

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145513.12 + phase: 0
(304 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
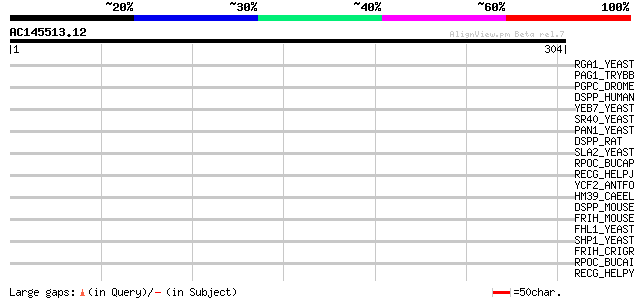
Score E
Sequences producing significant alignments: (bits) Value
RGA1_YEAST (P39083) Rho-type GTPase-activating protein 1 35 0.18
PAG1_TRYBB (Q01889) PAG1 protein precursor 35 0.23
PGPC_DROME (Q9GNK5) Peptidoglycan-recognition protein-LC (Immune... 35 0.30
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 33 0.88
YEB7_YEAST (P39996) Hypothetical 38.2 kDa protein in PMP2-VAC8 i... 33 1.1
SR40_YEAST (P32583) Suppressor protein SRP40 33 1.1
PAN1_YEAST (P32521) PAN1 protein 33 1.1
DSPP_RAT (Q62598) Dentin sialophosphoprotein precursor [Contains... 32 1.5
SLA2_YEAST (P33338) SLA2 protein (Transmembrane protein MOP2) 32 2.5
RPOC_BUCAP (P41185) DNA-directed RNA polymerase beta' chain (EC ... 31 3.3
RECG_HELPJ (Q9ZJA1) ATP-dependent DNA helicase recG (EC 3.6.1.-) 31 3.3
YCF2_ANTFO (Q859W7) Protein ycf2 31 4.3
HM39_CAEEL (Q22812) Homeobox protein ceh-39 31 4.3
DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin... 31 4.3
FRIH_MOUSE (P09528) Ferritin heavy chain (Ferritin H subunit) 30 5.7
FHL1_YEAST (P39521) Pre-rRNA processing protein FHL1 30 5.7
SHP1_YEAST (P34223) SHP1 protein 30 7.4
FRIH_CRIGR (P29389) Ferritin heavy chain (Ferritin H subunit) 30 7.4
RPOC_BUCAI (P57145) DNA-directed RNA polymerase beta' chain (EC ... 30 9.7
RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-) 30 9.7
>RGA1_YEAST (P39083) Rho-type GTPase-activating protein 1
Length = 1007
Score = 35.4 bits (80), Expect = 0.18
Identities = 20/79 (25%), Positives = 41/79 (51%)
Query: 29 LSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSV 88
++++ SGSE S E +SS + + G+ + DD + G++ + + +S + V
Sbjct: 178 VNEIPSGSEPSKDIETNSSDIVPHFITGYNDSDDNSGSSKFGSNVSIDVIGPEENSTEHV 237
Query: 89 EDLLRLSAENANADSYLNM 107
D ++ AE +A+ LN+
Sbjct: 238 NDDVKEEAEAPSANMSLNV 256
>PAG1_TRYBB (Q01889) PAG1 protein precursor
Length = 405
Score = 35.0 bits (79), Expect = 0.23
Identities = 28/124 (22%), Positives = 59/124 (47%), Gaps = 5/124 (4%)
Query: 33 SSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLL 92
SSG G + + ++L++ +L + + + S G+ ERV+ ++ + +S+ +LL
Sbjct: 163 SSGKGDFCVGNSEHHATRNQLLNCYLSDSEKDEFISLGDMHRMERVEEMNMTRESMNELL 222
Query: 93 RLSAENAN-ADSYLNMLRLHVSEAAVKFDFLKKQSVS---VYNRNVMSFLRE-KGHNAAI 147
RL+ N + Y++ H++ +L + S S ++ ++S R +G
Sbjct: 223 RLTGGGGNPLERYVHSANCHLTNTRPGGGYLSENSTSRRILWGDGIISLKRSGRGFVGGT 282
Query: 148 CKTR 151
KTR
Sbjct: 283 GKTR 286
>PGPC_DROME (Q9GNK5) Peptidoglycan-recognition protein-LC (Immune
response deficient 7 protein)
Length = 520
Score = 34.7 bits (78), Expect = 0.30
Identities = 22/92 (23%), Positives = 39/92 (41%)
Query: 12 PFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGN 71
PF++E + RR++D +G ++ T DS L++ V F E + +
Sbjct: 2 PFSNETEMSQCSNAKRRVNDPLTGPKNCSTSSTDSGVILNDNVAAFRPEKETKDRGTGEG 61
Query: 72 DFDSERVDSVSDSMDSVEDLLRLSAENANADS 103
F S+ + SVE + ++ EN S
Sbjct: 62 QFQSKSEEKTESKRISVEHTVNITTENVGKTS 93
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 33.1 bits (74), Expect = 0.88
Identities = 23/74 (31%), Positives = 37/74 (49%), Gaps = 1/74 (1%)
Query: 30 SDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVE 89
S S S+ + + + DSS S S D N DS+ + S+ DS SDS DS
Sbjct: 566 SSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDS-SDSSDSKS 624
Query: 90 DLLRLSAENANADS 103
D + ++++++DS
Sbjct: 625 DSSKSESDSSDSDS 638
Score = 30.4 bits (67), Expect = 5.7
Identities = 22/74 (29%), Positives = 34/74 (45%), Gaps = 2/74 (2%)
Query: 30 SDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVE 89
SD S+ S+ S + + DSS S S + D + DS +D S SDS DS +
Sbjct: 555 SDSSNSSDSSDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSD--SSNSSDSSDSSDSSD 612
Query: 90 DLLRLSAENANADS 103
+ ++ +DS
Sbjct: 613 SSDSSDSSDSKSDS 626
>YEB7_YEAST (P39996) Hypothetical 38.2 kDa protein in PMP2-VAC8
intergenic region
Length = 337
Score = 32.7 bits (73), Expect = 1.1
Identities = 33/124 (26%), Positives = 52/124 (41%), Gaps = 31/124 (25%)
Query: 30 SDVSSGSEH-SGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSV 88
++ SGS + SG+G CD++ + S+L +++EDD D S D
Sbjct: 77 NEYGSGSGNGSGSGSCDTATNDSDLEKAYIKEDD----------------DEKPQSGDET 120
Query: 89 EDLLRLSAENANADSYLNMLRLHVS-------EAAVKFDFLKKQSVS-------VYNRNV 134
LS+ NAN+++ N L S A KF F ++ +S N NV
Sbjct: 121 SATKPLSSRNANSNAKTNFNLLDFSTDNDSSTSAFTKFKFNFQEYLSDIRYQTQKLNENV 180
Query: 135 MSFL 138
+L
Sbjct: 181 QDYL 184
>SR40_YEAST (P32583) Suppressor protein SRP40
Length = 406
Score = 32.7 bits (73), Expect = 1.1
Identities = 27/98 (27%), Positives = 42/98 (42%), Gaps = 14/98 (14%)
Query: 6 KKRVTDPFNDEAKARLVGADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNV 65
KKR + N++AK + + SS SE S +G SS E + G+
Sbjct: 127 KKRARESDNEDAKET---KKAKTEPESSSSSESSSSGSSSSS-----------ESESGSE 172
Query: 66 CDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADS 103
DS + S DS SDS + S+ ++++DS
Sbjct: 173 SDSDSSSSSSSSSDSESDSESDSQSSSSSSSSDSSSDS 210
>PAN1_YEAST (P32521) PAN1 protein
Length = 1480
Score = 32.7 bits (73), Expect = 1.1
Identities = 27/116 (23%), Positives = 44/116 (37%), Gaps = 16/116 (13%)
Query: 69 TGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVS 128
TG +SE S+ D S E + R+ EN + +R +S+ + S
Sbjct: 890 TGKSTESEDSLSMEDEQQSAE-IKRIQQENGKNQEIIKDIRSSISDISASLKSTMTGSNM 948
Query: 129 VYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFIDVVRMRSGSSTWQNRYFV 184
+ N+ RW+ GL G EF+D ++ S S ++ FV
Sbjct: 949 ISNQEF---------------ERWEFGIGLEDGVREFLDDLKSNSNKSVTESSPFV 989
>DSPP_RAT (Q62598) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 687
Score = 32.3 bits (72), Expect = 1.5
Identities = 26/81 (32%), Positives = 34/81 (41%), Gaps = 6/81 (7%)
Query: 24 ADLRRLSDVSSGSEHSG-TGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVS 82
+D SD S G G + + DSS S S D + S +D DS+ DS S
Sbjct: 592 SDTSDSSDSSDGDSSDGDSSDSDSSDSDSSNSSDSDSSDSSDSSSSDSSDSDSDSKDSTS 651
Query: 83 DSMDSVEDLLRLSAENANADS 103
DS D + N N+DS
Sbjct: 652 DSSDD-----NSKSGNGNSDS 667
Score = 30.0 bits (66), Expect = 7.4
Identities = 24/80 (30%), Positives = 39/80 (48%), Gaps = 6/80 (7%)
Query: 24 ADLRRLSDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSD 83
+D SD S S+ S + + S+ G + DG+ DS +D DS S SD
Sbjct: 571 SDSSDTSDSSDSSDSSDSSNSSDTSDSSDSSDG--DSSDGDSSDSDSSDSDSSN-SSDSD 627
Query: 84 SMDSVEDLLRLSAENANADS 103
S DS + S++++++DS
Sbjct: 628 SSDSSDS---SSSDSSDSDS 644
>SLA2_YEAST (P33338) SLA2 protein (Transmembrane protein MOP2)
Length = 968
Score = 31.6 bits (70), Expect = 2.5
Identities = 22/75 (29%), Positives = 31/75 (41%), Gaps = 1/75 (1%)
Query: 37 EHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSA 96
EH D+ SL V G +E+D DF SE ++ VE +L+L
Sbjct: 887 EHCSKDVTDACRSLGNHVMGMIEDDHSTSQQQQPLDFTSEHTLKTAEMEQQVE-ILKLEQ 945
Query: 97 ENANADSYLNMLRLH 111
+NA L +R H
Sbjct: 946 SLSNARKRLGEIRRH 960
>RPOC_BUCAP (P41185) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta' chain)
(RNA polymerase beta' subunit)
Length = 1413
Score = 31.2 bits (69), Expect = 3.3
Identities = 25/100 (25%), Positives = 44/100 (44%), Gaps = 23/100 (23%)
Query: 71 NDFDSERV---DSVSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSV 127
N F+ ERV D +SD +S D+LRL A +N ++ + + Q V
Sbjct: 1197 NVFEGERVERGDVISDGPESPHDILRLRGVQAVTKYIVNEVQ----------EVYRLQGV 1246
Query: 128 SVYNRNVMSFLREKGHNAAICKTRWDSSGGLTAGNHEFID 167
+ ++++ +R+ A + K +GN EF+D
Sbjct: 1247 KINDKHIEVIIRQMLRKATVIK----------SGNSEFLD 1276
>RECG_HELPJ (Q9ZJA1) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 623
Score = 31.2 bits (69), Expect = 3.3
Identities = 20/67 (29%), Positives = 34/67 (49%), Gaps = 12/67 (17%)
Query: 74 DSERVDSVSDSMDSVE------------DLLRLSAENANADSYLNMLRLHVSEAAVKFDF 121
++ER++ +D +D + DLL+ ++ N+ Y+++ R A VK DF
Sbjct: 551 ENERLEKFADELDGFKIAELDLQYRKSGDLLQGGKQSGNSFEYIDLARDENIIAEVKQDF 610
Query: 122 LKKQSVS 128
LK SVS
Sbjct: 611 LKNASVS 617
>YCF2_ANTFO (Q859W7) Protein ycf2
Length = 2392
Score = 30.8 bits (68), Expect = 4.3
Identities = 24/124 (19%), Positives = 50/124 (39%), Gaps = 20/124 (16%)
Query: 94 LSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICKTRWD 153
++ N+ Y++ +RL + + F ++ Q + N V L K A I
Sbjct: 1852 INTTQGNSTIYIDAVRLALHRQTLGFTYINNQMLFCQNEGV---LLHKIGKAVI------ 1902
Query: 154 SSGGLTAGNHEFIDVVRMRS---GSSTWQNRYFVELDFAVQFEIARPTSRYSEIMSYVPG 210
+ FI + M GS W+ +++ ++ ++ + PT + ++SY+ G
Sbjct: 1903 --------QNIFIKIFPMNPLYIGSDLWKKKFYYLSEWFLEPSVTEPTIKELTLLSYILG 1954
Query: 211 IFVG 214
G
Sbjct: 1955 CLAG 1958
>HM39_CAEEL (Q22812) Homeobox protein ceh-39
Length = 399
Score = 30.8 bits (68), Expect = 4.3
Identities = 26/73 (35%), Positives = 30/73 (40%), Gaps = 2/73 (2%)
Query: 20 RLVGADLRRLSDV--SSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSER 77
R G D R S + SS S S + SSP+ G EE S FDSE
Sbjct: 50 RNYGTDFGRASSMCGSSSSSSSSSYSSGSSPNYLTENGGDTEEPRFEELVSRKMSFDSEN 109
Query: 78 VDSVSDSMDSVED 90
V VS+ MD D
Sbjct: 110 VAQVSNQMDYARD 122
>DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin
matrix protein-3) (DMP-3) [Contains: Dentin
phosphoprotein (Dentin phosphophoryn) (DPP); Dentin
sialoprotein (DSP)]
Length = 934
Score = 30.8 bits (68), Expect = 4.3
Identities = 19/61 (31%), Positives = 29/61 (47%)
Query: 30 SDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVE 89
SD S S S + + DS S S+ G + +GN ++ ++ DS+ SDS S
Sbjct: 873 SDSSDSSNSSDSSDSDSKDSSSDSSDGDSKSGNGNSDSNSDSNSDSDSDSEGSDSNHSTS 932
Query: 90 D 90
D
Sbjct: 933 D 933
>FRIH_MOUSE (P09528) Ferritin heavy chain (Ferritin H subunit)
Length = 181
Score = 30.4 bits (67), Expect = 5.7
Identities = 18/57 (31%), Positives = 30/57 (52%)
Query: 49 SLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYL 105
SL EL +++D ++CD + SE+V S+ + D V +L ++ A A YL
Sbjct: 113 SLLELHKLATDKNDPHLCDFIETYYLSEQVKSIKELGDHVTNLRKMGAPEAGMAEYL 169
>FHL1_YEAST (P39521) Pre-rRNA processing protein FHL1
Length = 936
Score = 30.4 bits (67), Expect = 5.7
Identities = 37/133 (27%), Positives = 56/133 (41%), Gaps = 13/133 (9%)
Query: 30 SDVSSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVE 89
+D+ S S S + SS S SE G DDG+ S N S +S SDS V+
Sbjct: 810 NDLESESGTSSSSS-SSSESGSESDSG---SDDGSASGSGDNSSTSSESESESDSGSEVD 865
Query: 90 DLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSVSVYNRNVMSFLREKGHNAAICK 149
E N + ++ + +E+ K D K S + +S E+GH++ K
Sbjct: 866 -------EKNNKNEKIDSESIKNNES--KDDIPSKDENSSNDNREISKTDEEGHDSKRRK 916
Query: 150 TRWDSSGGLTAGN 162
D + G+T N
Sbjct: 917 VSEDINEGITEVN 929
>SHP1_YEAST (P34223) SHP1 protein
Length = 423
Score = 30.0 bits (66), Expect = 7.4
Identities = 23/79 (29%), Positives = 36/79 (45%), Gaps = 9/79 (11%)
Query: 33 SSGSEHSGTGECDSSPSLSELVHGFLEEDDGNVCDST--GNDFDSERVDSVSDSMDSVED 90
+ GS SG+G S S++V G ++DD + +T G + V SD ++D
Sbjct: 112 TKGSSRSGSGNNSRFMSFSDMVRGQADDDDEDQPRNTFAGGETSGLEVTDPSDPNSLLKD 171
Query: 91 LL-------RLSAENANAD 102
LL ++ AEN D
Sbjct: 172 LLEKARRGGQMGAENGFRD 190
>FRIH_CRIGR (P29389) Ferritin heavy chain (Ferritin H subunit)
Length = 185
Score = 30.0 bits (66), Expect = 7.4
Identities = 17/57 (29%), Positives = 30/57 (51%)
Query: 49 SLSELVHGFLEEDDGNVCDSTGNDFDSERVDSVSDSMDSVEDLLRLSAENANADSYL 105
SL EL +++D ++CD + +E+V S+ + D V +L ++ A A YL
Sbjct: 118 SLLELHKLATDKNDPHLCDFIETHYLNEQVKSIKELGDHVTNLRKMGAPEAGMAEYL 174
>RPOC_BUCAI (P57145) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta' chain)
(RNA polymerase beta' subunit)
Length = 1407
Score = 29.6 bits (65), Expect = 9.7
Identities = 21/84 (25%), Positives = 39/84 (46%), Gaps = 13/84 (15%)
Query: 71 NDFDSERVDS---VSDSMDSVEDLLRLSAENANADSYLNMLRLHVSEAAVKFDFLKKQSV 127
N F+ ERVD +SD +S D+LRL A +N ++ + + Q V
Sbjct: 1197 NVFEGERVDRGDVISDGPESPHDILRLRGVQAVTRYIVNEVQ----------EVYRLQGV 1246
Query: 128 SVYNRNVMSFLREKGHNAAICKTR 151
+ ++++ +R+ A + K+R
Sbjct: 1247 KINDKHIEVIIRQMLRKATVVKSR 1270
>RECG_HELPY (O26051) ATP-dependent DNA helicase recG (EC 3.6.1.-)
Length = 623
Score = 29.6 bits (65), Expect = 9.7
Identities = 19/67 (28%), Positives = 34/67 (50%), Gaps = 12/67 (17%)
Query: 74 DSERVDSVSDSMDSVE------------DLLRLSAENANADSYLNMLRLHVSEAAVKFDF 121
++ER++ +D +D + DLL+ ++ N+ Y+++ + A VK DF
Sbjct: 551 ENERLEKFADELDGFKIAELDLEYRKSGDLLQGGEQSGNSFEYIDLAKDENIIAEVKRDF 610
Query: 122 LKKQSVS 128
LK SVS
Sbjct: 611 LKAASVS 617
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,823,863
Number of Sequences: 164201
Number of extensions: 1490755
Number of successful extensions: 3650
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 3569
Number of HSP's gapped (non-prelim): 89
length of query: 304
length of database: 59,974,054
effective HSP length: 110
effective length of query: 194
effective length of database: 41,911,944
effective search space: 8130917136
effective search space used: 8130917136
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC145513.12


