

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145767.4 + phase: 0
(218 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
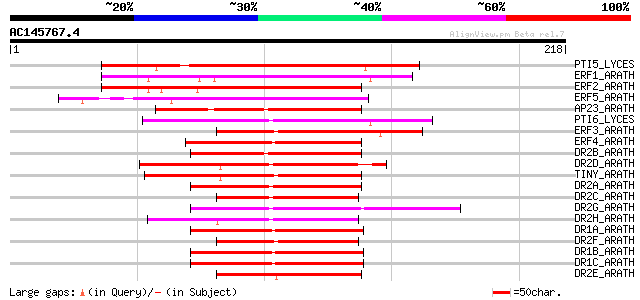
Score E
Sequences producing significant alignments: (bits) Value
PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional a... 118 1e-26
ERF1_ARATH (O80337) Ethylene responsive element binding factor 1... 105 6e-23
ERF2_ARATH (O80338) Ethylene responsive element binding factor 2... 105 8e-23
ERF5_ARATH (O80341) Ethylene responsive element binding factor 5... 101 2e-21
AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Rela... 87 3e-17
PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional a... 83 6e-16
ERF3_ARATH (O80339) Ethylene responsive element binding factor 3... 80 4e-15
ERF4_ARATH (O80340) Ethylene responsive element binding factor 4... 77 2e-14
DR2B_ARATH (O82133) Dehydration responsive element binding prote... 74 2e-13
DR2D_ARATH (Q9LQZ2) Putative dehydration responsive element bind... 74 3e-13
TINY_ARATH (Q39127) Transcriptional factor TINY 72 1e-12
DR2A_ARATH (O82132) Dehydration responsive element binding prote... 71 2e-12
DR2C_ARATH (Q8LFR2) Dehydration responsive element binding prote... 67 3e-11
DR2G_ARATH (P61827) Putative dehydration responsive element bind... 67 4e-11
DR2H_ARATH (Q9SIZ0) Putative dehydration responsive element bind... 66 7e-11
DR1A_ARATH (Q9M0L0) Dehydration responsive element binding prote... 66 7e-11
DR2F_ARATH (Q9SVX5) Dehydration responsive element binding prote... 65 9e-11
DR1B_ARATH (P93835) Dehydration responsive element binding prote... 65 2e-10
DR1C_ARATH (Q9SYS6) Dehydration responsive element binding prote... 64 3e-10
DR2E_ARATH (O80917) Dehydration responsive element binding prote... 64 4e-10
>PTI5_LYCES (O04681) Pathogenesis-related genes transcriptional
activator PTI5
Length = 161
Score = 118 bits (295), Expect = 1e-26
Identities = 69/140 (49%), Positives = 85/140 (60%), Gaps = 18/140 (12%)
Query: 37 LPFNENDPEEMLLYGMITSS--------PQEQTLSKTSNEKTTKSNKDKNNKSYRGVRRR 88
LP NEND +EM+LY ++ + PQ L +N K YRGVRRR
Sbjct: 9 LPLNENDSQEMVLYEVLNEANALNIPYLPQRNQLLPRNN---ILRPLQCIGKKYRGVRRR 65
Query: 89 PWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNF-------SVE 141
PWGK+AAEIRDS RHG RVWLGTF++AE AALAYD+AAF MRG+ A LNF SV
Sbjct: 66 PWGKYAAEIRDSARHGARVWLGTFETAEEAALAYDRAAFRMRGAKALLNFPSEIVNASVS 125
Query: 142 KVKESLRDMNYLLSNDNECS 161
K SL +Y +N+++ S
Sbjct: 126 VDKLSLCSNSYTTNNNSDSS 145
>ERF1_ARATH (O80337) Ethylene responsive element binding factor 1
(AtERF1) (EREBP-2 protein)
Length = 268
Score = 105 bits (263), Expect = 6e-23
Identities = 65/161 (40%), Positives = 92/161 (56%), Gaps = 39/161 (24%)
Query: 37 LPFNENDPEEMLLYGMI------------TSSPQEQTLSKTSNEKTTKS---------NK 75
LP END E+ML+YG++ +SS ++++ + +T +S K
Sbjct: 71 LPLKENDSEDMLVYGILNDAFHGGWEPSSSSSDEDRSSFPSVKIETPESFAAVDSVPVKK 130
Query: 76 DKNN-----------KSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQ 124
+K + K YRGVR+RPWGKFAAEIRD ++G RVWLGTF++AE AALAYD+
Sbjct: 131 EKTSPVSAAVTAAKGKHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDR 190
Query: 125 AAFSMRGSSATLNFSV-------EKVKESLRDMNYLLSNDN 158
AAF MRGS A LNF + + V+ + ++ SN+N
Sbjct: 191 AAFRMRGSRALLNFPLRVNSGEPDPVRIKSKRSSFSSSNEN 231
>ERF2_ARATH (O80338) Ethylene responsive element binding factor 2
(AtERF2)
Length = 243
Score = 105 bits (262), Expect = 8e-23
Identities = 63/126 (50%), Positives = 76/126 (60%), Gaps = 24/126 (19%)
Query: 37 LPFNENDPEEMLLYGMI-------TSSPQ------------EQTLSKTSNEKTTK----- 72
LP END E+ML+YG++ TSS E T + T+ E+ K
Sbjct: 48 LPLKENDSEDMLVYGLLKDAFHFDTSSSDLSCLFDFPAVKVEPTENFTAMEEKPKKAIPV 107
Query: 73 SNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGS 132
+ K YRGVR+RPWGKFAAEIRD ++G RVWLGTF++AE AALAYD AAF MRGS
Sbjct: 108 TETAVKAKHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDIAAFRMRGS 167
Query: 133 SATLNF 138
A LNF
Sbjct: 168 RALLNF 173
>ERF5_ARATH (O80341) Ethylene responsive element binding factor 5
(AtERF5)
Length = 300
Score = 101 bits (251), Expect = 2e-21
Identities = 60/135 (44%), Positives = 79/135 (58%), Gaps = 20/135 (14%)
Query: 20 FGSQDSIS---LNPKNHNEFLPFNENDPEEMLLYGMITSSPQEQTL----------SKTS 66
F S+ S+S P N N+ N+ +PE L I P + +L ++ +
Sbjct: 88 FDSEVSVSDFDFKPSNQNQ----NQFEPE---LKSQIRKPPLKISLPAKTEWIQFAAENT 140
Query: 67 NEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAA 126
+ TK ++ K YRGVR+RPWGKFAAEIRD + G RVWLGTFD+A AA AYD+AA
Sbjct: 141 KPEVTKPVSEEEKKHYRGVRQRPWGKFAAEIRDPNKRGSRVWLGTFDTAIEAARAYDEAA 200
Query: 127 FSMRGSSATLNFSVE 141
F +RGS A LNF +E
Sbjct: 201 FRLRGSKAILNFPLE 215
>AP23_ARATH (P42736) AP2 domain transcription factor RAP2.3 (Related
to AP2 protein 3) (Cadmium-induced protein AS30)
Length = 248
Score = 87.0 bits (214), Expect = 3e-17
Identities = 45/81 (55%), Positives = 57/81 (69%), Gaps = 3/81 (3%)
Query: 58 QEQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEA 117
+E+ + K + K K KN YRG+R+RPWGK+AAEIRD R G+RVWLGTF++AE
Sbjct: 57 KEEAVKKEQATEPGKRRKRKN--VYRGIRKRPWGKWAAEIRDP-RKGVRVWLGTFNTAEE 113
Query: 118 AALAYDQAAFSMRGSSATLNF 138
AA+AYD AA +RG A LNF
Sbjct: 114 AAMAYDVAAKQIRGDKAKLNF 134
>PTI6_LYCES (O04682) Pathogenesis-related genes transcriptional
activator PTI6
Length = 248
Score = 82.8 bits (203), Expect = 6e-16
Identities = 48/125 (38%), Positives = 64/125 (50%), Gaps = 12/125 (9%)
Query: 53 ITSSPQEQTLSKTSNEKTTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTF 112
I P +++ + + K +RGVR+RPWG++AAEIRD TR G RVWLGT+
Sbjct: 69 INLMPSTKSIGDRKRRSVSPDSDVTRRKKFRGVRQRPWGRWAAEIRDPTR-GKRVWLGTY 127
Query: 113 DSAEAAALAYDQAAFSMRGSSATLNFSV-----------EKVKESLRDMNYLLSNDNECS 161
D+ E AA+ YD+AA ++G A NF V E ES+ D ND S
Sbjct: 128 DTPEEAAVVYDKAAVKLKGPDAVTNFPVSTTAEVTVTVTETETESVADGGDKSENDVALS 187
Query: 162 PVIAL 166
P L
Sbjct: 188 PTSVL 192
>ERF3_ARATH (O80339) Ethylene responsive element binding factor 3
(AtERF3)
Length = 225
Score = 80.1 bits (196), Expect = 4e-15
Identities = 43/84 (51%), Positives = 53/84 (62%), Gaps = 4/84 (4%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLNFSVE 141
+RGVR+RPWG+FAAEIRD + RVWLGTFDSAE AA AYD AA ++RG A NF ++
Sbjct: 28 FRGVRKRPWGRFAAEIRDPWKKA-RVWLGTFDSAEEAARAYDSAARNLRGPKAKTNFPID 86
Query: 142 KVK---ESLRDMNYLLSNDNECSP 162
+LR N N+ P
Sbjct: 87 SSSPPPPNLRFNQIRNQNQNQVDP 110
>ERF4_ARATH (O80340) Ethylene responsive element binding factor 4
(AtERF4)
Length = 222
Score = 77.4 bits (189), Expect = 2e-14
Identities = 38/69 (55%), Positives = 48/69 (69%), Gaps = 1/69 (1%)
Query: 70 TTKSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSM 129
T +++ + YRGVR+RPWG++AAEIRD + RVWLGTFD+AE AA AYD AA
Sbjct: 13 TNQTHNNAKEIRYRGVRKRPWGRYAAEIRDPGKK-TRVWLGTFDTAEEAARAYDTAARDF 71
Query: 130 RGSSATLNF 138
RG+ A NF
Sbjct: 72 RGAKAKTNF 80
>DR2B_ARATH (O82133) Dehydration responsive element binding protein
2B (DREB2B protein)
Length = 330
Score = 74.3 bits (181), Expect = 2e-13
Identities = 38/67 (56%), Positives = 49/67 (72%), Gaps = 1/67 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K D ++ S+RGVR+R WGK+ AEIR+ + G R+WLGTF +AE AA AYD+AA +M G
Sbjct: 68 KGGPDNSHCSFRGVRQRIWGKWVAEIREP-KIGTRLWLGTFPTAEKAASAYDEAATAMYG 126
Query: 132 SSATLNF 138
S A LNF
Sbjct: 127 SLARLNF 133
>DR2D_ARATH (Q9LQZ2) Putative dehydration responsive element binding
protein 2D (DREB2D protein)
Length = 206
Score = 73.9 bits (180), Expect = 3e-13
Identities = 42/99 (42%), Positives = 62/99 (62%), Gaps = 8/99 (8%)
Query: 52 MITSSPQEQTLSKTSNEKTTKSNKDKNNKS--YRGVRRRPWGKFAAEIRDSTRHGIRVWL 109
M+ ++ +++T+ +S + + +N S Y+GVR+R WGK+ AEIR+ R G R+WL
Sbjct: 10 MVGANKKQRTVQASSRKGCMRGKGGPDNASCTYKGVRQRTWGKWVAEIREPNR-GARLWL 68
Query: 110 GTFDSAEAAALAYDQAAFSMRGSSATLNFSVEKVKESLR 148
GTFD++ AALAYD AA + G A LN + ESLR
Sbjct: 69 GTFDTSREAALAYDSAARKLYGPEAHLN-----LPESLR 102
>TINY_ARATH (Q39127) Transcriptional factor TINY
Length = 218
Score = 72.0 bits (175), Expect = 1e-12
Identities = 39/86 (45%), Positives = 54/86 (62%), Gaps = 2/86 (2%)
Query: 54 TSSPQEQTLSKTSNEKTTKSNKDKNNKS-YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTF 112
T S + + + + E+ K KD YRGVR+R WGK+ +EIR+ + R+WLGTF
Sbjct: 7 TKSWEASAVRQENEEEKKKPVKDSGKHPVYRGVRKRNWGKWVSEIREPRKKS-RIWLGTF 65
Query: 113 DSAEAAALAYDQAAFSMRGSSATLNF 138
S E AA A+D AA S++G+SA LNF
Sbjct: 66 PSPEMAARAHDVAALSIKGASAILNF 91
>DR2A_ARATH (O82132) Dehydration responsive element binding protein
2A (DREB2A protein)
Length = 335
Score = 70.9 bits (172), Expect = 2e-12
Identities = 36/67 (53%), Positives = 47/67 (69%), Gaps = 1/67 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K + + S+RGVR+R WGK+ AEIR+ R G R+WLGTF +A+ AA AYD+AA +M G
Sbjct: 69 KGGPENSRCSFRGVRQRIWGKWVAEIREPNR-GSRLWLGTFPTAQEAASAYDEAAKAMYG 127
Query: 132 SSATLNF 138
A LNF
Sbjct: 128 PLARLNF 134
>DR2C_ARATH (Q8LFR2) Dehydration responsive element binding protein
2C (DREB2C protein)
Length = 341
Score = 67.0 bits (162), Expect = 3e-11
Identities = 34/56 (60%), Positives = 42/56 (74%), Gaps = 1/56 (1%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLN 137
YRGVR+R WGK+ AEIR+ G R+WLGTF S+ AALAYD+AA ++ G SA LN
Sbjct: 72 YRGVRQRRWGKWVAEIREPDG-GARLWLGTFSSSYEAALAYDEAAKAIYGQSARLN 126
>DR2G_ARATH (P61827) Putative dehydration responsive element binding
protein 2G (DREB2G protein)
Length = 307
Score = 66.6 bits (161), Expect = 4e-11
Identities = 41/106 (38%), Positives = 59/106 (54%), Gaps = 2/106 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K + ++RGVR+R WGK+ AEIR+ R G R+WLGTF+++ AA+AYD+AA + G
Sbjct: 24 KGGPENATCTFRGVRQRTWGKWVAEIREPNR-GTRLWLGTFNTSVEAAMAYDEAAKKLYG 82
Query: 132 SSATLNFSVEKVKESLRDMNYLLSNDNECSPVIALKRKHSMKRKMD 177
A LN V ++ +N LS S A +K M +D
Sbjct: 83 HEAKLNL-VHPQQQQQVVVNRNLSFSGHGSGSWAYNKKLDMVHGLD 127
>DR2H_ARATH (Q9SIZ0) Putative dehydration responsive element binding
protein 2H (DREB2H protein)
Length = 164
Score = 65.9 bits (159), Expect = 7e-11
Identities = 38/85 (44%), Positives = 50/85 (58%), Gaps = 3/85 (3%)
Query: 55 SSPQEQTLSKTSNEKTTKSNKDKNNK--SYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTF 112
S P + K S + K N Y GVR+R WGK+ AEIR+ R G ++WLGTF
Sbjct: 45 SKPIRKAPPKRSRKGCMKGKGGPENGICDYTGVRQRTWGKWVAEIREPGR-GAKLWLGTF 103
Query: 113 DSAEAAALAYDQAAFSMRGSSATLN 137
S+ AALAYD+A+ ++ G SA LN
Sbjct: 104 SSSYEAALAYDEASKAIYGQSARLN 128
>DR1A_ARATH (Q9M0L0) Dehydration responsive element binding protein
1A (DREB1A protein) (C-repeat binding factor 3)
(C-repeat/dehydration responsive element binding factor
3) (CRT/DRE binding factor 3)
Length = 216
Score = 65.9 bits (159), Expect = 7e-11
Identities = 33/68 (48%), Positives = 46/68 (67%), Gaps = 1/68 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K ++ + YRGVRRR GK+ E+R+ + R+WLGTF +AE AA A+D AA ++RG
Sbjct: 41 KKFRETRHPIYRGVRRRNSGKWVCEVREPNKK-TRIWLGTFQTAEMAARAHDVAALALRG 99
Query: 132 SSATLNFS 139
SA LNF+
Sbjct: 100 RSACLNFA 107
>DR2F_ARATH (Q9SVX5) Dehydration responsive element binding protein
2F (DREB2F protein)
Length = 277
Score = 65.5 bits (158), Expect = 9e-11
Identities = 31/56 (55%), Positives = 41/56 (72%), Gaps = 1/56 (1%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRGSSATLN 137
YRGVR+R WGK+ AEIR+ + R+WLG+F +AE AA+AYD+AA + G A LN
Sbjct: 28 YRGVRQRTWGKWVAEIREPKKRA-RLWLGSFATAEEAAMAYDEAALKLYGHDAYLN 82
>DR1B_ARATH (P93835) Dehydration responsive element binding protein
1B (DREB1B protein) (C-repeat binding factor 1)
(C-repeat/dehydration responsive element binding factor
1) (CRT/DRE binding factor 1)
Length = 213
Score = 64.7 bits (156), Expect = 2e-10
Identities = 32/68 (47%), Positives = 47/68 (69%), Gaps = 1/68 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K ++ + YRGVR+R GK+ +E+R+ + R+WLGTF +AE AA A+D AA ++RG
Sbjct: 38 KKFRETRHPIYRGVRQRNSGKWVSEVREPNKK-TRIWLGTFQTAEMAARAHDVAALALRG 96
Query: 132 SSATLNFS 139
SA LNF+
Sbjct: 97 RSACLNFA 104
>DR1C_ARATH (Q9SYS6) Dehydration responsive element binding protein
1C (DREB1C protein) (C-repeat binding factor 2)
(C-repeat/dehydration responsive element binding factor
2) (CRT/DRE binding factor 2)
Length = 216
Score = 63.9 bits (154), Expect = 3e-10
Identities = 32/68 (47%), Positives = 46/68 (67%), Gaps = 1/68 (1%)
Query: 72 KSNKDKNNKSYRGVRRRPWGKFAAEIRDSTRHGIRVWLGTFDSAEAAALAYDQAAFSMRG 131
K ++ + YRGVR+R GK+ E+R+ + R+WLGTF +AE AA A+D AA ++RG
Sbjct: 41 KKFRETRHPIYRGVRQRNSGKWVCELREPNKK-TRIWLGTFQTAEMAARAHDVAAIALRG 99
Query: 132 SSATLNFS 139
SA LNF+
Sbjct: 100 RSACLNFA 107
>DR2E_ARATH (O80917) Dehydration responsive element binding protein
2E (DREB2E protein)
Length = 244
Score = 63.5 bits (153), Expect = 4e-10
Identities = 34/64 (53%), Positives = 41/64 (63%), Gaps = 7/64 (10%)
Query: 82 YRGVRRRPWGKFAAEIRDSTRH-------GIRVWLGTFDSAEAAALAYDQAAFSMRGSSA 134
+RGVR+R WGK+ AEIR+ H R+WLGTF +A AALAYD+AA M G A
Sbjct: 70 FRGVRQRVWGKWVAEIREPVSHRGANSSRSKRLWLGTFATAAEAALAYDRAASVMYGPYA 129
Query: 135 TLNF 138
LNF
Sbjct: 130 RLNF 133
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.128 0.365
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,586,304
Number of Sequences: 164201
Number of extensions: 983966
Number of successful extensions: 2749
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 2683
Number of HSP's gapped (non-prelim): 59
length of query: 218
length of database: 59,974,054
effective HSP length: 106
effective length of query: 112
effective length of database: 42,568,748
effective search space: 4767699776
effective search space used: 4767699776
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC145767.4


