: 243 PWGAVVAQCSERVD--MCFAEIDLSYVDTLREMQPV 276
>NIT1_HUMAN (Q86X76) Nitrilase homolog 1 (EC 3.5.-.-)
Length = 327
Score = 257 bits (656), Expect = 3e-68
Identities = 127/265 (47%), Positives = 178/265 (66%), Gaps = 7/265 (2%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGP 105
VA Q+TS D NF TC+ LV+EAA GA L PEAF F+ +++ +++PL G
Sbjct: 49 VAVCQVTSTPDKQQNFKTCAELVREAARLGACLAFLPEAFDFIARDPAETLHLSEPLGGK 108
Query: 106 IMDQYCSLARESSIWLSLGGFQEKGSD---PRHLFNTHVVVDDTGKIQTTYRKIHLFDVD 162
++++Y LARE +WLSLGGF E+G D + ++N HV+++ G + TYRK HL DV+
Sbjct: 109 LLEEYTQLARECGLWLSLGGFHERGQDWEQTQKIYNCHVLLNSKGAVVATYRKTHLCDVE 168
Query: 163 VPGGRVYKESNFTESGKDIVA-VDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPA 221
+PG ESN T G + + V +P G++GL+VCYD+RFPEL L Q GA+IL P+
Sbjct: 169 IPGQGPMCESNSTMPGPSLESPVSTPAGKIGLAVCYDMRFPEL-SLALAQAGAEILTYPS 227
Query: 222 AFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPD 281
AF +TG AHWE+LLRARAIE QCYV+AAAQ G H++KR SYG ++++DPWGTVV R +
Sbjct: 228 AFGSITGPAHWEVLLRARAIETQCYVVAAAQCGRHHEKRASYGHSMVVDPWGTVVARCSE 287
Query: 282 RLSTGIVVADIDLSLVDSVREKMPI 306
G+ +A IDL+ + +R +P+
Sbjct: 288 --GPGLCLARIDLNYLRQLRRHLPV 310
>NIT1_RAT (Q7TQ94) Nitrilase homolog 1 (EC 3.5.-.-)
Length = 292
Score = 253 bits (647), Expect = 4e-67
Identities = 126/265 (47%), Positives = 177/265 (66%), Gaps = 7/265 (2%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGP 105
VA Q+TS + NF TC+ LV+EA GA L PEAF F+ +++ +++PLDG
Sbjct: 14 VAVCQVTSTPNKQENFKTCAELVQEATRLGACLAFLPEAFDFIARNPAETLLLSEPLDGD 73
Query: 106 IMDQYCSLARESSIWLSLGGFQEKGSD---PRHLFNTHVVVDDTGKIQTTYRKIHLFDVD 162
++ QY LARE IWLSLGGF E+G D + ++N HV+++ G + +YRK HL DV+
Sbjct: 74 LLGQYSQLARECGIWLSLGGFHERGQDWEQTQKIYNCHVLLNSKGSVVASYRKTHLCDVE 133
Query: 163 VPGGRVYKESNFTESGKDIVA-VDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPA 221
+PG +ESN+T G + V +P G++GL++CYD+RFPEL L Q GA+IL P+
Sbjct: 134 IPGQGPMRESNYTMPGYALEPPVKTPAGKVGLAICYDMRFPEL-SLKLAQAGAEILTYPS 192
Query: 222 AFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPD 281
AF VTG AHWE+LLRARAIE+QCYVIAAAQ G H++ R SYG ++++DPWGTVV +
Sbjct: 193 AFGSVTGPAHWEVLLRARAIESQCYVIAAAQCGRHHETRASYGHSMVVDPWGTVVASCSE 252
Query: 282 RLSTGIVVADIDLSLVDSVREKMPI 306
G+ +A IDL + +R+ +P+
Sbjct: 253 --GPGLCLARIDLHFLQQMRQHLPV 275
>NIT1_MOUSE (Q8VDK1) Nitrilase homolog 1 (EC 3.5.-.-)
Length = 323
Score = 253 bits (647), Expect = 4e-67
Identities = 126/265 (47%), Positives = 179/265 (67%), Gaps = 7/265 (2%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGP 105
VA Q+TS + NF TC+ LV+EAA GA L PEAF F+ +++ +++PL+G
Sbjct: 45 VAVCQVTSTPNKQENFKTCAELVQEAARLGACLAFLPEAFDFIARNPAETLLLSEPLNGD 104
Query: 106 IMDQYCSLARESSIWLSLGGFQEKGSD---PRHLFNTHVVVDDTGKIQTTYRKIHLFDVD 162
++ QY LARE IWLSLGGF E+G D + ++N HV+++ G + +YRK HL DV+
Sbjct: 105 LLGQYSQLARECGIWLSLGGFHERGQDWEQNQKIYNCHVLLNSKGSVVASYRKTHLCDVE 164
Query: 163 VPGGRVYKESNFTESGKDIVA-VDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPA 221
+PG +ESN+T+ G + V +P G++GL++CYD+RFPEL L Q GA+IL +
Sbjct: 165 IPGQGPMRESNYTKPGGTLEPPVKTPAGKVGLAICYDMRFPEL-SLKLAQAGAEILTYSS 223
Query: 222 AFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPD 281
AF VTG AHWE+LLRARAIE+QCYVIAAAQ G H++ R SYG ++++DPWGTVV R +
Sbjct: 224 AFGSVTGPAHWEVLLRARAIESQCYVIAAAQCGRHHETRASYGHSMVVDPWGTVVARCSE 283
Query: 282 RLSTGIVVADIDLSLVDSVREKMPI 306
G+ +A IDL + +R+ +P+
Sbjct: 284 --GPGLCLARIDLHFLQQMRQHLPV 306
>NFHT_DROME (O76464) Nitrilase and fragile histidine triad fusion
protein NitFhit (NFT-1 protein) [Includes:
Bis(5'-adenosyl)-triphosphatase (EC 3.6.1.29)
(Diadenosine 5',5'''-P1,P3-triphosphate hydrolase)
(Dinucleosidetriphosphatase) (AP3A hydrolase) (AP
Length = 460
Score = 215 bits (548), Expect = 1e-55
Identities = 113/263 (42%), Positives = 164/263 (61%), Gaps = 4/263 (1%)
Query: 43 SVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPL 102
S +A QM S +D A+N S LV A S A +L PE FVG ++ +++ L
Sbjct: 32 SATIAVGQMRSTSDKAANLSQVIELVDRAKSQNACMLFLPECCDFVGESRTQTIELSEGL 91
Query: 103 DGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVD 162
DG +M QY LA+ + IW+SLGG E+ + +FN HV++++ G++ YRK+H+FDV
Sbjct: 92 DGELMAQYRELAKCNKIWISLGGVHERND--QKIFNAHVLLNEKGELAAVYRKLHMFDVT 149
Query: 163 VPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAA 222
R+ + T V +P+G++GL +CYDLRF E LLR + GA +L P+A
Sbjct: 150 TKEVRLRESDTVTPGYCLERPVSTPVGQIGLQICYDLRFAEPAVLLR-KLGANLLTYPSA 208
Query: 223 FTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDR 282
FT TG+AHWEILLRARAIE QC+V+AAAQ G HN KR+S+G ++I+ PWG V+ ++
Sbjct: 209 FTYATGKAHWEILLRARAIETQCFVVAAAQIGWHNQKRQSWGHSMIVSPWGNVLADCSEQ 268
Query: 283 LSTGIVVADIDLSLVDSVREKMP 305
I A++DLS++ S+ + MP
Sbjct: 269 -ELDIGTAEVDLSVLQSLYQTMP 290
>Y601_SYNY3 (P55175) Hypothetical UPF0012 protein sll0601 (EC
3.5.-.-)
Length = 272
Score = 182 bits (461), Expect = 1e-45
Identities = 102/262 (38%), Positives = 150/262 (56%), Gaps = 7/262 (2%)
Query: 47 AAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPI 106
AA QMTS +L N L+ A GA+L+ PE F+F+G + + + A +
Sbjct: 7 AALQMTSRPNLTENLQEAEELIDLAVRQGAELVGLPENFAFLG-NETEKLEQATAIATAT 65
Query: 107 MDQYCSLARESSIWLSLGGFQ-EKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPG 165
++A+ + + GGF + +NT ++ G+ Y K+HLFDV+VP
Sbjct: 66 EKFLQTMAQRFQVTILAGGFPFPVAGEAGKAYNTATLIAPNGQELARYHKVHLFDVNVPD 125
Query: 166 GRVYKESNFTESGKDIVAV--DSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAF 223
G Y ES +G+ V G LGLS+CYD+RFPELY+ L Q GA +L VPAAF
Sbjct: 126 GNTYWESATVMAGQKYPPVYHSDSFGNLGLSICYDVRFPELYRYLSRQ-GADVLFVPAAF 184
Query: 224 TKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRL 283
T TG+ HW++LL+ARAIEN CYVIA AQ G H ++R ++G +IIDPWG ++ ++
Sbjct: 185 TAYTGKDHWQVLLQARAIENTCYVIAPAQTGCHYERRHTHGHAMIIDPWGVILADAGEK- 243
Query: 284 STGIVVADIDLSLVDSVREKMP 305
G+ +A+I+ + VR++MP
Sbjct: 244 -PGLAIAEINPDRLKQVRQQMP 264
>NIT2_YEAST (P47016) Probable hydrolase NIT2 (EC 3.5.-.-)
Length = 307
Score = 179 bits (453), Expect = 1e-44
Identities = 110/298 (36%), Positives = 159/298 (52%), Gaps = 31/298 (10%)
Query: 39 MATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSI 98
M + RVA AQ+ S DL N L+ EA A ++ PEA ++ S +
Sbjct: 1 MTSKLKRVAVAQLCSSADLTKNLKVVKELISEAIQKKADVVFLPEASDYLSQNPLHSRYL 60
Query: 99 AQPLDGPIMDQYCS---LARESS--IWLSLG-----GFQEKGSDPRHLFNTHVVVDDTGK 148
AQ I S L R++S I +S+G Q+ + N + +D GK
Sbjct: 61 AQKSPKFIRQLQSSITDLVRDNSRNIDVSIGVHLPPSEQDLLEGNDRVRNVLLYIDHEGK 120
Query: 149 IQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIV-AVDSPIGRLGLSVCYDLRFPELYQL 207
I Y+K+HLFDVDVP G + KES + GK I ++SP+G+LG ++CYD+RFPE
Sbjct: 121 ILQEYQKLHLFDVDVPNGPILKESKSVQPGKAIPDIIESPLGKLGSAICYDIRFPEFSLK 180
Query: 208 LRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTH----------- 256
LR GA+IL P+AFT TGEAHWE+L RARA++ QCYV+ Q G H
Sbjct: 181 LRSM-GAEILCFPSAFTIKTGEAHWELLGRARAVDTQCYVLMPGQVGMHDLSDPEWEKQS 239
Query: 257 -------NDKRESYGDTLIIDPWGTVVGRL-PDRLSTGIVVADIDLSLVDSVREKMPI 306
+ +RES+G +++IDPWG ++ P + +++AD+D L+ +R KMP+
Sbjct: 240 HMSALEKSSRRESWGHSMVIDPWGKIIAHADPSTVGPQLILADLDRELLQEIRNKMPL 297
>YAUB_SCHPO (Q10166) Hypothetical UPF0012 protein C26A3.11 in
chromosome I (EC 3.5.-.-)
Length = 322
Score = 165 bits (418), Expect = 1e-40
Identities = 91/264 (34%), Positives = 147/264 (55%), Gaps = 8/264 (3%)
Query: 45 RVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLD- 103
R+ Q+ + D + N V EAA G+ ++ PE F+ G A+P++
Sbjct: 45 RIGLVQLANTKDKSENLQLARLKVLEAAKNGSNVIVLPEIFNSPYGT-GYFNQYAEPIEE 103
Query: 104 -GPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVD 162
P S+A+++ +L G E+ L+NT +V D +GK+ +RKIHLFD+D
Sbjct: 104 SSPSYQALSSMAKDTKTYLFGGSIPERKDGK--LYNTAMVFDPSGKLIAVHRKIHLFDID 161
Query: 163 VPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAA 222
+PGG ++ES+ G + VD+ G+ GL +CYD+RFPEL ++ ++G +++ P A
Sbjct: 162 IPGGVSFRESDSLSPGDAMTMVDTEYGKFGLGICYDIRFPEL-AMIAARNGCSVMIYPGA 220
Query: 223 FTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDR 282
F TG HWE+L RARA++N+ +V A A N S+G + ++DP+G V+ ++
Sbjct: 221 FNLSTGPLHWELLARARAVDNEMFVACCAPARDMNADYHSWGHSTVVDPFGKVIATTDEK 280
Query: 283 LSTGIVVADIDLSLVDSVREKMPI 306
S IV ADID S++ + R +PI
Sbjct: 281 PS--IVYADIDPSVMSTARNSVPI 302
>NIT3_YEAST (P49954) Probable hydrolase NIT3 (EC 3.5.-.-)
Length = 291
Score = 147 bits (371), Expect = 4e-35
Identities = 86/285 (30%), Positives = 149/285 (52%), Gaps = 9/285 (3%)
Query: 37 STMATNSVRVAAAQMT-SITDLASNFSTCSRLVKEAASA--GAKLLCFPEAFSFVGAKDG 93
S + + ++VA Q++ S D +N + ++ A KL+ PE F+ + D
Sbjct: 4 SKILSQKIKVALVQLSGSSPDKMANLQRAATFIERAMKEQPDTKLVVLPECFNSPYSTDQ 63
Query: 94 --DSVSIAQPLDGPIMDQYCS-LARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQ 150
+ P + Q+ S LA + I L G E ++NT ++ ++ GK+
Sbjct: 64 FRKYSEVINPKEPSTSVQFLSNLANKFKIILVGGTIPELDPKTDKIYNTSIIFNEDGKLI 123
Query: 151 TTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRF 210
+RK+HLFDVD+P G + ES G+ +D+ G+ G+ +CYD+RFPEL +L
Sbjct: 124 DKHRKVHLFDVDIPNGISFHESETLSPGEKSTTIDTKYGKFGVGICYDMRFPEL-AMLSA 182
Query: 211 QHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIID 270
+ GA ++ P+AF VTG HW +L R+RA++NQ YV+ + A +YG ++++D
Sbjct: 183 RKGAFAMIYPSAFNTVTGPLHWHLLARSRAVDNQVYVMLCSPARNLQSSYHAYGHSIVVD 242
Query: 271 PWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPIAKVIVCNVY 315
P G +V + I+ A++D +++S R+ +P+ K +VY
Sbjct: 243 PRGKIVAEAGE--GEEIIYAELDPEVIESFRQAVPLTKQRRFDVY 285
>Y480_MYCTU (Q11146) Hypothetical UPF0012 protein Rv0480c/MT0498 (EC
3.5.-.-)
Length = 340
Score = 112 bits (281), Expect = 1e-24
Identities = 92/281 (32%), Positives = 141/281 (49%), Gaps = 32/281 (11%)
Query: 44 VRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSV-SIAQPL 102
+R+A AQ+ S TD A+N + EAA+AGA+L+ FPEA + G + +A+P+
Sbjct: 61 MRIALAQIRSGTDPAANLQLVGKYAGEAATAGAQLVVFPEA---TMCRLGVPLRQVAEPV 117
Query: 103 DGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVV--DDTGKIQTTYRKIHLFD 160
DGP + +A E+ I + G F G + NT + + Y KIHL+D
Sbjct: 118 DGPWANGVRRIATEAGITVIAGMFTPTGDG--RVTNTLIAAGPGTPNQPDAHYHKIHLYD 175
Query: 161 VDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVP 220
+ ES G++ V V R+GL+VCYD+RFP LY L + GAQ++ V
Sbjct: 176 -----AFGFTESRTVAPGREPVVVVVDGVRVGLTVCYDIRFPALYTELA-RRGAQLIAVC 229
Query: 221 AAFTKVTGE-AHWEILLRARAIENQCYVIAAAQA---------GTHNDKRESYGDTLIID 270
A++ G+ W +L RARA+++ YV AA QA G + G +L+
Sbjct: 230 ASWGSGPGKLEQWTLLARARALDSMSYVAAAGQADPGDARTGVGASSAAPTGVGGSLVAS 289
Query: 271 PWGTVV---GRLPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
P G VV G P ++VADID+ V + R+++ + +
Sbjct: 290 PLGEVVVSAGTQPQ-----LLVADIDVDNVAAARDRIAVLR 325
>YBEM_ECOLI (P39874) Hypothetical UPF0012 protein ybeM (EC 3.5.-.-)
Length = 262
Score = 108 bits (269), Expect = 2e-23
Identities = 83/264 (31%), Positives = 128/264 (48%), Gaps = 19/264 (7%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGD-SVSIAQPLDG 104
VAA Q + N C+ L+ +AA A L PEA D D SV AQ L+G
Sbjct: 3 VAAGQFAVTSVWEKNAEICASLMAQAAENDASLFALPEALLARDDHDADLSVKSAQLLEG 62
Query: 105 PIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVP 164
+ + ++ + + L S P +N V + G I Y K+HL+D
Sbjct: 63 EFLGRLRRESKRNMMTTILT--IHVPSTPGRAWNMLVALQ-AGNIVARYAKLHLYDAFA- 118
Query: 165 GGRVYKESNFTESGKDIVAVDSPIG-RLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAF 223
+ES ++G +I + G ++GL CYDLRFPEL L + GA+IL++PAA+
Sbjct: 119 ----IQESRRVDAGNEIAPLLEVEGMKVGLMTCYDLRFPEL-ALAQALQGAEILVLPAAW 173
Query: 224 TK-VTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDR 282
+ E HW LL ARA++ CY++AA + G N G + IIDP+G + +
Sbjct: 174 VRGPLKEHHWSTLLAARALDTTCYMVAAGECGNKN-----IGQSRIIDPFGVTIAAASE- 227
Query: 283 LSTGIVVADIDLSLVDSVREKMPI 306
+++A++ V VR ++P+
Sbjct: 228 -MPALIMAEVTPERVRQVRAQLPV 250
>YBEM_ECO57 (P58054) Hypothetical UPF0012 protein ybeM (EC 3.5.-.-)
Length = 262
Score = 106 bits (264), Expect = 9e-23
Identities = 82/264 (31%), Positives = 127/264 (48%), Gaps = 19/264 (7%)
Query: 46 VAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGD-SVSIAQPLDG 104
VAA Q + N C+ L+ +AA L PEA D D SV AQ L+G
Sbjct: 3 VAAGQFAVTSVWEKNAEICASLMAQAAENDVSLFVLPEALLARDDHDADLSVKSAQLLEG 62
Query: 105 PIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVP 164
+ + ++ + + L S P +N V + G I Y K+HL+D
Sbjct: 63 EFLGRLRRESKRNMMTTILT--IHVPSTPGRAWNMLVALQ-AGNIVARYAKLHLYDAFA- 118
Query: 165 GGRVYKESNFTESGKDIVAVDSPIG-RLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAF 223
+ES ++G +I + G ++GL CYDLRFPEL L + GA+IL++PAA+
Sbjct: 119 ----IQESRRVDAGNEIAPLLEVEGMKVGLMTCYDLRFPEL-ALAQALQGAEILVLPAAW 173
Query: 224 TK-VTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDR 282
+ E HW LL ARA++ CY++AA + G N G + IIDP+G + +
Sbjct: 174 VRGPLKEHHWSTLLAARALDTTCYMVAAGECGNKN-----IGQSRIIDPFGVTIAAASE- 227
Query: 283 LSTGIVVADIDLSLVDSVREKMPI 306
+++A++ V VR ++P+
Sbjct: 228 -MPALIMAEVTPERVRQVRAQLPV 250
>YPQQ_PSEFL (P55176) Hypothetical UPF0012 protein in pqqF 5'region
(EC 3.5.-.-) (ORF2)
Length = 285
Score = 86.3 bits (212), Expect = 1e-16
Identities = 80/278 (28%), Positives = 129/278 (45%), Gaps = 22/278 (7%)
Query: 28 ASLSVTTGESTMATNSVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAFS 86
A+ + G S ++RVA Q D+A N ++ EA A LL PE F
Sbjct: 6 ATAATVAGLSVSGVKTMRVALYQCPPRPLDVAGNLQRLHQVAMEATDAD--LLVLPEMF- 62
Query: 87 FVGAKDGDSV--SIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVD 144
G G ++A+ DGP + ++A+ + + L G+ E+ D + ++N ++D
Sbjct: 63 LSGYNIGLEAVGALAEAQDGPSAQRIAAIAQAAGTAI-LYGYPERSVDGQ-IYNAVQLID 120
Query: 145 DTGKIQTTYRKIHLF-DVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPE 203
G+ YRK HLF D+D S F+ D V+ +LG +CYD+ FPE
Sbjct: 121 AQGQRLCNYRKTHLFGDLD--------HSMFSAGEDDFPLVELDGWKLGFLICYDIEFPE 172
Query: 204 LYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESY 263
+ L GA+++LVP A + + ++ +RARA ENQCYV A G H ++
Sbjct: 173 NARRLALA-GAELILVPTA-NMIPYDFVADVTIRARAFENQCYVAYANYCG-HEEQIRYC 229
Query: 264 GDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVR 301
G + I P G+ + L +++ +D L+ R
Sbjct: 230 GQSSIAAPDGSRIALA--GLDEALIIGTLDRQLMGESR 265
>YAG5_STALU (P55178) Hypothetical UPF0012 protein in agr operon (ORF
5) (Fragment)
Length = 234
Score = 84.0 bits (206), Expect = 5e-16
Identities = 67/208 (32%), Positives = 104/208 (49%), Gaps = 20/208 (9%)
Query: 112 SLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKE 171
+LA + + + G K D H+FNT +D TGK+ Y K+HL VP + E
Sbjct: 42 NLALQYQVDIIAGSVSNKHHD--HIFNTAFAIDKTGKVINQYDKMHL----VP---MLDE 92
Query: 172 SNFTESGKDIVAVDSPIGRLGLS--VCYDLRFPELYQLLRF--QHGAQILLVPAAFTKVT 227
F +GK++ + ++ +CYDLRFPEL LR+ + GA I A +
Sbjct: 93 PAFLTAGKNVPETFKLSNGVKVTQMICYDLRFPEL---LRYPARSGATIAFYVAQWPSAR 149
Query: 228 GEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGI 287
HW++LL+ARAIEN YVI G ++ K + G ++ I+P G ++ L
Sbjct: 150 LN-HWQVLLKARAIENNMYVIGCNGCG-YDGKTQYAGHSVAINPNGEIIQELSTTEKELT 207
Query: 288 VVADIDLSLVDSVREKMPIAKVIVCNVY 315
V DID V+ R+ +P+ +V ++Y
Sbjct: 208 VTIDID--AVEQQRKAIPVFDSLVPHLY 233
>YAG5_STAAU (P55177) Hypothetical UPF0012 protein in agr operon (EC
3.5.-.-) (ORF 5)
Length = 261
Score = 80.5 bits (197), Expect = 5e-15
Identities = 62/187 (33%), Positives = 103/187 (54%), Gaps = 24/187 (12%)
Query: 136 LFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDI-----VAVDSPIGR 190
+FNT V+ +G++ Y K+HL VP + +E F +G+ + ++ + + +
Sbjct: 91 IFNTAFSVNKSGQLINEYDKVHL----VP---MLREHEFLTAGEYVAEPFQLSDGTYVTQ 143
Query: 191 LGLSVCYDLRFPELYQLLRF--QHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVI 248
L +CYDLRFPEL LR+ + GA+I A + ++ HW LL+ARAIEN +VI
Sbjct: 144 L---ICYDLRFPEL---LRYPARSGAKIAFYVAQWP-MSRLQHWHSLLKARAIENNMFVI 196
Query: 249 AAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
G + E G +++I+P G +VG L + S I+ D++L+ V+ RE +P+ K
Sbjct: 197 GTNSTG-FDGNTEYAGHSIVINPNGDLVGELNE--SADILTVDLNLNEVEQQRENIPVFK 253
Query: 309 VIVCNVY 315
I ++Y
Sbjct: 254 SIKLDLY 260
>NRL1_ARATH (P32961) Nitrilase 1 (EC 3.5.5.1)
Length = 346
Score = 80.5 bits (197), Expect = 5e-15
Identities = 78/302 (25%), Positives = 137/302 (44%), Gaps = 56/302 (18%)
Query: 41 TNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFV---GAKDGDSV 96
+ +VRV Q +++ D + + + EAAS GA+L+ FPE F G + G +V
Sbjct: 22 STTVRVTIVQSSTVYNDTPATIDKAEKYIVEAASKGAELVLFPEGFIGGYPRGFRFGLAV 81
Query: 97 SI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHV 141
+ A + GP + + +AR++ ++L +G +++G L+ T +
Sbjct: 82 GVHNEEGRDEFRKYHASAIHVPGPEVARLADVARKNHVYLVMGAIEKEGYT---LYCTVL 138
Query: 142 VVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRF 201
G+ +RK+ ++ ++ + + G I D+PIG+LG ++C++ R
Sbjct: 139 FFSPQGQFLGKHRKLMPTSLE---RCIWGQGD----GSTIPVYDTPIGKLGAAICWENRM 191
Query: 202 PELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ--------- 252
P LY+ + G ++ P A G W+ + AIE C+V++A Q
Sbjct: 192 P-LYRTALYAKGIELYCAPTA----DGSKEWQSSMLHIAIEGGCFVLSACQFCQRKHFPD 246
Query: 253 ------AGTHNDKRE----SYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVRE 302
++DK S G ++II P G V+ P+ S G+V ADIDL D R
Sbjct: 247 HPDYLFTDWYDDKEHDSIVSQGGSVIISPLGQVLAG-PNFESEGLVTADIDLG--DIARA 303
Query: 303 KM 304
K+
Sbjct: 304 KL 305
>NRL2_ARATH (P32962) Nitrilase 2 (EC 3.5.5.1)
Length = 339
Score = 78.6 bits (192), Expect = 2e-14
Identities = 78/316 (24%), Positives = 143/316 (44%), Gaps = 61/316 (19%)
Query: 32 VTTGEST-----MATNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAF 85
++T E+T ++ VR Q +++ D + ++ + EAAS G++L+ FPEAF
Sbjct: 1 MSTSENTPFNGVASSTIVRATIVQASTVYNDTPATLEKANKFIVEAASKGSELVVFPEAF 60
Query: 86 SFV---GAKDGDSVSI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQ 127
G + G V + A + GP +++ LA +++++L +G +
Sbjct: 61 IGGYPRGFRFGLGVGVHNEEGRDEFRKYHASAIKVPGPEVEKLAELAGKNNVYLVMGAIE 120
Query: 128 EKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSP 187
+ G L+ T + G+ +RK+ ++ ++ + + G I D+P
Sbjct: 121 KDGYT---LYCTALFFSPQGQFLGKHRKLMPTSLE---RCIWGQGD----GSTIPVYDTP 170
Query: 188 IGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYV 247
IG+LG ++C++ R P LY+ + G ++ P A G W+ + AIE C+V
Sbjct: 171 IGKLGAAICWENRMP-LYRTALYAKGIELYCAPTA----DGSKEWQSSMLHIAIEGGCFV 225
Query: 248 IAAAQ---------------AGTHNDKRE----SYGDTLIIDPWGTVVGRLPDRLSTGIV 288
++A Q ++DK S G ++II P G V+ P+ S G++
Sbjct: 226 LSACQFCLRKDFPDHPDYLFTDWYDDKEPDSIVSQGGSVIISPLGQVLAG-PNFESEGLI 284
Query: 289 VADIDLSLVDSVREKM 304
AD+DL D R K+
Sbjct: 285 TADLDLG--DVARAKL 298
>NRL3_ARATH (P46010) Nitrilase 3 (EC 3.5.5.1)
Length = 346
Score = 77.8 bits (190), Expect = 3e-14
Identities = 76/302 (25%), Positives = 136/302 (44%), Gaps = 56/302 (18%)
Query: 41 TNSVRVAAAQMTSI-TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSF---VGAKDGDSV 96
+++VRV Q +++ D + + + EAAS GAKL+ FPEAF G + G +V
Sbjct: 22 SSTVRVTIVQSSTVYNDTPATLDKAEKFIVEAASKGAKLVLFPEAFIGGYPRGFRFGLAV 81
Query: 97 SI---------------AQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHV 141
+ A + GP +++ LA ++++ L +G ++ G L+ T +
Sbjct: 82 GVHNEEGRDEFRNYHASAIKVPGPEVERLAELAGKNNVHLVMGAIEKDGYT---LYCTAL 138
Query: 142 VVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRF 201
G+ +RK+ ++ ++ + + G I D+PIG++G ++C++ R
Sbjct: 139 FFSPQGQFLGKHRKVMPTSLE---RCIWGQGD----GSTIPVYDTPIGKIGAAICWENRM 191
Query: 202 PELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ--------- 252
P LY+ + G +I P A + W+ + A+E C+V++A Q
Sbjct: 192 P-LYRTALYAKGIEIYCAPTADYSL----EWQASMIHIAVEGGCFVLSAHQFCKRREFPE 246
Query: 253 ----------AGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVRE 302
+D S G ++II P G V+ P+ S G+V AD+DL D R
Sbjct: 247 HPDYLFNDIVDTKEHDPTVSGGGSVIISPLGKVLAG-PNYESEGLVTADLDLG--DIARA 303
Query: 303 KM 304
K+
Sbjct: 304 KL 305
>NRL4_TOBAC (Q42965) Nitrilase 4 (EC 3.5.5.1)
Length = 349
Score = 73.6 bits (179), Expect = 7e-13
Identities = 75/293 (25%), Positives = 128/293 (43%), Gaps = 54/293 (18%)
Query: 40 ATNSVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAF-------SFVGAK 91
+T +VR Q ++I D + RL+ EAAS GA+L+ FPEAF S G
Sbjct: 25 STPTVRATVVQASTIFYDTPATLVKAERLLAEAASYGAQLVVFPEAFIGGYPRGSTFGVS 84
Query: 92 DGDSV-----------SIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTH 140
G+ + A + GP +D+ ++A + ++L +G + G L+ T
Sbjct: 85 IGNRTAKGKEEFRKYHASAIDVPGPEVDRLAAMAGKYKVYLVMGVIERDGYT---LYCTV 141
Query: 141 VVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLR 200
+ D G +RKI +P F + G I D+P+G++G ++C++ R
Sbjct: 142 LFFDSQGHFLGKHRKI------MPTALERIIWGFGD-GSTIPVYDTPLGKIGAAICWENR 194
Query: 201 FPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ-------- 252
P L + + G +I P A ++ W+ + A+E C+V++A Q
Sbjct: 195 MP-LLRTAMYAKGIEIYCAPTADSRDV----WQASMTHIALEGGCFVLSANQFCRRKDYP 249
Query: 253 -------AGTHNDKRES----YGDTLIIDPWGTVVGRLPDRLSTGIVVADIDL 294
+GT D G ++II P G V+ P+ + ++ AD+DL
Sbjct: 250 PPPEYVFSGTEEDLTPDSIVCAGGSVIISPSGAVLAG-PNYVGEALISADLDL 301
>BUP1_HUMAN (Q9UBR1) Beta-ureidopropionase (EC 3.5.1.6)
(Beta-alanine synthase) (N-carbamoyl-beta-alanine
amidohydrolase) (BUP-1)
Length = 384
Score = 70.5 bits (171), Expect = 6e-12
Identities = 63/256 (24%), Positives = 109/256 (41%), Gaps = 26/256 (10%)
Query: 67 LVKEAASAGAKLLCFPEA----FSFVGAKDGDSVSIAQPLDGPIMDQYCSLARESSIWLS 122
+V+ AA G ++CF EA F+F + A+ + ++C ++ +
Sbjct: 103 IVEVAAMCGVNIICFQEAWTMPFAFCTREKLPWTEFAESAEDGPTTRFCQKLAKNHDMVV 162
Query: 123 LGGFQEKGSDPRH-LFNTHVVVDDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDI 181
+ E+ S+ L+NT VV+ ++G + RK H+ V G + + + E
Sbjct: 163 VSPILERDSEHGDVLWNTAVVISNSGAVLGKTRKNHIPRV----GDFNESTYYMEGNLGH 218
Query: 182 VAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAI 241
+ GR+ +++CY P L L+ +GA+I+ P+A E+ W I R AI
Sbjct: 219 PVFQTQFGRIAVNICYGRHHP-LNWLMYSINGAEIIFNPSATIGALSESLWPIEARNAAI 277
Query: 242 ENQCYVIAAAQAGT---------------HNDKRESYGDTLIIDPWGTVVGRLPDRLSTG 286
N C+ A + GT H D YG + + P + L R G
Sbjct: 278 ANHCFTCAINRVGTEHFPNEFTSGDGKKAHQDFGYFYGSSYVAAPDSSRTPGL-SRSRDG 336
Query: 287 IVVADIDLSLVDSVRE 302
++VA +DL+L V +
Sbjct: 337 LLVAKLDLNLCQQVND 352
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,286,451
Number of Sequences: 164201
Number of extensions: 1535457
Number of successful extensions: 3603
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 3480
Number of HSP's gapped (non-prelim): 85
length of query: 327
length of database: 59,974,054
effective HSP length: 110
effective length of query: 217
effective length of database: 41,911,944
effective search space: 9094891848
effective search space used: 9094891848
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146329.11


