

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145512.11 - phase: 0
(127 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
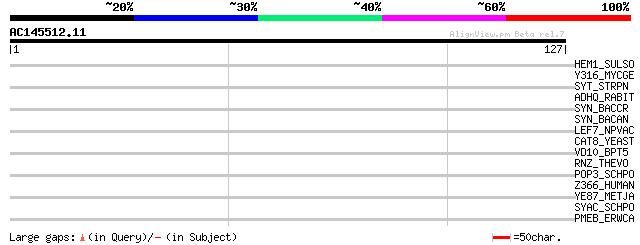
Score E
Sequences producing significant alignments: (bits) Value
HEM1_SULSO (Q980U7) Glutamyl-tRNA reductase (EC 1.2.1.-) (GluTR) 30 1.2
Y316_MYCGE (P47558) Hypothetical protein MG316 29 1.6
SYT_STRPN (Q97PI4) Threonyl-tRNA synthetase (EC 6.1.1.3) (Threon... 28 2.7
ADHQ_RABIT (O46650) Alcohol dehydrogenase class II isozyme 2 (EC... 28 2.7
SYN_BACCR (Q817I8) Asparaginyl-tRNA synthetase (EC 6.1.1.22) (As... 28 3.6
SYN_BACAN (Q81L32) Asparaginyl-tRNA synthetase (EC 6.1.1.22) (As... 28 3.6
LEF7_NPVAC (P41677) Late expression factor 7 28 3.6
CAT8_YEAST (P39113) Regulatory protein CAT8 28 3.6
VD10_BPT5 (P11107) Probable helicase (D10 protein) 28 4.7
RNZ_THEVO (Q979A8) Ribonuclease Z (EC 3.1.26.11) (RNase Z) (tRNA... 28 4.7
POP3_SCHPO (O74184) WD-repeat protein pop3 (WD-repeat protein wat1) 28 4.7
Z366_HUMAN (Q8N895) Zinc finger protein 366 27 6.1
YE87_METJA (Q58882) Hypothetical protein MJ1487 27 6.1
SYAC_SCHPO (O13914) Probable alanyl-tRNA synthetase, cytoplasmic... 27 6.1
PMEB_ERWCA (P55743) Pectinesterase B (EC 3.1.1.11) (Pectin methy... 27 6.1
>HEM1_SULSO (Q980U7) Glutamyl-tRNA reductase (EC 1.2.1.-) (GluTR)
Length = 426
Score = 29.6 bits (65), Expect = 1.2
Identities = 16/43 (37%), Positives = 24/43 (55%), Gaps = 2/43 (4%)
Query: 78 LENKWNHAEVDFGFPFMFSGIHV--LKEKSNMKDIRFTNPEND 118
LENKWN VD P +F+G +V L+E + ++ F E +
Sbjct: 256 LENKWNTLIVDITVPPLFTGNNVITLEELERISNLNFKAREEE 298
>Y316_MYCGE (P47558) Hypothetical protein MG316
Length = 369
Score = 29.3 bits (64), Expect = 1.6
Identities = 26/118 (22%), Positives = 47/118 (39%), Gaps = 19/118 (16%)
Query: 6 WFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQ 65
W+ NK A+ ++ + G+ RV I+ T +F Q
Sbjct: 147 WYLNKLSGFAVLLIYLFLVGFAFSALRVFIS-------------------TLLKQVFKKQ 187
Query: 66 VEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPENDANIVL 123
+ N ++ L+ NHA +FGF F F VL + +K ++ P ++++L
Sbjct: 188 LPEDNLSLTALLIILISNHALNNFGFNFSFLACFVLLFVNKLKLLKALKPLVSSSLIL 245
>SYT_STRPN (Q97PI4) Threonyl-tRNA synthetase (EC 6.1.1.3)
(Threonine--tRNA ligase) (ThrRS)
Length = 647
Score = 28.5 bits (62), Expect = 2.7
Identities = 15/53 (28%), Positives = 22/53 (41%), Gaps = 6/53 (11%)
Query: 42 YTHGRKIDRCSRPDTYH------LHLFHMQVEYFNGNMDKALLENKWNHAEVD 88
Y G +D C P HL H+ Y+ GN D A+++ + A D
Sbjct: 172 YRQGEYVDLCRGPHVPSTGRIQIFHLLHVAGAYWRGNSDNAMMQRIYGTAWFD 224
>ADHQ_RABIT (O46650) Alcohol dehydrogenase class II isozyme 2 (EC
1.1.1.1)
Length = 378
Score = 28.5 bits (62), Expect = 2.7
Identities = 20/81 (24%), Positives = 36/81 (43%), Gaps = 3/81 (3%)
Query: 41 FYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHV 100
+ H RK C P T F E+ N +++ L+++K + + F GI
Sbjct: 94 YVPHCRKCKFCQSPLTNFCTKFS---EHKNPIIEQELMDDKTSRFTCKGKSIYHFLGISA 150
Query: 101 LKEKSNMKDIRFTNPENDANI 121
+ + +KDI ++DAN+
Sbjct: 151 FSQYTVVKDINLAKIDDDANL 171
>SYN_BACCR (Q817I8) Asparaginyl-tRNA synthetase (EC 6.1.1.22)
(Asparagine--tRNA ligase) (AsnRS)
Length = 463
Score = 28.1 bits (61), Expect = 3.6
Identities = 15/39 (38%), Positives = 21/39 (53%)
Query: 65 QVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKE 103
++E+FN +DK +LE N DFG I VL+E
Sbjct: 281 EMEFFNSFVDKTVLERMNNVINSDFGRITYTEAIKVLQE 319
>SYN_BACAN (Q81L32) Asparaginyl-tRNA synthetase (EC 6.1.1.22)
(Asparagine--tRNA ligase) (AsnRS)
Length = 463
Score = 28.1 bits (61), Expect = 3.6
Identities = 15/39 (38%), Positives = 21/39 (53%)
Query: 65 QVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKE 103
++E+FN +DK +LE N DFG I VL+E
Sbjct: 281 EMEFFNSFVDKTVLERMNNVINSDFGRITYTEAIKVLQE 319
>LEF7_NPVAC (P41677) Late expression factor 7
Length = 226
Score = 28.1 bits (61), Expect = 3.6
Identities = 10/27 (37%), Positives = 16/27 (59%)
Query: 44 HGRKIDRCSRPDTYHLHLFHMQVEYFN 70
H R++DRC P T ++L+ +Y N
Sbjct: 140 HERRMDRCVTPSTVQINLYDDNEDYLN 166
>CAT8_YEAST (P39113) Regulatory protein CAT8
Length = 1433
Score = 28.1 bits (61), Expect = 3.6
Identities = 24/96 (25%), Positives = 41/96 (42%), Gaps = 17/96 (17%)
Query: 43 THGRKIDRCSRPDT-------YHLHLFHMQ--VEYFNGNMDKALLENKWNHAEVDFGFPF 93
T+ +D C+R T +L +M ++ N N+ + +ENK H + F P
Sbjct: 1056 TNNGNVDVCNRASTDATDANIENLSFLNMAPFLQTGNSNIGQNTIENKPMHMDAIFSLP- 1114
Query: 94 MFSGIHVLKEKSNMKDIRF-----TNPENDANIVLH 124
S + ++K+ + K + NPEN N H
Sbjct: 1115 --SNLDLMKDNMDSKPEQLEPVIKQNPENSKNNQFH 1148
>VD10_BPT5 (P11107) Probable helicase (D10 protein)
Length = 450
Score = 27.7 bits (60), Expect = 4.7
Identities = 13/51 (25%), Positives = 21/51 (40%)
Query: 32 RVIINDNTFFYTHGRKIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKW 82
+V+I++ +F D CS+ TYH+ + N E KW
Sbjct: 2 KVVISNKAYFKPDDELWDYCSKQTTYHIETMTSKYPIMYKNSGVVAKEIKW 52
>RNZ_THEVO (Q979A8) Ribonuclease Z (EC 3.1.26.11) (RNase Z) (tRNA 3
endonuclease)
Length = 304
Score = 27.7 bits (60), Expect = 4.7
Identities = 14/47 (29%), Positives = 23/47 (48%)
Query: 75 KALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPENDANI 121
K ++++KW+ +D F F G H L ++ + F N D NI
Sbjct: 45 KQIMKSKWSFMSIDNIFITHFHGDHFLGLIGLVQSMSFNNRTKDLNI 91
>POP3_SCHPO (O74184) WD-repeat protein pop3 (WD-repeat protein wat1)
Length = 314
Score = 27.7 bits (60), Expect = 4.7
Identities = 14/41 (34%), Positives = 17/41 (41%), Gaps = 1/41 (2%)
Query: 20 SAYTWGYLEHPARVIINDNTFFYTHGRKIDRC-SRPDTYHL 59
+ Y W L H ++ F H R I RC PD HL
Sbjct: 190 NCYVWRMLNHQGASLLQPVVKFQAHQRYITRCVLSPDVKHL 230
>Z366_HUMAN (Q8N895) Zinc finger protein 366
Length = 744
Score = 27.3 bits (59), Expect = 6.1
Identities = 12/36 (33%), Positives = 18/36 (49%)
Query: 90 GFPFMFSGIHVLKEKSNMKDIRFTNPENDANIVLHS 125
GFP +F G K KS + + +P + + LHS
Sbjct: 66 GFPGVFEGAGSRKRKSMPTKMPYNHPAEEVTLALHS 101
>YE87_METJA (Q58882) Hypothetical protein MJ1487
Length = 426
Score = 27.3 bits (59), Expect = 6.1
Identities = 25/89 (28%), Positives = 38/89 (42%), Gaps = 13/89 (14%)
Query: 35 INDNTFFYTHGRK-IDRCSRPDTYHLHLFHMQVEYFNGNMDKA-------LLENKWNHAE 86
+NDN F YT RK +D P H +E G K + K H +
Sbjct: 145 LNDNEFIYTGRRKPVDLNKYPPFPVKHNKFGHIEITRGCPYKCYFCQTPRIFGKKIRHRD 204
Query: 87 VDFGFPFMFSGIHVLKEKSNMKDIRFTNP 115
V+ ++ + ++ E+ N+KDIRF P
Sbjct: 205 VE----NIYKYVEIMAER-NLKDIRFITP 228
>SYAC_SCHPO (O13914) Probable alanyl-tRNA synthetase, cytoplasmic
(EC 6.1.1.7) (Alanine--tRNA ligase) (AlaRS)
Length = 959
Score = 27.3 bits (59), Expect = 6.1
Identities = 21/71 (29%), Positives = 28/71 (38%), Gaps = 4/71 (5%)
Query: 57 YHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNPE 116
YH F E FN LL ++ N G + I + + D R TN +
Sbjct: 512 YHKGGFQKSTEGFNSGEQLGLLLDRTNFYAEQGGQEYDTGHIVI----DGVADFRVTNVQ 567
Query: 117 NDANIVLHSGF 127
A VLH+GF
Sbjct: 568 VYAGYVLHTGF 578
>PMEB_ERWCA (P55743) Pectinesterase B (EC 3.1.1.11) (Pectin
methylesterase B) (PE B) (Fragment)
Length = 178
Score = 27.3 bits (59), Expect = 6.1
Identities = 17/72 (23%), Positives = 34/72 (46%), Gaps = 4/72 (5%)
Query: 50 RCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGF---PFMFSGIHVLKEKS- 105
R R DT+ ++ +++ EY +A +++ + +VD+ F +F +H S
Sbjct: 1 RIGRQDTFFVNTSNLRNEYVTDRYSRAYIKDSYIEGDVDYVFGRATAVFDRVHFHTVSSR 60
Query: 106 NMKDIRFTNPEN 117
KDI P++
Sbjct: 61 GAKDIHVFAPDS 72
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,472,168
Number of Sequences: 164201
Number of extensions: 674382
Number of successful extensions: 1515
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 1504
Number of HSP's gapped (non-prelim): 17
length of query: 127
length of database: 59,974,054
effective HSP length: 103
effective length of query: 24
effective length of database: 43,061,351
effective search space: 1033472424
effective search space used: 1033472424
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC145512.11


