

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145753.3 - phase: 0
(383 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
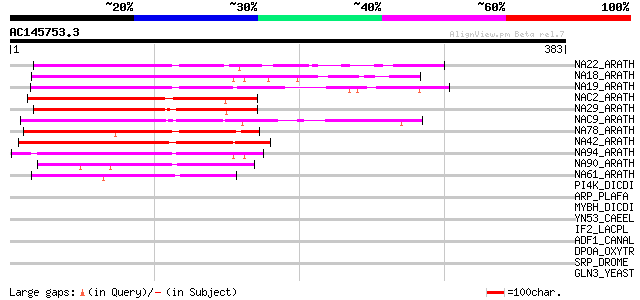
Score E
Sequences producing significant alignments: (bits) Value
NA22_ARATH (Q84TE6) NAC-domain containing protein 21/22 (ANAC021... 216 1e-55
NA18_ARATH (Q9ZNU2) NAC-domain containing protein 18 (ANAC018) (... 180 5e-45
NA19_ARATH (Q9C932) NAC-domain containing protein 19 (ANAC019) (... 177 3e-44
NAC2_ARATH (Q39013) NAC-domain containing protein 2 (ANAC002) 174 3e-43
NA29_ARATH (O49255) NAC-domain containing protein 29 (ANAC029) (... 173 8e-43
NAC9_ARATH (Q9ZVH0) Putative NAC-domain containing protein 9 (AN... 172 2e-42
NA78_ARATH (Q84K00) NAC-domain containing protein 78 (ANAC078) 169 1e-41
NA42_ARATH (Q9SK55) Putative NAC-domain containing protein 42 (A... 164 3e-40
NA94_ARATH (Q9FIW5) Putative NAC-domain containing protein 94 (A... 162 2e-39
NA90_ARATH (Q9FMR3) NAC-domain containing protein 90 (ANAC090) 95 3e-19
NA61_ARATH (Q9M290) Putative NAC-domain containing protein 61 (A... 91 4e-18
PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67) ... 40 0.013
ARP_PLAFA (P04931) Asparagine-rich protein (AG319) (ARP) (Fragment) 38 0.049
MYBH_DICDI (P34127) Myb-like protein (Fragment) 36 0.14
YN53_CAEEL (P34587) Hypothetical protein ZC21.3 in chromosome III 35 0.42
IF2_LACPL (Q88VK7) Translation initiation factor IF-2 35 0.42
ADF1_CANAL (P46589) Adherence factor (Adhesion and aggregation m... 35 0.42
DPOA_OXYTR (Q27152) DNA polymerase alpha catalytic subunit (EC 2... 34 0.54
SRP_DROME (P52172) Box A-binding factor (ABF) (Serpent protein) ... 33 0.93
GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3 33 0.93
>NA22_ARATH (Q84TE6) NAC-domain containing protein 21/22 (ANAC021)
(ANAC022)
Length = 324
Score = 216 bits (549), Expect = 1e-55
Identities = 126/289 (43%), Positives = 168/289 (57%), Gaps = 57/289 (19%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEK-VKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYF 75
LPPGFRFHP D E++ YL + + N+ + + DLNKCEPWD+PK A +G K+WYF
Sbjct: 19 LPPGFRFHPKDDELVCDYLMRRSLHNNHRPPLVLIQVDLNKCEPWDIPKMACVGGKDWYF 78
Query: 76 FCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGE 135
+ Q+DRKY TG+RTNRAT +GYWKATGKD+ I KGK LVGM+KTLVFY+GRAP+G
Sbjct: 79 YSQRDRKYATGLRTNRATATGYWKATGKDRTILRKGK----LVGMRKTLVFYQGRAPRGR 134
Query: 136 KTNWVMHEFRLEGKFATHNLPN----KEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIG 191
KT+WVMHEFRL+G +H+ PN K++WV+ RVFHKNT+ + R N
Sbjct: 135 KTDWVMHEFRLQG---SHHPPNHSLSSPKEDWVLCRVFHKNTE-------GVICRDNMGS 184
Query: 192 HDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKP 251
D +SLPPLMDP Y N D Q YL
Sbjct: 185 CFDETASASLPPLMDP-YINFD---------------QEPSSYL---------------- 212
Query: 252 EDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPN 300
D++H I IN + ++++S LN+ + N++ + +I +NPN
Sbjct: 213 SDDHHYI------INEHVPCFSNLSQNQTLNSNLTNSVSELKIPCKNPN 255
>NA18_ARATH (Q9ZNU2) NAC-domain containing protein 18 (ANAC018) (NO
APICAL MERISTEM protein) (AtNAM)
Length = 320
Score = 180 bits (457), Expect = 5e-45
Identities = 106/281 (37%), Positives = 149/281 (52%), Gaps = 31/281 (11%)
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYF 75
+LPPGFRFHPTD E+++ YL K + I + DL K +PW+LP KA GE+EWYF
Sbjct: 16 NLPPGFRFHPTDEELVIHYLKRKADSVPLPVAIIADVDLYKFDPWELPAKASFGEQEWYF 75
Query: 76 FCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGE 135
F +DRKYP G R NRA SGYWKATG DK + G G VG+KK LVFY G+ PKG
Sbjct: 76 FSPRDRKYPNGARPNRAATSGYWKATGTDKPVISTGGGGSKKVGVKKALVFYSGKPPKGV 135
Query: 136 KTNWVMHEFRLEGKFATH--NLPNKEK----DEWVVSRVFHKNTDVKK---PQISSGLLR 186
K++W+MHE+RL TH + NK+ D+WV+ R++ KN + L
Sbjct: 136 KSDWIMHEYRLTDNKPTHICDFGNKKNSLRLDDWVLCRIYKKNNSTASRHHHHLHHIHLD 195
Query: 187 INSIGHDDLLD---YSSLPP-LMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINN 242
+ HD ++D + +PP L P+ +D+ N + GY S+S+N
Sbjct: 196 NDHHRHDMMIDDDRFRHVPPGLHFPAIFSDN------NDPTAIYDGGGGGYGGGSYSMN- 248
Query: 243 NQHQFLIKPEDNYHRINYDQHEINPTMMNYTSISNQSNLNN 283
H F + Q ++ P +M TS++ S + +
Sbjct: 249 --HCFASGSK---------QEQLFPPVMMMTSLNQDSGIGS 278
>NA19_ARATH (Q9C932) NAC-domain containing protein 19 (ANAC019)
(ANAC) (Abscicic-acid-responsive NAC)
Length = 317
Score = 177 bits (450), Expect = 3e-44
Identities = 112/300 (37%), Positives = 150/300 (49%), Gaps = 50/300 (16%)
Query: 15 LDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWY 74
L LPPGFRF+PTD E++V YL K FS I E DL K +PW LP KA GEKEWY
Sbjct: 12 LSLPPGFRFYPTDEELMVQYLCRKAAGYDFSLQLIAEIDLYKFDPWVLPNKALFGEKEWY 71
Query: 75 FFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKG 134
FF +DRKYP G R NR SGYWKATG DK I +G+ VG+KK LVFY G+APKG
Sbjct: 72 FFSPRDRKYPNGSRPNRVAGSGYWKATGTDKIISTEGQ----RVGIKKALVFYIGKAPKG 127
Query: 135 EKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQISSGLLRINSIGHDD 194
KTNW+MHE+RL + + + D+WV+ R++ K + +K +G+
Sbjct: 128 TKTNWIMHEYRLIEPSRRNG--STKLDDWVLCRIYKKQSSAQKQVYDNGIANAREF---- 181
Query: 195 LLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGY--YLPSF--SINNNQHQFLIK 250
+N SST S S + L SF I+N QF
Sbjct: 182 ------------------------SNNGTSSTTSSSSHFEDVLDSFHQEIDNRNFQF--- 214
Query: 251 PEDNYHRINYDQHEINPTMMNYTSISNQSNL-------NNPIGNTLPQPQIRIQNPNLNY 303
N +RI+ + ++ + +++ SN N N++P+ + PNL Y
Sbjct: 215 --SNPNRISSLRPDLTEQKTGFHGLADTSNFDWASFAGNVEHNNSVPELGMSHVVPNLEY 272
>NAC2_ARATH (Q39013) NAC-domain containing protein 2 (ANAC002)
Length = 289
Score = 174 bits (442), Expect = 3e-43
Identities = 83/161 (51%), Positives = 105/161 (64%), Gaps = 7/161 (4%)
Query: 13 QTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKE 72
+ L LPPGFRFHPTD E+++ YL K + + I E DL K +PW+LP A GEKE
Sbjct: 3 ELLQLPPGFRFHPTDEELVMHYLCRKCASQSIAVPIIAEIDLYKYDPWELPGLALYGEKE 62
Query: 73 WYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAP 132
WYFF +DRKYP G R NR+ SGYWKATG DK I G+ VG+KK LVFY G+AP
Sbjct: 63 WYFFSPRDRKYPNGSRPNRSAGSGYWKATGADKPI-----GLPKPVGIKKALVFYAGKAP 117
Query: 133 KGEKTNWVMHEFRLE--GKFATHNLPNKEKDEWVVSRVFHK 171
KGEKTNW+MHE+RL + + D+WV+ R+++K
Sbjct: 118 KGEKTNWIMHEYRLADVDRSVRKKKNSLRLDDWVLCRIYNK 158
>NA29_ARATH (O49255) NAC-domain containing protein 29 (ANAC029)
(NAC2) (NAC-LIKE, ACTIVATED BY AP3/PI protein) (NAP)
Length = 268
Score = 173 bits (438), Expect = 8e-43
Identities = 84/157 (53%), Positives = 105/157 (66%), Gaps = 6/157 (3%)
Query: 17 LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMGEKEWYFF 76
LPPGFRFHPTD E+IV YL + + + I E D+ K +PW LP+K + GE EWYFF
Sbjct: 9 LPPGFRFHPTDEELIVYYLRNQTMSKPCPVSIIPEVDIYKFDPWQLPEKTEFGENEWYFF 68
Query: 77 CQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPKGEK 136
++RKYP G+R NRA SGYWKATG DK I H G + VG+KK LVFYKGR PKG K
Sbjct: 69 SPRERKYPNGVRPNRAAVSGYWKATGTDKAI-HSG---SSNVGVKKALVFYKGRPPKGIK 124
Query: 137 TNWVMHEFRLEG--KFATHNLPNKEKDEWVVSRVFHK 171
T+W+MHE+RL K +T + DEWV+ R++ K
Sbjct: 125 TDWIMHEYRLHDSRKASTKRNGSMRLDEWVLCRIYKK 161
>NAC9_ARATH (Q9ZVH0) Putative NAC-domain containing protein 9
(ANAC009)
Length = 418
Score = 172 bits (435), Expect = 2e-42
Identities = 107/293 (36%), Positives = 149/293 (50%), Gaps = 45/293 (15%)
Query: 8 NKGEDQTLD-LPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKA 66
N G+ + D L PGFRFHPTD E++ YL KV+++ S I + D+ K +PWDLPK A
Sbjct: 6 NDGDQKMEDVLLPGFRFHPTDEELVSFYLKRKVQHNPLSIELIRQLDIYKYDPWDLPKFA 65
Query: 67 KMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVF 126
GEKEWYF+C +DRKY R NR T +G+WKATG D+ IY +G +G+KK+LVF
Sbjct: 66 MTGEKEWYFYCPRDRKYRNSSRPNRVTGAGFWKATGTDRPIY-SSEG-NKCIGLKKSLVF 123
Query: 127 YKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKE--------KDEWVVSRVFHKNTDVKKP 178
YKGRA KG KT+W+MHEFRL + + P+K D W + R+F K
Sbjct: 124 YKGRAAKGVKTDWMMHEFRLP-SLSEPSPPSKRFFDSPVSPNDSWAICRIFKKTNTTTLR 182
Query: 179 QISSGLLRINSIGHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSF 238
+S + SSLPP T+ S + QS+ Y+ S
Sbjct: 183 ALSHSFV-------------SSLPP--------------ETSTDTMSNQKQSNTYHFSSD 215
Query: 239 SINNNQHQFLIKPED-NYHRINYDQHEINPTM-----MNYTSISNQSNLNNPI 285
I F E+ N + + PT+ +++TS +N+ NP+
Sbjct: 216 KILKPSSHFQFHHENMNTPKTSNSTTPSVPTISPFSYLDFTSYDKPTNVFNPV 268
>NA78_ARATH (Q84K00) NAC-domain containing protein 78 (ANAC078)
Length = 567
Score = 169 bits (428), Expect = 1e-41
Identities = 84/166 (50%), Positives = 109/166 (65%), Gaps = 10/166 (6%)
Query: 10 GEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKKAKMG 69
G L PGFRFHPTD E++ YL KV N F AI D+ K EPWDLP K+K+
Sbjct: 2 GRGSVTSLAPGFRFHPTDEELVRYYLKRKVCNKPFKFDAISVTDIYKSEPWDLPDKSKLK 61
Query: 70 EK--EWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFY 127
+ EWYFF D+KY G +TNRATE GYWK TGKD+EI + + +VGMKKTLV++
Sbjct: 62 SRDLEWYFFSMLDKKYSNGSKTNRATEKGYWKTTGKDREIRNGSR----VVGMKKTLVYH 117
Query: 128 KGRAPKGEKTNWVMHEFRLEGK-FATHNLPNKEKDEWVVSRVFHKN 172
KGRAP+GE+TNWVMHE+RL + +P ++ +V+ R+F K+
Sbjct: 118 KGRAPRGERTNWVMHEYRLSDEDLKKAGVP---QEAYVLCRIFQKS 160
>NA42_ARATH (Q9SK55) Putative NAC-domain containing protein 42
(ANAC042)
Length = 275
Score = 164 bits (416), Expect = 3e-40
Identities = 79/175 (45%), Positives = 110/175 (62%), Gaps = 6/175 (3%)
Query: 7 VNKGEDQTLDLP-PGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWDLPKK 65
+ K ++ + P PGFRFHPTD E++ YL KV+N I + D+ K +PWDLP+
Sbjct: 7 LGKDHEEENEAPLPGFRFHPTDEELLGYYLRRKVENKTIKLELIKQIDIYKYDPWDLPRV 66
Query: 66 AKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLV 125
+ +GEKEWYFFC + RKY +R NR T SG+WKATG DK +Y + VG+KK+LV
Sbjct: 67 SSVGEKEWYFFCMRGRKYRNSVRPNRVTGSGFWKATGIDKPVYSN----LDCVGLKKSLV 122
Query: 126 FYKGRAPKGEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVFHKNTDVKKPQI 180
+Y G A KG KT+W+MHEFRL T + P ++ + W + R+F + T + P I
Sbjct: 123 YYLGSAGKGTKTDWMMHEFRLPSTTKTDS-PAQQAEVWTLCRIFKRVTSQRNPTI 176
>NA94_ARATH (Q9FIW5) Putative NAC-domain containing protein 94
(ANAC094)
Length = 337
Score = 162 bits (409), Expect = 2e-39
Identities = 86/193 (44%), Positives = 109/193 (55%), Gaps = 24/193 (12%)
Query: 2 EEPIVVNKGEDQTLDLPPGFRFHPTDAEIIVCYLTEKVKNSKFSATAIGEADLNKCEPWD 61
EE V + +D L PGFRFHPTD E++ YL KV + I + D+ K +PWD
Sbjct: 8 EESNNVERYDDVVL---PGFRFHPTDEELVSFYLKRKVLHKSLPFDLIKKVDIYKYDPWD 64
Query: 62 LPKKAKMGEKEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMK 121
LPK A MGEKEWYF+C +DRKY R NR T G+WKATG D+ IY +G+K
Sbjct: 65 LPKLAAMGEKEWYFYCPRDRKYRNSTRPNRVTGGGFWKATGTDRPIYSLDS--TRCIGLK 122
Query: 122 KTLVFYKGRAPKGEKTNWVMHEFRLEGKFATH-----NLPNKEK--------------DE 162
K+LVFY+GRA KG KT+W+MHEFRL +H N NK++ D
Sbjct: 123 KSLVFYRGRAAKGVKTDWMMHEFRLPSLSDSHHSSYPNYNNKKQHLNNNNNSKELPSNDA 182
Query: 163 WVVSRVFHKNTDV 175
W + R+F K V
Sbjct: 183 WAICRIFKKTNAV 195
>NA90_ARATH (Q9FMR3) NAC-domain containing protein 90 (ANAC090)
Length = 235
Score = 94.7 bits (234), Expect = 3e-19
Identities = 52/156 (33%), Positives = 83/156 (52%), Gaps = 9/156 (5%)
Query: 20 GFRFHPTDAEIIVCYLTEKVKNSKFSAT--AIGEADLNKCEPWDLPKKAKM----GEKEW 73
GFRF+PT+ E++ YL +++ + I D+ + EP LP A + ++W
Sbjct: 8 GFRFYPTEEELVSFYLRNQLEGRSDDSMHRVIPVLDVFEVEPSHLPNVAGVRCRGDAEQW 67
Query: 74 YFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGRAPK 133
+FF + + G R +R T SGYWKATG ++ K ++G KKT+VFY G+AP
Sbjct: 68 FFFVPRQEREARGGRPSRTTGSGYWKATGSPGPVFSKDN---KMIGAKKTMVFYTGKAPT 124
Query: 134 GEKTNWVMHEFRLEGKFATHNLPNKEKDEWVVSRVF 169
G KT W M+E+ + + K + E+ + RV+
Sbjct: 125 GRKTKWKMNEYHAVDETVNASTIPKLRREFSLCRVY 160
>NA61_ARATH (Q9M290) Putative NAC-domain containing protein 61
(ANAC061)
Length = 228
Score = 91.3 bits (225), Expect = 4e-18
Identities = 48/146 (32%), Positives = 78/146 (52%), Gaps = 8/146 (5%)
Query: 16 DLPPGFRFHPTDAEIIVCYLTEKVKNSKFSA-TAIGEADLNKCEPWDLP----KKAKMGE 70
+L GFRF+PT+ E++ YL ++ + + I D+ EP LP ++ +
Sbjct: 4 ELSVGFRFYPTEVELLTYYLRIQLGGGNATIHSLIPILDVFSVEPTQLPNLAGERCRGDA 63
Query: 71 KEWYFFCQKDRKYPTGMRTNRATESGYWKATGKDKEIYHKGKGIQNLVGMKKTLVFYKGR 130
++W FF + + G R +R T SGYWKATG ++ + +G+KKT+VFY G+
Sbjct: 64 EQWIFFVPRQEREARGGRPSRTTGSGYWKATGSPGPVFSPDNRV---IGVKKTMVFYTGK 120
Query: 131 APKGEKTNWVMHEFRLEGKFATHNLP 156
AP G KT W M+E++ + +P
Sbjct: 121 APTGRKTKWKMNEYKAVETASVSTIP 146
>PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67)
(PI4-kinase) (PtdIns-4-kinase) (PI4K-alpha)
Length = 1093
Score = 39.7 bits (91), Expect = 0.013
Identities = 40/170 (23%), Positives = 71/170 (41%), Gaps = 19/170 (11%)
Query: 200 SLPPLMDPSYTNDDFKGITTNQQISSTKSQSDGYYLPSFS-----INNNQHQFLIKPEDN 254
S P P Y+ND I ++ + ++ D + L I H + E++
Sbjct: 117 STSPKDVPMYSNDQIVPIDLDKIYNQSQYDDDDFDLSDDDGGFEIIKKKCHHY----END 172
Query: 255 YHRINYDQHEINPTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSSM 314
+H N + +IN N +I+N ++ N+ N +I + N +N +
Sbjct: 173 HHIENDPKKDINSNNNNNNNINNNNSNNDDNNNN----EILPNENSDNSINDENNQYGNS 228
Query: 315 PTNVYGSGKNNECKVEQFSSNQSQ------DTGLSNDTSSAVSKLDMERN 358
N SG+NN K++ S N+S ++ L +T ++ K DME N
Sbjct: 229 NNNNNISGENNNIKIDINSQNKSDSNIETLNSTLCEETKTSPIKDDMENN 278
>ARP_PLAFA (P04931) Asparagine-rich protein (AG319) (ARP) (Fragment)
Length = 537
Score = 37.7 bits (86), Expect = 0.049
Identities = 47/208 (22%), Positives = 89/208 (42%), Gaps = 17/208 (8%)
Query: 171 KNTDVKKPQIS---SGLLRINSIGHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQISSTK 227
KNTD K S G + ++ D L + +++ S N + + T+
Sbjct: 124 KNTDNNKTDTSYNMKGTINNDNNNMDYLRNINNINEYKG-SAKNKFYTNYMNKNNLKFTQ 182
Query: 228 SQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYDQHEINPTMMNYTSISNQSNLNNPIGN 287
+ +D + + NNN + +N NY + +N N +I N NN I N
Sbjct: 183 NNNDNMNINEDNNNNNNNN----NNNNGVFSNYQNNNMNRN--NSINIKRNLNNNNNINN 236
Query: 288 TLPQ--PQIRIQNPNLNYFM---YQNRMQSSMPTNVYGSGKNNECKVEQFSSNQSQDTGL 342
+ + Q + QN N N++M YQNR ++SM N+ + NN + N + + +
Sbjct: 237 NMNKMGSQDKNQNSNNNFYMNYNYQNR-KNSMNNNMNNNMNNNMNHNMNNNMNHNMNNNM 295
Query: 343 SNDTSSAVSKLDMERNRALYDDLEGPSS 370
+++ ++ ++ +M N + L+ S
Sbjct: 296 NHNMNNNMNH-NMNNNMNNINSLDSDMS 322
>MYBH_DICDI (P34127) Myb-like protein (Fragment)
Length = 451
Score = 36.2 bits (82), Expect = 0.14
Identities = 24/86 (27%), Positives = 41/86 (46%), Gaps = 10/86 (11%)
Query: 241 NNNQHQFLIKPEDNYHRINYDQHEIN-PTMMNYTSISNQSNLNNPIGNTLPQPQIRIQNP 299
NNN + + +D+ ++H I+ P +N SI+N +NL N I N +
Sbjct: 58 NNNNEEEEEEDDDDDDDSQQNRHNISIPFTINKNSINNNNNLINNINNNI---------N 108
Query: 300 NLNYFMYQNRMQSSMPTNVYGSGKNN 325
N+N M N ++M N+ +G +N
Sbjct: 109 NMNNNMNNNNNNNNMNNNINNNGNSN 134
>YN53_CAEEL (P34587) Hypothetical protein ZC21.3 in chromosome III
Length = 321
Score = 34.7 bits (78), Expect = 0.42
Identities = 31/140 (22%), Positives = 56/140 (39%), Gaps = 15/140 (10%)
Query: 208 SYTNDDFKGITTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYDQHEINP 267
+Y+ G ++ S+ S S+ Y PS S N Q + N + + YD N
Sbjct: 88 TYSPTSTYGYSSQNYNPSSASSSNVYRNPSTSYGNTQ-------QSNQNYVAYDTTNQNS 140
Query: 268 TMMNYTSISNQSNLNNPIGNTLPQPQ-------IRIQNPNLNYFMYQNRMQSSMPTNVYG 320
Y + + + N+ +G PQ +R Q+ N Y M ++ T YG
Sbjct: 141 PYY-YQRVPSTAQYNSVLGVNAGFPQSSNQNQVLRDQSNNQGYMNNNQGMYNTQTTQGYG 199
Query: 321 SGKNNECKVEQFSSNQSQDT 340
+ + + +++N Q+T
Sbjct: 200 NNQGSYQNQMMYNNNNPQNT 219
>IF2_LACPL (Q88VK7) Translation initiation factor IF-2
Length = 858
Score = 34.7 bits (78), Expect = 0.42
Identities = 40/140 (28%), Positives = 61/140 (43%), Gaps = 16/140 (11%)
Query: 218 TTNQQISSTKSQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYDQHEINPTMMNYTSISN 277
T NQQ S+++ Q G YL + + NNNQ+ + N +R N + N N ++ ++
Sbjct: 78 TANQQQSNSRHQQSGDYLNN-NRNNNQNNSQSR-NSNGNRSNNGGNRTNSNNGNRSANAS 135
Query: 278 QSN--LNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSS--------MPTNVYGSGKNNEC 327
+SN NN NT N N N NR +S TN G +NN
Sbjct: 136 RSNSARNNGGSNT----NRSNNNNNNNRSNNNNRSNTSNNRTNSTGNRTNNGGQNRNNNR 191
Query: 328 KVEQFSSNQSQDTGLSNDTS 347
++N S + SN++S
Sbjct: 192 TTTNNNNNNSNRSNGSNNSS 211
>ADF1_CANAL (P46589) Adherence factor (Adhesion and aggregation
mediating surface antigen)
Length = 612
Score = 34.7 bits (78), Expect = 0.42
Identities = 43/175 (24%), Positives = 70/175 (39%), Gaps = 29/175 (16%)
Query: 200 SLPPLM--DPSYTNDDFKGITT----NQQISSTKSQSDGYYLPSFSINNNQHQFLIKPED 253
++PPL PS ++ + NQQ S + Q Y NN QF
Sbjct: 90 AMPPLQTQQPSSSSATTNNVVPPHHYNQQQSQQQQQQQQQYQQMQPQPNNM-QFFDNTIP 148
Query: 254 NYHRINYDQHEINPTMMNYTSISNQSNLN----------NPIGNTLPQPQIRIQNPNLNY 303
NY +N I+P+ T+ N S N PI ++ PQPQ + P N
Sbjct: 149 NYLIMN---QTISPSQTQTTAQPNISYYNYSTQPQLQSAQPISHSQPQPQPQATQPRSN- 204
Query: 304 FMYQNRMQSSM-----PTNVYGSGKNNECKVEQFSSNQSQDTGLSNDTSSAVSKL 353
++R Q+S V GSG++ K + ++ S TG + + + ++ +
Sbjct: 205 ---RSRSQTSFSKPRGSRQVSGSGRSTGAKKQSAITSGSTGTGPARNADTGMTSV 256
>DPOA_OXYTR (Q27152) DNA polymerase alpha catalytic subunit (EC
2.7.7.7)
Length = 1513
Score = 34.3 bits (77), Expect = 0.54
Identities = 29/114 (25%), Positives = 49/114 (42%), Gaps = 8/114 (7%)
Query: 275 ISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSSMPTNVYGSGKN------NECK 328
+SNQ+N N T +++ NP + RM SM + + KN N K
Sbjct: 248 VSNQTNDNLNKVETQVISMLKVANPQIKVTSKLARMDDSMKIDQTSAQKNQIEQTVNHSK 307
Query: 329 VEQFSSNQSQD-TGLSNDTSSAVSKLDMERNRALYDDLEGPSSVAPLSDLDSFW 381
+++++ NQ L N TS + L++ N+ L + E P + L +W
Sbjct: 308 LQRYNQNQMMSGIKLDNRTSKPIRNLEI-WNQVLANTQEFPLPLNENGTLSFYW 360
>SRP_DROME (P52172) Box A-binding factor (ABF) (Serpent protein)
(GATA-binding factor-B) (Transcription factor GATA-B)
(dGATA-B)
Length = 779
Score = 33.5 bits (75), Expect = 0.93
Identities = 55/256 (21%), Positives = 97/256 (37%), Gaps = 27/256 (10%)
Query: 107 IYHKGKGIQNLVGMKKTLVFYKGRAPKGEKTNWVMHEFR--LEGKFATHNLPNKEKDEWV 164
+Y+K + + MKK + + R PKG K+ + + L + +L + V
Sbjct: 345 LYYKLHSVPRPLTMKKDTIQKRKRKPKGTKSEKSKSKSKNALNAIMESGSLVTNCHNVGV 404
Query: 165 V--SRVFHKNTDVKKPQISSGLLRINSIGHDDLLDYSSLPPLMDPSYTNDDFKGITTNQQ 222
V S N D+ KPQ+ L NS YSS P P Y + + Q
Sbjct: 405 VLDSSQMDVNDDM-KPQLD--LKPYNS--------YSSQPQQQLPQYQQQ--QQLVMADQ 451
Query: 223 ISSTKSQSDGYYLPSFSINNNQHQFLIKPEDNYHRINYDQHEINPTM---MNYTSISNQS 279
SS S S S + HQ P + PT ++ S+ S
Sbjct: 452 HSSAASSPHSMGSTSLSPSAMSHQHQTHPHQQQQQQLCSGWTCRPTQTTKCRRSTCSSIS 511
Query: 280 NLNNPIGNTLPQPQIRIQNPNLNYFMYQNR-------MQSSMPTNVYGSGKNNECKVEQF 332
+ N +T P + +Q+P+ + ++ N ++ N S +NN ++++
Sbjct: 512 SSNRAACSTHPAHPLHLQHPSPTHQLHNNNNNNNSSLFNNNNNNNNSSSNENNNKLIQKY 571
Query: 333 SSNQSQDTGLSNDTSS 348
Q + ++D++S
Sbjct: 572 LQAQQLSSSSNSDSTS 587
>GLN3_YEAST (P18494) Nitrogen regulatory protein GLN3
Length = 730
Score = 33.5 bits (75), Expect = 0.93
Identities = 21/86 (24%), Positives = 41/86 (47%), Gaps = 2/86 (2%)
Query: 268 TMMNYTSISNQSNLNNPIGNTLPQPQIRIQNPNLNYFMYQNRMQSSMPTNVYGSGKNNEC 327
++ Y S ++ SN+N P N + + + N + QN SS N+ + +N
Sbjct: 181 SLFPYNSSTSNSNINQPSINNNSNTNAQSHH-SFNIYKLQNNNSSSSAMNITNNNNSNNS 239
Query: 328 KVEQFSSNQSQDTGL-SNDTSSAVSK 352
++ +S GL S++T+++V K
Sbjct: 240 NIQHPFLKKSDSIGLSSSNTTNSVRK 265
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.132 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,236,871
Number of Sequences: 164201
Number of extensions: 2202848
Number of successful extensions: 3618
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 3545
Number of HSP's gapped (non-prelim): 73
length of query: 383
length of database: 59,974,054
effective HSP length: 112
effective length of query: 271
effective length of database: 41,583,542
effective search space: 11269139882
effective search space used: 11269139882
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC145753.3


