

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145470.6 + phase: 2 /partial
(753 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
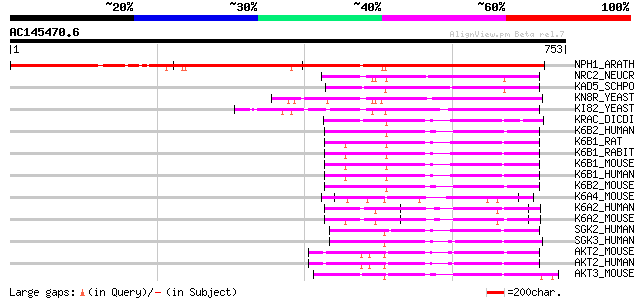
Score E
Sequences producing significant alignments: (bits) Value
NPH1_ARATH (O48963) Nonphototropic hypocotyl protein 1 (EC 2.7.1... 835 0.0
NRC2_NEUCR (O42626) Serine/threonine-protein kinase nrc-2 (EC 2.... 256 1e-67
KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase C4G... 243 1e-63
KN8R_YEAST (P53739) Probable serine/threonine-protein kinase YNR... 239 3e-62
KI82_YEAST (P25341) Probable serine/threonine-protein kinase KIN... 238 3e-62
KRAC_DICDI (P54644) RAC-family serine/threonine-protein kinase h... 180 1e-44
K6B2_HUMAN (Q9UBS0) Ribosomal protein S6 kinase 2 (EC 2.7.1.37) ... 172 2e-42
K6B1_RAT (P67999) Ribosomal protein S6 kinase I (EC 2.7.1.37) (S... 172 2e-42
K6B1_RABIT (P67998) Ribosomal protein S6 kinase I (EC 2.7.1.37) ... 172 2e-42
K6B1_MOUSE (Q8BSK8) Ribosomal protein S6 kinase I (EC 2.7.1.37) ... 172 2e-42
K6B1_HUMAN (P23443) Ribosomal protein S6 kinase 1 (EC 2.7.1.37) ... 172 2e-42
K6B2_MOUSE (Q9Z1M4) Ribosomal protein S6 kinase beta 2 (EC 2.7.1... 170 1e-41
K6A4_MOUSE (Q9Z2B9) Ribosomal protein S6 kinase alpha 4 (EC 2.7.... 169 3e-41
K6A2_HUMAN (Q15349) Ribosomal protein S6 kinase alpha 2 (EC 2.7.... 168 4e-41
K6A2_MOUSE (Q9WUT3) Ribosomal protein S6 kinase alpha 2 (EC 2.7.... 167 7e-41
SGK2_HUMAN (Q9HBY8) Serine/threonine-protein kinase Sgk2 (EC 2.7... 167 9e-41
SGK3_HUMAN (Q96BR1) Serine/threonine-protein kinase Sgk3 (EC 2.7... 166 2e-40
AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC... 166 2e-40
AKT2_HUMAN (P31751) RAC-beta serine/threonine-protein kinase (EC... 166 3e-40
AKT3_MOUSE (Q9WUA6) RAC-gamma serine/threonine-protein kinase (E... 165 4e-40
>NPH1_ARATH (O48963) Nonphototropic hypocotyl protein 1 (EC
2.7.1.37) (Phototropin)
Length = 996
Score = 835 bits (2158), Expect = 0.0
Identities = 438/753 (58%), Positives = 547/753 (72%), Gaps = 43/753 (5%)
Query: 1 RFLQGPETDQNEVAKIRDATKNGKSYCGRLLNYKKNGTPFWNLLTVTPIKDDRGNTIKFI 60
RFLQG TD +E+AKIR+ G +YCGR+LNYKK+GT FWNLLT+ PIKD+ G +KFI
Sbjct: 235 RFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFI 294
Query: 61 GMQVEVSKYTEGVNEKALRPNGLPKSLIRYDARQKEEAMGSITEVVQTVRNPKSIIRSKN 120
GMQVEVSK+TEG EKALRPNGLP+SLIRYDARQK+ A S+TE+V+ V+ P+++ S N
Sbjct: 295 GMQVEVSKHTEGAKEKALRPNGLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTN 354
Query: 121 DDTATIMHEEPENLNHDFVLPKSVEPVNDTTTP-GRQTPLKFHGDNNNMSRFSSYEERNN 179
+H D + K +++ P GR+ G N+M R + E
Sbjct: 355 ------LHPFMTKSESDELPKKPARRMSENVVPSGRRN--SGGGRRNSMQRINEIPE--- 403
Query: 180 KSSRKSGITSLKGVKGKSMSSVGRDKDKTIVE-----PEVLMTKEIEWSKYE-LRERDIR 233
K SRKS + S G+K KS S+ D +E E+ E S + +R++++R
Sbjct: 404 KKSRKSSL-SFMGIKKKS-ESLDESIDDGFIEYGEEDDEISDRDERPESVDDKVRQKEMR 461
Query: 234 QGIDLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEYTREEILGRNCRFLQGPETD 293
+GIDLATTLERIEKNFVI+DPRLPD PIIFASDSFLELTEY+REEILGRNCRFLQGPETD
Sbjct: 462 KGIDLATTLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETD 521
Query: 294 QATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDH 353
TV +IR+AI +Q E+TVQLINYTKSGKKFWN+FHLQPMRDQKGE+QYFIGVQLDGS H
Sbjct: 522 LTTVKKIRNAIDNQTEVTVQLINYTKSGKKFWNIFHLQPMRDQKGEVQYFIGVQLDGSKH 581
Query: 354 LEPLRNRLSEGSEIQSAKLVKATAENVDGAVRELPDANLRPEDLWAIHSQAVSPRPHKRD 413
+EP+RN + E + + LVK TA N+D AVRELPDAN+ PEDLWA HS+ V +PH++D
Sbjct: 582 VEPVRNVIEETAVKEGEDLVKKTAVNIDEAVRELPDANMTPEDLWANHSKVVHCKPHRKD 641
Query: 414 NPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLN 473
+P W+AIQK+ GE IGL HF P++PLG GDTGSVHLVEL GT +L+AMKAM+K+VMLN
Sbjct: 642 SPPWIAIQKVLESGEPIGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLN 701
Query: 474 RNKSVFYYRFIERALKERLYHYWIILFFQHY---------TPHF----------KQPMKI 514
RNK + ER + + L H ++ + + T ++ +QP K+
Sbjct: 702 RNK--VHRARAEREILDLLDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKV 759
Query: 515 LKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQV 574
LKED+ RFYAA+VV+ LEYLHC GIIYRDLKPEN+L+Q +G I L+DFDLS +TSCKPQ+
Sbjct: 760 LKEDAVRFYAAQVVVALEYLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQL 819
Query: 575 VKQSLPGNRRR--SRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLG 632
+ S+ +++ +SQ PIF++EP+ SNSFVGTEEYIAPEII+GA HTSA+DWW LG
Sbjct: 820 LIPSIDEKKKKKQQKSQQTPIFMAEPMRASNSFVGTEEYIAPEIISGAGHTSAVDWWALG 879
Query: 633 ILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGS 692
IL+YEMLYG TPFRGK RQKTF+N+L KDL FP+SIPASL +QLI LLQRDP RLG
Sbjct: 880 ILMYEMLYGYTPFRGKTRQKTFTNVLQKDLKFPASIPASLQVKQLIFRLLQRDPKKRLGC 939
Query: 693 ATGSNEIKQHPFFRGINWPLIRNMSPPPLDVPL 725
G+NE+KQH FF+GINW LIR +PP L+ P+
Sbjct: 940 FEGANEVKQHSFFKGINWALIRCTNPPELETPI 972
Score = 112 bits (281), Expect = 3e-24
Identities = 65/186 (34%), Positives = 102/186 (53%), Gaps = 12/186 (6%)
Query: 223 SKYELRERDI---RQGI-----DLATTLERIEKNFVISDPRLPDCPIIFASDSFLELTEY 274
S E+ + D+ R GI DL L ++ FV+SD PD PI++AS F +T Y
Sbjct: 165 SSGEMSDGDVPGGRSGIPRVSEDLKDALSTFQQTFVVSDATKPDYPIMYASAGFFNMTGY 224
Query: 275 TREEILGRNCRFLQGPETDQATVNRIRDAIKDQREITVQLINYTKSGKKFWNLFHLQPMR 334
T +E++GRNCRFLQG TD + +IR+ + +++NY K G FWNL + P++
Sbjct: 225 TSKEVVGRNCRFLQGSGTDADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIK 284
Query: 335 DQKGELQYFIGVQLDGSDHLEPLRNRLSEGSEIQSAKLVKATAENVD---GAVRELPDAN 391
D+ G++ FIG+Q++ S H E + + + + + L++ A D +V EL +A
Sbjct: 285 DESGKVLKFIGMQVEVSKHTEGAKEKALRPNGLPES-LIRYDARQKDMATNSVTELVEAV 343
Query: 392 LRPEDL 397
RP L
Sbjct: 344 KRPRAL 349
>NRC2_NEUCR (O42626) Serine/threonine-protein kinase nrc-2 (EC
2.7.1.37) (Nonrepressible conidiation protein 2)
Length = 623
Score = 256 bits (655), Expect = 1e-67
Identities = 150/326 (46%), Positives = 194/326 (59%), Gaps = 38/326 (11%)
Query: 423 ITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYR 482
I R ++G F I+ +G GD G V+LV+ + +G LYAMK + K M+ RNK
Sbjct: 230 IKVRNVEVGPQSFDKIKLIGKGDVGKVYLVKEKKSGRLYAMKVLSKKEMIKRNK------ 283
Query: 483 FIERALKER-------------LYHY-----WIILFFQHYT--PHFK----QPMKILKED 518
I+RAL E+ LYH ++ L ++ + F+ +P K + ED
Sbjct: 284 -IKRALAEQEILATSNHPFIVTLYHSFQSEDYLYLCMEYCSGGEFFRALQTRPGKCIPED 342
Query: 519 SARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQS 578
ARFYAAEV LEYLH +G IYRDLKPEN+LL + GHI+L+DFDLS P
Sbjct: 343 DARFYAAEVTAALEYLHLMGFIYRDLKPENILLHQSGHIMLSDFDLS--KQSDPGGKPTM 400
Query: 579 LPGNRRRSRSQPPPIFVSEPVT--QSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLY 636
+ G S S P I + ++NSFVGTEEYIAPE+I G+ HTSA+DWWTLGIL+Y
Sbjct: 401 IIGKNGTSTSSLPTIDTKSCIANFRTNSFVGTEEYIAPEVIKGSGHTSAVDWWTLGILIY 460
Query: 637 EMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPA---SLAARQLINALLQRDPASRLGSA 693
EMLYG TPF+GKNR TF+NIL +D+ FP A S + LI LL +D RLG+
Sbjct: 461 EMLYGTTPFKGKNRNATFANILREDIPFPDHAGAPQISNLCKSLIRKLLIKDENRRLGAR 520
Query: 694 TGSNEIKQHPFFRGINWPLIRNMSPP 719
G+++IK HPFFR W LIR+M PP
Sbjct: 521 AGASDIKTHPFFRTTQWALIRHMKPP 546
>KAD5_SCHPO (Q09831) Probable serine/threonine-protein kinase
C4G8.05 (EC 2.7.1.37)
Length = 566
Score = 243 bits (620), Expect = 1e-63
Identities = 134/315 (42%), Positives = 185/315 (58%), Gaps = 28/315 (8%)
Query: 429 KIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERAL 488
++G F + LG GD G V+LV + +G+ YAMK + K M+ RNKS F E+ +
Sbjct: 189 EVGPSSFEKVFLLGKGDVGRVYLVREKKSGKFYAMKVLSKQEMIKRNKSK--RAFAEQHI 246
Query: 489 KERLYHYWIILFFQHYTPHF-------------------KQPMKILKEDSARFYAAEVVI 529
H +I+ + + ++P + L E+ A+FY AEV
Sbjct: 247 LATSNHPFIVTLYHSFQSDEYLYLCMEYCMGGEFFRALQRRPGRCLSENEAKFYIAEVTA 306
Query: 530 GLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCK--PQVVKQSLPGNRRRSR 587
LEYLH +G IYRDLKPEN+LL + GHI+L+DFDLS ++ P V++ + + +
Sbjct: 307 ALEYLHLMGFIYRDLKPENILLHESGHIMLSDFDLSKQSNSAGAPTVIQARNAPSAQNAY 366
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
+ +++ ++NSFVGTEEYIAPE+I G HTSA+DWWTLGIL YEMLY TPF+G
Sbjct: 367 ALDTKSCIAD--FRTNSFVGTEEYIAPEVIKGCGHTSAVDWWTLGILFYEMLYATTPFKG 424
Query: 648 KNRQKTFSNILHKDLTFPSSIPA---SLAARQLINALLQRDPASRLGSATGSNEIKQHPF 704
KNR TFSNILHKD+ FP A S + LI LL +D RLGS G+ ++K HPF
Sbjct: 425 KNRNMTFSNILHKDVIFPEYADAPSISSLCKNLIRKLLVKDENDRLGSQAGAADVKLHPF 484
Query: 705 FRGINWPLIRNMSPP 719
F+ + W L+R+ PP
Sbjct: 485 FKNVQWALLRHTEPP 499
>KN8R_YEAST (P53739) Probable serine/threonine-protein kinase
YNR047W (EC 2.7.1.37)
Length = 893
Score = 239 bits (609), Expect = 3e-62
Identities = 150/403 (37%), Positives = 214/403 (52%), Gaps = 50/403 (12%)
Query: 356 PLRNRLSEGSEIQSAKLVKAT---AENVDGAVR---ELPDANLRPEDLWAIHSQAVSPRP 409
PLR SE + +Q A T A G+ ++P ++ P + + Q SP+
Sbjct: 408 PLRRAASEPNGLQLASATSPTSSSARKTSGSSNINDKIPGQSVPPPN--SFFPQEPSPKI 465
Query: 410 HKRDNPSWVAIQKITARGEK-----IGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMK 464
P + + K +G F IR LG GD G V LV + T +YA+K
Sbjct: 466 SDFPEPRRSRRLRTKSFSNKFQDIMVGPQSFEKIRLLGQGDVGKVFLVREKKTNRVYALK 525
Query: 465 AMEKSVMLNRNKSVFYYRFIERALKER-------------LYHY-----WIILFFQH--- 503
+ K M+ RNK I+R L E+ LYH ++ L ++
Sbjct: 526 VLSKDEMIKRNK-------IKRVLTEQEILATSNHPFIVTLYHSFQSEDYLYLCMEYCMG 578
Query: 504 ---YTPHFKQPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLT 560
+ + K + ED ARFYA+EV LEYLH LG IYRDLKPEN+LL + GHI+L+
Sbjct: 579 GEFFRALQTRKTKCICEDDARFYASEVTAALEYLHLLGFIYRDLKPENILLHQSGHIMLS 638
Query: 561 DFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGA 620
DFDLS Q +P + ++S + ++NSFVGTEEYIAPE+I G
Sbjct: 639 DFDLSI------QAKDSKVPVVKGSAQSTLVDTKICSDGFRTNSFVGTEEYIAPEVIRGN 692
Query: 621 RHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINA 680
HT+A+DWWTLGIL+YEML+G TPF+G N +TF+NIL +++FP++ S + LI
Sbjct: 693 GHTAAVDWWTLGILIYEMLFGFTPFKGDNTNETFTNILKNEVSFPNNNEISRTCKDLIKK 752
Query: 681 LLQRDPASRLGSATGSNEIKQHPFFRGINWPLIRNMSPPPLDV 723
LL ++ + RLG G+ ++K+HPFF+ + W L+RN PP + V
Sbjct: 753 LLTKNESKRLGCKMGAADVKKHPFFKKVQWSLLRNQEPPLIPV 795
>KI82_YEAST (P25341) Probable serine/threonine-protein kinase KIN82
(EC 2.7.1.37)
Length = 720
Score = 238 bits (608), Expect = 3e-62
Identities = 161/449 (35%), Positives = 230/449 (50%), Gaps = 63/449 (14%)
Query: 305 KDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHLEPLRNRLSEG 364
KD R ++ +N ++ N FH + + G + L S P +N +
Sbjct: 197 KDSRNLSNGSLNDINENEELQN-FHRKI--SENGSASPLANLSLSNSPIDSPRKNSETRK 253
Query: 365 SEIQ---SAKLVKATAENV----DGAVRELPDANLRPEDLWAIHSQAVSPRPHKRDNPSW 417
+I + +L +A +E DG +RE A +P L I V PR +R
Sbjct: 254 DQIPMNITPRLRRAASEPFNTAKDGLMREDYIALKQPPSLGDI----VEPRRSRRLRTKS 309
Query: 418 VA--IQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRN 475
Q IT + F IR LG GD G V+L+ + T +++A+K + K M+ R
Sbjct: 310 FGNKFQDITVEPQS-----FEKIRLLGQGDVGKVYLMRERDTNQIFALKVLNKHEMIKRK 364
Query: 476 KSVFYYRFIERALKER-------------LYHYWIILFFQHYTPHF-----------KQP 511
K I+R L E+ LYH + + + + +
Sbjct: 365 K-------IKRVLTEQEILATSDHPFIVTLYHSFQTKDYLYLCMEYCMGGEFFRALQTRK 417
Query: 512 MKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSF-ITSC 570
K + E+ A+FYA+EVV LEYLH LG IYRDLKPEN+LL + GH++L+DFDLS T
Sbjct: 418 SKCIAEEDAKFYASEVVAALEYLHLLGFIYRDLKPENILLHQSGHVMLSDFDLSIQATGS 477
Query: 571 KPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWT 630
K +K S + + + ++NSFVGTEEY+APE+I G HT+A+DWWT
Sbjct: 478 KKPTMKDSTYLDTK----------ICSDGFRTNSFVGTEEYLAPEVIRGNGHTAAVDWWT 527
Query: 631 LGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRL 690
LGIL+YEML+G TPF+G N +TFSNIL KD+ FP S + LI LL ++ A RL
Sbjct: 528 LGILIYEMLFGCTPFKGDNSNETFSNILTKDVKFPHDKEVSKNCKDLIKKLLNKNEAKRL 587
Query: 691 GSATGSNEIKQHPFFRGINWPLIRNMSPP 719
GS +G+ +IK+HPFF+ + W +RN PP
Sbjct: 588 GSKSGAADIKRHPFFKKVQWSFLRNQDPP 616
>KRAC_DICDI (P54644) RAC-family serine/threonine-protein kinase
homolog (EC 2.7.1.37)
Length = 444
Score = 180 bits (457), Expect = 1e-44
Identities = 110/316 (34%), Positives = 162/316 (50%), Gaps = 50/316 (15%)
Query: 426 RGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIE 485
+ EK+G+ F + +G G G V V + TGE+YAMK + K ++ N+ + E
Sbjct: 111 KSEKVGVADFELLNLVGKGSFGKVIQVRKKDTGEVYAMKVLSKKHIVEHNE--VEHTLSE 168
Query: 486 RALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEVV 528
R + +++ H +++ + K Q K ED R+Y AE+V
Sbjct: 169 RNILQKINHPFLVNLNYSFQTEDKLYFILDYVNGGELFYHLQKDKKFTEDRVRYYGAEIV 228
Query: 529 IGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRS 588
+ LE+LH G+IYRDLKPENLLL +GHI +TDF L CK ++
Sbjct: 229 LALEHLHLSGVIYRDLKPENLLLTNEGHICMTDFGL-----CKEGLL------------- 270
Query: 589 QPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGK 648
P ++ +F GT EY+APE++ G + +DWW+ G LLYEML G PF +
Sbjct: 271 --------TPTDKTGTFCGTPEYLAPEVLQGNGYGKQVDWWSFGSLLYEMLTGLPPFYNQ 322
Query: 649 NRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGI 708
+ Q+ + I+ + L+FP I S AR L+ LL+RDP RL N IK+HPFFR I
Sbjct: 323 DVQEMYRKIMMEKLSFPHFI--SPDARSLLEQLLERDPEKRLAD---PNLIKRHPFFRSI 377
Query: 709 NWPLIRNMSPPPLDVP 724
+W + + PP +P
Sbjct: 378 DWEQLFQKNIPPPFIP 393
>K6B2_HUMAN (Q9UBS0) Ribosomal protein S6 kinase 2 (EC 2.7.1.37)
(S6K2) (70 kDa ribosomal protein S6 kinase 2) (p70-S6KB)
(p70 ribosomal S6 kinase beta) (p70 S6Kbeta) (p70 S6
kinase beta) (S6K-beta) (p70-beta) (S6 kinase-related
kinase) (SRK) (Serine/thre
Length = 482
Score = 172 bits (437), Expect = 2e-42
Identities = 111/315 (35%), Positives = 164/315 (51%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLV-ELQGT--GELYAMKAMEKSVMLNRNKSVFYYRFI 484
E+IG H F +R LG G G V V ++QGT G++YAMK + K+ ++ K + R
Sbjct: 60 ERIGPHCFELLRVLGKGGYGKVFQVRKVQGTNLGKIYAMKVLRKAKIVRNAKDTAHTR-A 118
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 119 ERNILESVKHPFIVELAYAFQTGGKLYLILECLSGGELFTHLEREGIFLEDTACFYLAEI 178
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GHI LTDF L CK + + ++
Sbjct: 179 TLALGHLHSQGIIYRDLKPENIMLSSQGHIKLTDFGL-----CKESIHEGAV-------- 225
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
+++F GT EY+APEI+ + H A+DWW+LG L+Y+ML G PF
Sbjct: 226 --------------THTFCGTIEYMAPEILVRSGHNRAVDWWSLGALMYDMLTGSPPFTA 271
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT I+ L P + AR L+ L+R+P+ R+G G + ++++HPFFR
Sbjct: 272 ENRKKTMDKIIRGKLALPPYLTPD--ARDLVKKFLKRNPSQRIGGGPGDAADVQRHPFFR 329
Query: 707 GINWP--LIRNMSPP 719
+NW L + PP
Sbjct: 330 HMNWDDLLAWRVDPP 344
>K6B1_RAT (P67999) Ribosomal protein S6 kinase I (EC 2.7.1.37) (S6K)
(p70-S6K)
Length = 525
Score = 172 bits (437), Expect = 2e-42
Identities = 114/315 (36%), Positives = 162/315 (51%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVEL---QGTGELYAMKAMEKSVMLNRNKSVFYYRFI 484
EKI F +R LG G G V V TG+++AMK ++K++++ K + +
Sbjct: 84 EKIRPECFELLRVLGKGGYGKVFQVRKVTGANTGKIFAMKVLKKAMIVRNAKDTAHTK-A 142
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 143 ERNILEEVKHPFIVDLIYAFQTGGKLYLILEYLSGGELFMQLEREGIFMEDTACFYLAEI 202
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GH+ LTDF L CK +
Sbjct: 203 SMALGHLHQKGIIYRDLKPENIMLNHQGHVKLTDFGL-----CKESI------------- 244
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
T +++F GT EY+APEI+ + H A+DWW+LG L+Y+ML G PF G
Sbjct: 245 ---------HDGTVTHTFCGTIEYMAPEILMRSGHNRAVDWWSLGALMYDMLTGAPPFTG 295
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT IL L P + + AR L+ LL+R+ ASRLG+ G + E++ HPFFR
Sbjct: 296 ENRKKTIDKILKCKLNLPPYL--TQEARDLLKKLLKRNAASRLGAGPGDAGEVQAHPFFR 353
Query: 707 GINWP--LIRNMSPP 719
INW L R + PP
Sbjct: 354 HINWEELLARKVEPP 368
>K6B1_RABIT (P67998) Ribosomal protein S6 kinase I (EC 2.7.1.37)
(S6K) (p70-S6K)
Length = 525
Score = 172 bits (437), Expect = 2e-42
Identities = 114/315 (36%), Positives = 162/315 (51%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVEL---QGTGELYAMKAMEKSVMLNRNKSVFYYRFI 484
EKI F +R LG G G V V TG+++AMK ++K++++ K + +
Sbjct: 84 EKIRPECFELLRVLGKGGYGKVFQVRKVTGANTGKIFAMKVLKKAMIVRNAKDTAHTK-A 142
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 143 ERNILEEVKHPFIVDLIYAFQTGGKLYLILEYLSGGELFMQLEREGIFMEDTACFYLAEI 202
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GH+ LTDF L CK +
Sbjct: 203 SMALGHLHQKGIIYRDLKPENIMLNHQGHVKLTDFGL-----CKESI------------- 244
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
T +++F GT EY+APEI+ + H A+DWW+LG L+Y+ML G PF G
Sbjct: 245 ---------HDGTVTHTFCGTIEYMAPEILMRSGHNRAVDWWSLGALMYDMLTGAPPFTG 295
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT IL L P + + AR L+ LL+R+ ASRLG+ G + E++ HPFFR
Sbjct: 296 ENRKKTIDKILKCKLNLPPYL--TQEARDLLKKLLKRNAASRLGAGPGDAGEVQAHPFFR 353
Query: 707 GINWP--LIRNMSPP 719
INW L R + PP
Sbjct: 354 HINWEELLARKVEPP 368
>K6B1_MOUSE (Q8BSK8) Ribosomal protein S6 kinase I (EC 2.7.1.37)
(S6K) (p70-S6K)
Length = 525
Score = 172 bits (437), Expect = 2e-42
Identities = 114/315 (36%), Positives = 162/315 (51%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVEL---QGTGELYAMKAMEKSVMLNRNKSVFYYRFI 484
EKI F +R LG G G V V TG+++AMK ++K++++ K + +
Sbjct: 84 EKIRPECFELLRVLGKGGYGKVFQVRKVTGANTGKIFAMKVLKKAMIVRNAKDTAHTK-A 142
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 143 ERNILEEVKHPFIVDLIYAFQTGGKLYLILEYLSGGELFMQLEREGIFMEDTACFYLAEI 202
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GH+ LTDF L CK +
Sbjct: 203 SMALGHLHQKGIIYRDLKPENIMLNHQGHVKLTDFGL-----CKESI------------- 244
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
T +++F GT EY+APEI+ + H A+DWW+LG L+Y+ML G PF G
Sbjct: 245 ---------HDGTVTHTFCGTIEYMAPEILMRSGHNRAVDWWSLGALMYDMLTGAPPFTG 295
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT IL L P + + AR L+ LL+R+ ASRLG+ G + E++ HPFFR
Sbjct: 296 ENRKKTIDKILKCKLNLPPYL--TQEARDLLKKLLKRNAASRLGAGPGDAGEVQAHPFFR 353
Query: 707 GINWP--LIRNMSPP 719
INW L R + PP
Sbjct: 354 HINWEELLARKVEPP 368
>K6B1_HUMAN (P23443) Ribosomal protein S6 kinase 1 (EC 2.7.1.37)
(S6K) (S6K1) (70 kDa ribosomal protein S6 kinase 1) (p70
S6 kinase alpha) (p70(S6K)-alpha) (p70-S6K) (p70-alpha)
Length = 525
Score = 172 bits (437), Expect = 2e-42
Identities = 114/315 (36%), Positives = 162/315 (51%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVEL---QGTGELYAMKAMEKSVMLNRNKSVFYYRFI 484
EKI F +R LG G G V V TG+++AMK ++K++++ K + +
Sbjct: 84 EKIRPECFELLRVLGKGGYGKVFQVRKVTGANTGKIFAMKVLKKAMIVRNAKDTAHTK-A 142
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 143 ERNILEEVKHPFIVDLIYAFQTGGKLYLILEYLSGGELFMQLEREGIFMEDTACFYLAEI 202
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GH+ LTDF L CK +
Sbjct: 203 SMALGHLHQKGIIYRDLKPENIMLNHQGHVKLTDFGL-----CKESI------------- 244
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
T +++F GT EY+APEI+ + H A+DWW+LG L+Y+ML G PF G
Sbjct: 245 ---------HDGTVTHTFCGTIEYMAPEILMRSGHNRAVDWWSLGALMYDMLTGAPPFTG 295
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT IL L P + + AR L+ LL+R+ ASRLG+ G + E++ HPFFR
Sbjct: 296 ENRKKTIDKILKCKLNLPPYL--TQEARDLLKKLLKRNAASRLGAGPGDAGEVQAHPFFR 353
Query: 707 GINWP--LIRNMSPP 719
INW L R + PP
Sbjct: 354 HINWEELLARKVEPP 368
>K6B2_MOUSE (Q9Z1M4) Ribosomal protein S6 kinase beta 2 (EC
2.7.1.37) (S6K-beta 2) (70 kDa ribosomal protein S6
kinase 2) (p70-S6KB) (p70 ribosomal S6 kinase beta) (p70
S6Kbeta) (S6K2)
Length = 485
Score = 170 bits (431), Expect = 1e-41
Identities = 112/315 (35%), Positives = 161/315 (50%), Gaps = 53/315 (16%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLV-ELQGT--GELYAMKAMEKSVMLNRNKSVFYYRFI 484
E+IG H F + LG G G V V ++QGT G++YAMK + K+ ++ K + R
Sbjct: 60 ERIGPHCFELLSVLGKGGYGKVFQVRKVQGTNLGKIYAMKVLRKAKIVCSAKDTAHTR-A 118
Query: 485 ERALKERLYHYWIILFFQHYTPHFK-----------------QPMKILKEDSARFYAAEV 527
ER + E + H +I+ + K + I ED+A FY AE+
Sbjct: 119 ERNILESVKHPFIVELAYAFQTGGKLYLILECLSGGELFTHLEREGIFLEDTACFYLAEI 178
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L +LH GIIYRDLKPEN++L GHI LTDF L CK + + ++
Sbjct: 179 TLALGHLHSHGIIYRDLKPENIMLSSQGHIKLTDFGL-----CKESIHEGAI-------- 225
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
+++F GT EY+APEI+ H A+DWW+LG L+Y+ML G PF
Sbjct: 226 --------------THTFCGTIEYMAPEILVRTGHNRAVDWWSLGALMYDMLTGSPPFTA 271
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATG-SNEIKQHPFFR 706
+NR+KT I+ L P + AR L L+R+P R+G G + ++++HPFFR
Sbjct: 272 ENRKKTMDKIIKGKLVLPPYLTPD--ARDLAKKFLKRNPTQRIGGGLGDAADVQRHPFFR 329
Query: 707 GINWP--LIRNMSPP 719
INW L R + PP
Sbjct: 330 HINWDDLLARRVDPP 344
>K6A4_MOUSE (Q9Z2B9) Ribosomal protein S6 kinase alpha 4 (EC
2.7.1.37) (Nuclear mitogen-and stress-activated protein
kinase-2) (90 kDa ribosomal protein S6 kinase 4)
(RSK-like protein kinase) (RLSK)
Length = 773
Score = 169 bits (427), Expect = 3e-41
Identities = 113/314 (35%), Positives = 164/314 (51%), Gaps = 54/314 (17%)
Query: 423 ITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQG---TGELYAMKAMEKSVMLNRNKSVF 479
+T EK+ + +F+ ++ LG G G V LV G G+LYAMK + K+ ++ R K+
Sbjct: 21 LTGHEEKVSVENFALLKVLGTGAYGKVFLVRKTGGHDAGKLYAMKVLRKAALVQRAKTQE 80
Query: 480 YYRFIERALKE--RLYHYWIILFFQHYTP-----------------HFKQPMKILKEDSA 520
+ R ER++ E R + + L + T H Q + KE
Sbjct: 81 HTR-TERSVLELVRQAPFLVTLHYAFQTDAKLHLILDYVSGGEMFTHLYQ-RQYFKEAEV 138
Query: 521 RFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLP 580
R Y E+V+ LE+LH LGIIYRDLK EN+LL +GHIVLTDF LS
Sbjct: 139 RVYGGEIVLALEHLHKLGIIYRDLKLENVLLDSEGHIVLTDFGLS--------------- 183
Query: 581 GNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEII-TGARHTSAIDWWTLGILLYEML 639
F++E ++ SF GT EY+APEII + A H A+DWW+LGILL+E+L
Sbjct: 184 -----------KEFLTEEKERTFSFCGTIEYMAPEIIRSKAGHGKAVDWWSLGILLFELL 232
Query: 640 YGRTPFRGKNRQKTFSNILHKDLTFPSSIPASL--AARQLINALLQRDPASRLGSA-TGS 696
G +PF + + T + + + L P + A+ L+ LL +DP RLG+ G+
Sbjct: 233 TGASPFTLEGERNTQAEVSRRILKCSPPFPLRIGPVAQDLLQRLLCKDPKKRLGAGPQGA 292
Query: 697 NEIKQHPFFRGINW 710
E+K HPFF+G++W
Sbjct: 293 QEVKSHPFFQGLDW 306
Score = 67.8 bits (164), Expect = 1e-10
Identities = 68/274 (24%), Positives = 119/274 (42%), Gaps = 51/274 (18%)
Query: 441 LGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRF---------IERALKER 491
LG G + +G+ +A+K + + + N + V R + L ++
Sbjct: 417 LGQGSFSVCRRCRQRQSGQEFAVKILSRRLEENTQREVAALRLCQSHPNVVNLHEVLHDQ 476
Query: 492 LYHYWIILFFQ--HYTPHFKQPMKILKEDSARFYAAEVVIGLEYLHC-LGIIYRDLKPEN 548
L+ Y ++ + H ++ ++ E A +V + ++H G+++RDLKPEN
Sbjct: 477 LHTYLVLELLRGGELLEHIRKK-RLFSESEASQILRSLVSAVSFMHEEAGVVHRDLKPEN 535
Query: 549 LLLQKD---GHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSF 605
+L D + + DF + R R Q P +EP+ Q+ F
Sbjct: 536 ILYADDTPGAPVKIIDFGFA-------------------RLRPQSP----AEPM-QTPCF 571
Query: 606 VGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFR---GKNRQKTFSNILHK-- 660
T +Y APE++ + + D W+LG++LY ML G+ PF+ G+ Q + I+ K
Sbjct: 572 --TLQYAAPELLAQQGYDESCDLWSLGVILYMMLSGQVPFQGASGQGGQSQAAEIMCKIR 629
Query: 661 ----DLTFPSSIPASLAARQLINALLQRDPASRL 690
L + S A++L+ LL DPA RL
Sbjct: 630 EGRFSLDGEAWQGVSEEAKELVRGLLTVDPAKRL 663
>K6A2_HUMAN (Q15349) Ribosomal protein S6 kinase alpha 2 (EC
2.7.1.37) (S6K-alpha 2) (90 kDa ribosomal protein S6
kinase 2) (p90-RSK 2) (Ribosomal S6 kinase 3) (RSK-3)
(pp90RSK3)
Length = 733
Score = 168 bits (426), Expect = 4e-41
Identities = 113/315 (35%), Positives = 171/315 (53%), Gaps = 54/315 (17%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLV-ELQGT--GELYAMKAMEKSVMLNRNKSVFYYRFI 484
EK F ++ LG G G V LV +++G+ G+LYAMK ++K+ + R++ +
Sbjct: 52 EKADPSQFELLKVLGQGSYGKVFLVRKVKGSDAGQLYAMKVLKKATLKVRDR---VRSKM 108
Query: 485 ERALKERLYH---------------YWIILFFQHYTPHFKQPMK--ILKEDSARFYAAEV 527
ER + + H ++IL F F + K + E+ +FY AE+
Sbjct: 109 ERDILAEVNHPFIVKLHYAFQTEGKLYLILDFLRGGDLFTRLSKEVMFTEEDVKFYLAEL 168
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L++LH LGIIYRDLKPEN+LL ++GHI +TDF LS K+++ ++R
Sbjct: 169 ALALDHLHSLGIIYRDLKPENILLDEEGHIKITDFGLS----------KEAIDHDKR--- 215
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
+ SF GT EY+APE++ HT + DWW+ G+L++EML G PF+G
Sbjct: 216 --------------AYSFCGTIEYMAPEVVNRRGHTQSADWWSFGVLMFEMLTGSLPFQG 261
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSA-TGSNEIKQHPFFR 706
K+R++T + IL L P + S A+ L+ AL +R+P +RLG+ G EIK+HPFF
Sbjct: 262 KDRKETMALILKAKLGMPQFL--SGEAQSLLRALFKRNPCNRLGAGIDGVEEIKRHPFFV 319
Query: 707 GINW-PLIRNMSPPP 720
I+W L R PP
Sbjct: 320 TIDWNTLYRKEIKPP 334
Score = 59.3 bits (142), Expect = 4e-08
Identities = 51/180 (28%), Positives = 81/180 (44%), Gaps = 34/180 (18%)
Query: 531 LEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQP 590
++YLH G+++RDLKP N+L + + I C KQ GN
Sbjct: 520 MDYLHSQGVVHRDLKPSNILYRDESG------SPESIRVCDFGFAKQLRAGNG------- 566
Query: 591 PPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNR 650
+ P +N ++APE++ + +A D W+LGILLY ML G TPF
Sbjct: 567 ---LLMTPCYTAN-------FVAPEVLKRQGYDAACDIWSLGILLYTMLAGFTPF-ANGP 615
Query: 651 QKTFSNILHK------DLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPF 704
T IL + L+ + S AA+ +++ +L DP RL + ++ +HP+
Sbjct: 616 DDTPEEILARIGSGKYALSGGNWDSISDAAKDVVSKMLHVDPHQRLTAM----QVLKHPW 671
>K6A2_MOUSE (Q9WUT3) Ribosomal protein S6 kinase alpha 2 (EC
2.7.1.37) (S6K-alpha 2) (90 kDa ribosomal protein S6
kinase 2) (p90-RSK 2) (Ribosomal S6 kinase 3) (RSK-3)
(pp90RSK3) (Protein-tyrosine kinase Mpk-9) (MAP
kinase-activated protein kinase 1c) (MA
Length = 733
Score = 167 bits (424), Expect = 7e-41
Identities = 112/315 (35%), Positives = 165/315 (51%), Gaps = 54/315 (17%)
Query: 428 EKIGLHHFSPIRPLGCGDTGSVHLVEL---QGTGELYAMKAMEKSVMLNRNKSVFYYRFI 484
EK F ++ LG G G V LV G+LYAMK ++K+ + R++ +
Sbjct: 52 EKADPSQFELLKVLGQGSYGKVFLVRKVTGSDAGQLYAMKVLKKATLKVRDR---VRSKM 108
Query: 485 ERALKERLYH---------------YWIILFFQHYTPHFKQPMK--ILKEDSARFYAAEV 527
ER + + H ++IL F F + K + E+ +FY AE+
Sbjct: 109 ERDILAEVNHPFIVKLHYAFQTEGKLYLILDFLRGGDLFTRLSKEVMFTEEDVKFYLAEL 168
Query: 528 VIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSR 587
+ L++LH LGIIYRDLKPEN+LL ++GHI +TDF LS K++ ++R
Sbjct: 169 ALALDHLHGLGIIYRDLKPENILLDEEGHIKITDFGLS----------KEATDHDKR--- 215
Query: 588 SQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRG 647
+ SF GT EY+APE++ HT + DWW+ G+L++EML G PF+G
Sbjct: 216 --------------AYSFCGTIEYMAPEVVNRRGHTQSADWWSFGVLMFEMLTGSLPFQG 261
Query: 648 KNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSAT-GSNEIKQHPFFR 706
K+R++T + IL L P + A A+ L+ AL +R+P +RLG+ G EIK+HPFF
Sbjct: 262 KDRKETMALILKAKLGMPQFLSAE--AQSLLRALFKRNPCNRLGAGVDGVEEIKRHPFFV 319
Query: 707 GINW-PLIRNMSPPP 720
I+W L R PP
Sbjct: 320 TIDWNKLYRKEIKPP 334
Score = 58.9 bits (141), Expect = 5e-08
Identities = 48/180 (26%), Positives = 83/180 (45%), Gaps = 34/180 (18%)
Query: 531 LEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQP 590
++YLH G+++RDLKP N+L + S P+ ++ G ++ R++
Sbjct: 520 MDYLHSQGVVHRDLKPSNILYMDE--------------SGNPESIRICDFGFAKQLRAEN 565
Query: 591 PPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNR 650
+ T ++APE++ + +A D W+LGILLY ML G TPF
Sbjct: 566 GLLMTP---------CYTANFVAPEVLKRQGYDAACDVWSLGILLYTMLAGFTPF-ANGP 615
Query: 651 QKTFSNILHK------DLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPF 704
T IL + L+ + S AA+ +++ +L DP RL + ++ +HP+
Sbjct: 616 DDTPEEILARIGSGKYALSGGNWDSISDAAKDVVSKMLHVDPQQRLTAV----QVLKHPW 671
>SGK2_HUMAN (Q9HBY8) Serine/threonine-protein kinase Sgk2 (EC
2.7.1.37) (Serum/glucocorticoid regulated kinase 2)
Length = 427
Score = 167 bits (423), Expect = 9e-41
Identities = 106/304 (34%), Positives = 152/304 (49%), Gaps = 49/304 (16%)
Query: 435 FSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERALKERLYH 494
F ++ +G G+ G V L + + G YA+K ++K +L + + LK +
Sbjct: 95 FDFLKVIGKGNYGKVLLAKRKSDGAFYAVKVLQKKSILKKKEQSHIMAERSVLLKNVRHP 154
Query: 495 YWIILFFQHYTP-----------------HFKQPMKILKEDSARFYAAEVVIGLEYLHCL 537
+ + L + TP H ++ + L E ARFYAAEV + YLH L
Sbjct: 155 FLVGLRYSFQTPEKLYFVLDYVNGGELFFHLQRERRFL-EPRARFYAAEVASAIGYLHSL 213
Query: 538 GIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSE 597
IIYRDLKPEN+LL GH+VLTDF L CK V E
Sbjct: 214 NIIYRDLKPENILLDCQGHVVLTDFGL-----CKEGV----------------------E 246
Query: 598 PVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNI 657
P +++F GT EY+APE++ + A+DWW LG +LYEML+G PF ++ + + NI
Sbjct: 247 PEDTTSTFCGTPEYLAPEVLRKEPYDRAVDWWCLGAVLYEMLHGLPPFYSQDVSQMYENI 306
Query: 658 LHKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINWPLI--RN 715
LH+ L P ++AA L+ +LL +D RLGS EIK H FF INW + +
Sbjct: 307 LHQPLQIPGG--RTVAACDLLQSLLHKDQRQRLGSKADFLEIKNHVFFSPINWDDLYHKR 364
Query: 716 MSPP 719
++PP
Sbjct: 365 LTPP 368
>SGK3_HUMAN (Q96BR1) Serine/threonine-protein kinase Sgk3 (EC
2.7.1.37) (Serum/glucocorticoid regulated kinase 3)
(Serum/glucocorticoid regulated kinase-like)
Length = 496
Score = 166 bits (421), Expect = 2e-40
Identities = 103/305 (33%), Positives = 153/305 (49%), Gaps = 46/305 (15%)
Query: 435 FSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVMLNRNKSVFYYRFIERALKERLYH 494
F ++ +G G G V L + + G+ YA+K ++K ++LNR + LK +
Sbjct: 162 FDFLKVIGKGSFGKVLLAKRKLDGKFYAVKVLQKKIVLNRKEQKHIMAERNVLLKNVKHP 221
Query: 495 YWIILFFQHYTPH----------------FKQPMKILKEDSARFYAAEVVIGLEYLHCLG 538
+ + L + T Q + E ARFYAAE+ L YLH +
Sbjct: 222 FLVGLHYSFQTTEKLYFVLDFVNGGELFFHLQRERSFPEHRARFYAAEIASALGYLHSIK 281
Query: 539 IIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVVKQSLPGNRRRSRSQPPPIFVSEP 598
I+YRDLKPEN+LL GH+VLTDF L CK + +S+
Sbjct: 282 IVYRDLKPENILLDSVGHVVLTDFGL-----CKEGIA-------------------ISDT 317
Query: 599 VTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNIL 658
T +F GT EY+APE+I + + +DWW LG +LYEMLYG PF ++ + + NIL
Sbjct: 318 TT---TFCGTPEYLAPEVIRKQPYDNTVDWWCLGAVLYEMLYGLPPFYCRDVAEMYDNIL 374
Query: 659 HKDLTFPSSIPASLAARQLINALLQRDPASRLGSATGSNEIKQHPFFRGINW-PLIRNMS 717
HK L+ + SL A ++ LL++D +RLG+ EI+ HPFF ++W L++
Sbjct: 375 HKPLSLRPGV--SLTAWSILEELLEKDRQNRLGAKEDFLEIQNHPFFESLSWADLVQKKI 432
Query: 718 PPPLD 722
PPP +
Sbjct: 433 PPPFN 437
>AKT2_MOUSE (Q60823) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 166 bits (420), Expect = 2e-40
Identities = 110/333 (33%), Positives = 169/333 (50%), Gaps = 52/333 (15%)
Query: 406 SPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKA 465
SP VA+ K A K+ ++ F ++ LG G G V LV + TG YAMK
Sbjct: 126 SPSDSSTSEMMEVAVNKARA---KVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKI 182
Query: 466 MEKSVMLNRNK---SVFYYRFIER-------ALKERLYHYWIILFFQHYTP------HFK 509
+ K V++ +++ +V R ++ ALK + + F Y H
Sbjct: 183 LRKEVIIAKDEVAHTVTESRVLQNTRHPFLTALKYAFQTHDRLCFVMEYANGGELFFHLS 242
Query: 510 QPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITS 569
+ ++ ED ARFY AE+V LEYLH ++YRD+K ENL+L KDGHI +TDF L
Sbjct: 243 RE-RVFTEDRARFYGAEIVSALEYLHSRDVVYRDIKLENLMLDKDGHIKITDFGL----- 296
Query: 570 CKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWW 629
CK +S+ T +F GT EY+APE++ + A+DWW
Sbjct: 297 CKEG---------------------ISDGATM-KTFCGTPEYLAPEVLEDNDYGRAVDWW 334
Query: 630 TLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASR 689
LG+++YEM+ GR PF ++ ++ F IL +++ FP ++ A+ L+ LL++DP R
Sbjct: 335 GLGVVMYEMMCGRLPFYNQDHERLFELILMEEIRFPRTLGPE--AKSLLAGLLKKDPKQR 392
Query: 690 LGSA-TGSNEIKQHPFFRGINWPLI--RNMSPP 719
LG + + E+ +H FF INW + + + PP
Sbjct: 393 LGGGPSDAKEVMEHRFFLSINWQDVVQKKLLPP 425
>AKT2_HUMAN (P31751) RAC-beta serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-beta) (Protein kinase Akt-2) (Protein
kinase B, beta) (PKB beta)
Length = 481
Score = 166 bits (419), Expect = 3e-40
Identities = 110/333 (33%), Positives = 170/333 (51%), Gaps = 52/333 (15%)
Query: 406 SPRPHKRDNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKA 465
SP VA+ K A K+ ++ F ++ LG G G V LV + TG YAMK
Sbjct: 126 SPSDSSTTEEMEVAVSKARA---KVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAMKI 182
Query: 466 MEKSVMLNRNK---SVFYYRFIER-------ALKERLYHYWIILFFQHYTP------HFK 509
+ K V++ +++ +V R ++ ALK + + F Y H
Sbjct: 183 LRKEVIIAKDEVAHTVTESRVLQNTRHPFLTALKYAFQTHDRLCFVMEYANGGELFFHLS 242
Query: 510 QPMKILKEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITS 569
+ ++ E+ ARFY AE+V LEYLH ++YRD+K ENL+L KDGHI +TDF L
Sbjct: 243 RE-RVFTEERARFYGAEIVSALEYLHSRDVVYRDIKLENLMLDKDGHIKITDFGL----- 296
Query: 570 CKPQVVKQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWW 629
CK +S+ T +F GT EY+APE++ + A+DWW
Sbjct: 297 CKEG---------------------ISDGATM-KTFCGTPEYLAPEVLEDNDYGRAVDWW 334
Query: 630 TLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASR 689
LG+++YEM+ GR PF ++ ++ F IL +++ FP ++ S A+ L+ LL++DP R
Sbjct: 335 GLGVVMYEMMCGRLPFYNQDHERLFELILMEEIRFPRTL--SPEAKSLLAGLLKKDPKQR 392
Query: 690 LGSA-TGSNEIKQHPFFRGINWPLI--RNMSPP 719
LG + + E+ +H FF INW + + + PP
Sbjct: 393 LGGGPSDAKEVMEHRFFLSINWQDVVQKKLLPP 425
>AKT3_MOUSE (Q9WUA6) RAC-gamma serine/threonine-protein kinase (EC
2.7.1.37) (RAC-PK-gamma) (Protein kinase Akt-3) (Protein
kinase B, gamma) (PKB gamma)
Length = 479
Score = 165 bits (418), Expect = 4e-40
Identities = 112/363 (30%), Positives = 180/363 (48%), Gaps = 62/363 (17%)
Query: 413 DNPSWVAIQKITARGEKIGLHHFSPIRPLGCGDTGSVHLVELQGTGELYAMKAMEKSVML 472
DN + T ++ ++ F ++ LG G G V LV + +G+ YAMK ++K V++
Sbjct: 126 DNIGEEEMDASTTHHKRKTMNDFDYLKLLGKGTFGKVILVREKASGKYYAMKILKKEVII 185
Query: 473 NRNKSVFYYRFIERALKERLYHYWIILFFQHYTP-----------------HFKQPMKIL 515
+++ V + R LK + + L + T H + ++
Sbjct: 186 AKDE-VAHTLTESRVLKNTRHPFLTSLKYSFQTKDRLCFVMEYVNGGELFFHLSRE-RVF 243
Query: 516 KEDSARFYAAEVVIGLEYLHCLGIIYRDLKPENLLLQKDGHIVLTDFDLSFITSCKPQVV 575
ED RFY AE+V L+YLH I+YRDLK ENL+L KDGHI +TDF L CK +
Sbjct: 244 SEDRTRFYGAEIVSALDYLHSGKIVYRDLKLENLMLDKDGHIKITDFGL-----CKEGIT 298
Query: 576 KQSLPGNRRRSRSQPPPIFVSEPVTQSNSFVGTEEYIAPEIITGARHTSAIDWWTLGILL 635
+ +F GT EY+APE++ + A+DWW LG+++
Sbjct: 299 DAAT----------------------MKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVM 336
Query: 636 YEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQLINALLQRDPASRLGSA-T 694
YEM+ GR PF ++ +K F IL +D+ FP ++ S A+ L++ LL +DP RLG
Sbjct: 337 YEMMCGRLPFYNQDHEKLFELILMEDIKFPRTL--SSDAKSLLSGLLIKDPNKRLGGGPD 394
Query: 695 GSNEIKQHPFFRGINWPLI--RNMSPP-----PLDVPLQFIGKDPTAK------DKKWED 741
+ EI +H FF G+NW + + + PP + ++ ++ TA+ +K++D
Sbjct: 395 DAKEIMRHSFFSGVNWQDVYDKKLVPPFKPQVTSETDTRYFDEEFTAQTITITPPEKYDD 454
Query: 742 DGV 744
DG+
Sbjct: 455 DGM 457
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.136 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 92,346,972
Number of Sequences: 164201
Number of extensions: 4095134
Number of successful extensions: 13738
Number of sequences better than 10.0: 1556
Number of HSP's better than 10.0 without gapping: 1274
Number of HSP's successfully gapped in prelim test: 282
Number of HSP's that attempted gapping in prelim test: 10151
Number of HSP's gapped (non-prelim): 2593
length of query: 753
length of database: 59,974,054
effective HSP length: 118
effective length of query: 635
effective length of database: 40,598,336
effective search space: 25779943360
effective search space used: 25779943360
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC145470.6


