

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.12 - phase: 0
(419 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
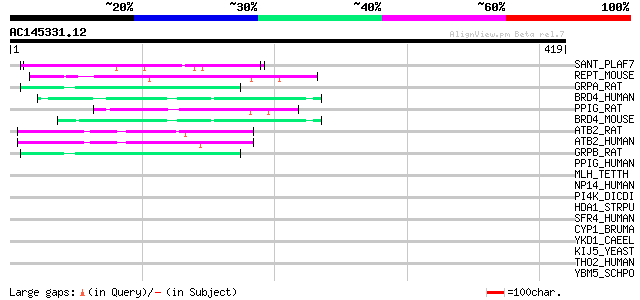
Sequences producing significant alignments: (bits) Value
SANT_PLAF7 (Q03400) S-antigen protein precursor 54 6e-07
REPT_MOUSE (P97347) Repetin 51 6e-06
GRPA_RAT (P08568) Submandibular gland secretory Glx-rich protein... 45 3e-04
BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1 prot... 45 3e-04
PPIG_RAT (O55035) Peptidyl-prolyl cis-trans isomerase G (EC 5.2.... 45 5e-04
BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic ch... 45 5e-04
ATB2_RAT (P11506) Plasma membrane calcium-transporting ATPase 2 ... 44 6e-04
ATB2_HUMAN (Q01814) Plasma membrane calcium-transporting ATPase ... 44 6e-04
GRPB_RAT (P08462) Submandibular gland secretory Glx-rich protein... 44 8e-04
PPIG_HUMAN (Q13427) Peptidyl-prolyl cis-trans isomerase G (EC 5.... 44 0.001
MLH_TETTH (P40631) Micronuclear linker histone polyprotein (MIC ... 42 0.004
NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130... 41 0.005
PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67) ... 41 0.007
HDA1_STRPU (P56518) Histone deacetylase 1 (HD1) 40 0.009
SFR4_HUMAN (Q08170) Splicing factor, arginine/serine-rich 4 (Pre... 40 0.015
CYP1_BRUMA (Q27450) Peptidylprolyl isomerase 1 (EC 5.2.1.8) (Pep... 40 0.015
YKD1_CAEEL (Q03560) Hypothetical protein B0464.2 in chromosome III 39 0.032
KIJ5_YEAST (P40494) Probable serine/threonine-protein kinase YIL... 39 0.032
THO2_HUMAN (Q8NI27) THO complex subunit 2 (Tho2) 38 0.042
YBM5_SCHPO (Q10337) Hypothetical protein C582.05c in chromosome II 37 0.072
>SANT_PLAF7 (Q03400) S-antigen protein precursor
Length = 593
Score = 54.3 bits (129), Expect = 6e-07
Identities = 48/186 (25%), Positives = 78/186 (41%), Gaps = 12/186 (6%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 298 SNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 357
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E ++ E + G + N + S V
Sbjct: 358 EDEVSNGREDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKV 417
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N G+D+ VSNG + + ++ SN R+ ++S + + G DKV
Sbjct: 418 SNG--GEDE-----VSNGR---EDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDKVSN 467
Query: 187 EGESSL 192
GE +
Sbjct: 468 GGEDEV 473
Score = 52.4 bits (124), Expect = 2e-06
Identities = 43/183 (23%), Positives = 76/183 (41%), Gaps = 4/183 (2%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 378 SNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 437
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS +E K+ E + G + N + S V
Sbjct: 438 EDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEV 497
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
N + +DK ++ ++ + + ++ SN R+ ++S + + G D+V
Sbjct: 498 SNGR--EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDEVSN 555
Query: 187 EGE 189
E
Sbjct: 556 GRE 558
Score = 51.2 bits (121), Expect = 5e-06
Identities = 50/185 (27%), Positives = 73/185 (39%), Gaps = 18/185 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 194 SNGREDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGG 253
Query: 69 MKEI-EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
E+ G D V E EV E S ++ V N S+ G + V
Sbjct: 254 EDEVSNGREDKVSNGG-----EDEVSNGREDEVSNGREDEVSNGREDKVSNGGEDE--VS 306
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGE 187
N + K VSNG S R+ + SN R+ ++S + G DKV
Sbjct: 307 NGREDK-------VSNGGEDEVSNGRE---DEVSNGREDKVSNGGEDEVSNGREDKVSNG 356
Query: 188 GESSL 192
GE +
Sbjct: 357 GEDEV 361
Score = 51.2 bits (121), Expect = 5e-06
Identities = 50/202 (24%), Positives = 82/202 (39%), Gaps = 20/202 (9%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENI-NQLESTATGNS 67
++G +D NG DE SN GED VSN + V+ E V +G + + N E +
Sbjct: 106 SNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGR 165
Query: 68 AMKEIEGSNDNV---------DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSS 118
K G D V +G VS +E ++ E + G + N +
Sbjct: 166 EDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDEV 225
Query: 119 SGVNASLVKNSKIGKDKQAS---PAVSNG-----TSALDSRPRQPIKNRSSNDRQSQLSK 170
S V N G+D+ ++ VSNG ++ + + ++ SN R+ ++S
Sbjct: 226 SNGREDKVSNG--GEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSN 283
Query: 171 KKPKSLKKGPLDKVQGEGESSL 192
+ + G DKV GE +
Sbjct: 284 GREDEVSNGREDKVSNGGEDEV 305
Score = 50.8 bits (120), Expect = 6e-06
Identities = 47/192 (24%), Positives = 76/192 (39%), Gaps = 8/192 (4%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 338 SNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGREDEVSNGREDKVSNGG 397
Query: 69 MKEI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
E+ G D V +G VS E ++ E + G + N + S V
Sbjct: 398 EDEVSNGREDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEV 457
Query: 127 KNSKIGK-DKQASPAVSNG-----TSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGP 180
N + K VSNG ++ + + ++ SN R+ ++S + G
Sbjct: 458 SNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGR 517
Query: 181 LDKVQGEGESSL 192
DKV GE +
Sbjct: 518 EDKVSNGGEDEV 529
Score = 49.7 bits (117), Expect = 1e-05
Identities = 48/187 (25%), Positives = 76/187 (39%), Gaps = 18/187 (9%)
Query: 11 GLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMK 70
G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 100 GTEDEVSNGREDKVSNGGEDEVSNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGED 159
Query: 71 EI-EGSNDNV-DGSNLTVSKEKEVKIKVSTEQ--SRAQKGPVKN-KNAKVGSSSGVNASL 125
E+ G D V +G VS +E K+ E S ++ V N + KV + S
Sbjct: 160 EVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGGEDEVSN 219
Query: 126 VKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQ 185
+ ++ ++ VSNG ++ SN R+ ++S + G DKV
Sbjct: 220 GREDEVSNGRE--DKVSNGG-----------EDEVSNGREDKVSNGGEDEVSNGREDKVS 266
Query: 186 GEGESSL 192
GE +
Sbjct: 267 NGGEDEV 273
Score = 49.7 bits (117), Expect = 1e-05
Identities = 47/184 (25%), Positives = 75/184 (40%), Gaps = 16/184 (8%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 154 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDKVSNGG 213
Query: 69 MKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN 128
E+ ++ ++ +E +V E S ++ V N G N K
Sbjct: 214 EDEVSNGRED----EVSNGREDKVSNGGEDEVSNGREDKVSNG----GEDEVSNGREDKV 265
Query: 129 SKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEG 188
S G+D+ VSNG S R+ + SN R+ ++S + G DKV G
Sbjct: 266 SNGGEDE-----VSNGREDEVSNGRE---DEVSNGREDKVSNGGEDEVSNGREDKVSNGG 317
Query: 189 ESSL 192
E +
Sbjct: 318 EDEV 321
Score = 38.5 bits (88), Expect = 0.032
Identities = 25/93 (26%), Positives = 41/93 (43%), Gaps = 4/93 (4%)
Query: 9 ADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSA 68
++G +D NG D+ SN GED VSN + V+ E V +G + ++ N
Sbjct: 490 SNGGEDEVSNGREDKVSNGGEDEVSNGREDKVSNGGEDEVSNGREDEVSNGREDEVSNGR 549
Query: 69 MKEIEGSNDNVDGS----NLTVSKEKEVKIKVS 97
E+ ++ G+ L+ + E K K S
Sbjct: 550 EDEVSNGREDKGGAGTDGELSHNSESHTKNKKS 582
>REPT_MOUSE (P97347) Repetin
Length = 1130
Score = 50.8 bits (120), Expect = 6e-06
Identities = 53/231 (22%), Positives = 98/231 (41%), Gaps = 26/231 (11%)
Query: 16 HQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGS 75
HQN HD+ +D+ +D DPH +ETF D + G+S ++ + S
Sbjct: 142 HQNVQHDQSQRQDKDSERHDTDPHCG-QSETFHGDSH-----------YGHSERQDTDYS 189
Query: 76 NDNVDGSNLTVSKEKEVKIKVSTEQSRAQ---------KGPVKNKNAKVGSSSGVNASLV 126
+D + N + S + + K S EQ + Q K P + + +SG +S
Sbjct: 190 SDQSESDNESSSSSQRLGYKSSHEQPKGQGYVFALSQSKNPEQAFHYGQSKTSGQQSSHG 249
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPL---DK 183
++ + KD +S + + + Q K+ +S + + + ++ KKG ++
Sbjct: 250 QSGRFRKDSYSSQTSQQESDSYEQYGSQHQKSGNSQTERQGQNSQYGQTNKKGHSSYHEQ 309
Query: 184 VQGEGESSLYPFSLRKGES--SSLRLRKRFRQRRRRRVACKQNPRRVKKPR 232
+G+G+S Y RK +S + RK ++ Q+P R +K R
Sbjct: 310 TEGQGQSFHYGQKGRKDQSFQQGQKGRKDQSPHLGQKGRQDQSPHRGQKGR 360
>GRPA_RAT (P08568) Submandibular gland secretory Glx-rich protein CA
precursor (GRP-CA)
Length = 246
Score = 45.4 bits (106), Expect = 3e-04
Identities = 34/167 (20%), Positives = 66/167 (39%), Gaps = 9/167 (5%)
Query: 9 ADGLDDVHQNG-VHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNS 67
A G D+ N D P++S + V + DP D ++EN+ + ES A N
Sbjct: 16 AQGTDEEVNNAETSDVPADSEQQPVDSGSDPPSA--------DADAENVQEGESAAPANE 67
Query: 68 AMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
GS + T ++ +E +E+ + Q+ P + +N + ++SG +
Sbjct: 68 EPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQ 127
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPK 174
+ Q PA S + +N+ +D + + +P+
Sbjct: 128 QEPTQAEDQQPPATSGSEEEQQQQESTQAENQEPSDSAGEGQETQPE 174
>BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1
protein)
Length = 1362
Score = 45.4 bits (106), Expect = 3e-04
Identities = 52/214 (24%), Positives = 81/214 (37%), Gaps = 26/214 (12%)
Query: 22 DEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDG 81
DEP E+ V P V T+ P +S++ + S + + S D+ +
Sbjct: 459 DEP----EEPVVAVSSPAVPPPTKVVAPPSSSDSSSDSSSDS---------DSSTDDSEE 505
Query: 82 SNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAV 141
E + ++K EQ A P +NK K K K K K+
Sbjct: 506 ERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKRKEEVE 558
Query: 142 SNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGE 201
N S P P K + +N S +SKK+P +K P + E E P S +
Sbjct: 559 ENKKSKAKEPP--PKKTKKNNSSNSNVSKKEPAPMKSKPPPTYESEEEDKCKPMSYEEKR 616
Query: 202 SSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
SL + K ++ R V Q+ ++P LK+
Sbjct: 617 QLSLDINKLPGEKLGRVVHIIQS----REPSLKN 646
>PPIG_RAT (O55035) Peptidyl-prolyl cis-trans isomerase G (EC
5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G)
(Rotamase G) (Cyclophilin G) (Matrin cyclophilin)
(Matrin-cyp)
Length = 752
Score = 44.7 bits (104), Expect = 5e-04
Identities = 40/170 (23%), Positives = 71/170 (41%), Gaps = 24/170 (14%)
Query: 64 TGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNA 123
+G ++EIE N D ++ ++ + + +S+ +K K + SSS
Sbjct: 146 SGQEVVREIE--NQKTDAASKPFAEVRILSCGELVPKSKVKKEEKKRHKSSSSSSSS--- 200
Query: 124 SLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGP--- 180
+S D Q+S S+ +A + + R+ K N R+ + KKK K KK P
Sbjct: 201 ----DSDSSSDSQSSSDSSDSETASEEKSRKRKKKHRKNSRKHKKEKKKRKKSKKSPSSE 256
Query: 181 --LDKVQGEGESSLYP----------FSLRKGESSSLRLRKRFRQRRRRR 218
D V + +S++ P F +RK + ++ R+R R R
Sbjct: 257 SEADNVDAQPQSTVRPEEIPPIPENRFLMRKSPPKADDKERKNRERERER 306
>BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic
chromosome-associated protein) (MCAP)
Length = 1400
Score = 44.7 bits (104), Expect = 5e-04
Identities = 47/199 (23%), Positives = 79/199 (39%), Gaps = 15/199 (7%)
Query: 37 DPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKV 96
+P VTV++ P ++ + S+ + + + + + S D+ + E + ++K
Sbjct: 464 EPVVTVSSPAVPPP--TKVVAPPSSSDSSSDSSSDSDSSTDDSEEERAQRLAELQEQLKA 521
Query: 97 STEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPI 156
EQ A P +NK K K K K K+ N S P P
Sbjct: 522 VHEQLAALSQPQQNKPKKKEKDK-------KEKKKEKHKKKEEVEENKKSKTKELP--PK 572
Query: 157 KNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRR 216
K + +N S +SKK+P K P + E E P S + SL + K ++
Sbjct: 573 KTKKNNSSNSNVSKKEPVPTKTKPPPTYESEEEDKCKPMSYEEKRQLSLDINKLPGEKLG 632
Query: 217 RRVACKQNPRRVKKPRLKS 235
R V Q+ ++P LK+
Sbjct: 633 RVVHIIQS----REPSLKN 647
>ATB2_RAT (P11506) Plasma membrane calcium-transporting ATPase 2 (EC
3.6.3.8) (PMCA2) (Plasma membrane calcium pump isoform
2) (Plasma membrane calcium ATPase isoform 2)
Length = 1243
Score = 44.3 bits (103), Expect = 6e-04
Identities = 47/183 (25%), Positives = 82/183 (44%), Gaps = 16/183 (8%)
Query: 7 LPADGLDDVHQNGVHDEPSNSGE-DAVSNDLDPHVTVNTETFVPDGNSENINQLESTATG 65
LPADGL + DE S +GE D V +D + + T V +G+ + TA G
Sbjct: 222 LPADGLFIQGNDLKIDESSLTGESDQVRKSVDKDPMLLSGTHVMEGSGRMV----VTAVG 277
Query: 66 NSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASL 125
++ I + G +E+E K K ++ + P + A ++ NASL
Sbjct: 278 VNSQTGIIFTLLGAGG------EEEEKKDKKGVKKGDGLQLPAADGAAPANAAGSANASL 331
Query: 126 VKNSKI----GKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPL 181
V N K+ Q+ +G +A++ +P + + ++D++ KK KS+ +G L
Sbjct: 332 V-NGKMQDGSADSSQSKAKQQDGAAAMEMQPLKSAEGGDADDKKKANMHKKEKSVLQGKL 390
Query: 182 DKV 184
K+
Sbjct: 391 TKL 393
>ATB2_HUMAN (Q01814) Plasma membrane calcium-transporting ATPase 2
(EC 3.6.3.8) (PMCA2) (Plasma membrane calcium pump
isoform 2) (Plasma membrane calcium ATPase isoform 2)
Length = 1243
Score = 44.3 bits (103), Expect = 6e-04
Identities = 47/182 (25%), Positives = 82/182 (44%), Gaps = 14/182 (7%)
Query: 7 LPADGLDDVHQNGVHDEPSNSGE-DAVSNDLDPHVTVNTETFVPDGNSENINQLESTATG 65
LPADGL + DE S +GE D V +D + + T V +G+ + TA G
Sbjct: 222 LPADGLFIQGNDLKIDESSLTGESDQVRKSVDKDPMLLSGTHVMEGSGRML----VTAVG 277
Query: 66 NSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASL 125
++ I + G +E+E K K ++ + P + A ++ NASL
Sbjct: 278 VNSQTGIIFTLLGAGG------EEEEKKDKKGVKKGDGLQLPAADGAAASNAADSANASL 331
Query: 126 VKNSKIGKDKQASPAVS---NGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLD 182
V + AS + + +G +A++ +P + + ++DR+ KK KS+ +G L
Sbjct: 332 VNGKMQDGNVDASQSKAKQQDGAAAMEMQPLKSAEGGDADDRKKASMHKKEKSVLQGKLT 391
Query: 183 KV 184
K+
Sbjct: 392 KL 393
>GRPB_RAT (P08462) Submandibular gland secretory Glx-rich protein CB
precursor (GRP-CB) (Contiguous repeat polypeptide) (CRP)
Length = 247
Score = 43.9 bits (102), Expect = 8e-04
Identities = 33/167 (19%), Positives = 65/167 (38%), Gaps = 9/167 (5%)
Query: 9 ADGLDDVHQNG-VHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNS 67
A G D+ N D P++S + V + DP D ++EN+ + ES N
Sbjct: 16 AQGTDEEVNNAETSDVPADSEQQPVDSGSDPPSA--------DADAENVQEGESAPPANE 67
Query: 68 AMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVK 127
GS + T ++ +E +E+ + Q+ P + +N + ++SG +
Sbjct: 68 EPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQQEPTQAENQEPPATSGSEEEQQQ 127
Query: 128 NSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPK 174
+ Q PA S + +N+ +D + + +P+
Sbjct: 128 QQPTQAENQEPPATSGSEEEQQQQESTQAENQEPSDSAGEGQETQPE 174
>PPIG_HUMAN (Q13427) Peptidyl-prolyl cis-trans isomerase G (EC
5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G)
(Rotamase G) (Cyclophilin G) (Clk associating
RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SRcyp)
(CASP10)
Length = 754
Score = 43.5 bits (101), Expect = 0.001
Identities = 38/170 (22%), Positives = 73/170 (42%), Gaps = 22/170 (12%)
Query: 64 TGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNA 123
+G ++EIE N D ++ ++ +++ + K VK + K SS ++
Sbjct: 146 SGQEVVREIE--NQKTDAASKPFAE-----VRILSCGELIPKSKVKKEEKKRHKSSSSSS 198
Query: 124 SLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKK----- 178
S +S D Q+S S+ SA + + ++ K N R+ + KKK K KK
Sbjct: 199 SSSSDSDSSSDSQSSSDSSDSESATEEKSKKRKKKHRKNSRKHKKEKKKRKKSKKSASSE 258
Query: 179 GPLDKVQGEGESSLYP----------FSLRKGESSSLRLRKRFRQRRRRR 218
+ ++ + +S++ P F +RK + ++ R+R R R
Sbjct: 259 SEAENLEAQPQSTVRPEEIPPIPENRFLMRKSPPKADEKERKNRERERER 308
Score = 31.6 bits (70), Expect = 4.0
Identities = 45/212 (21%), Positives = 81/212 (37%), Gaps = 22/212 (10%)
Query: 35 DLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKI 94
+L P V E ++ + S+++ + + + + S+D+ D + T K K+ K
Sbjct: 176 ELIPKSKVKKE---EKKRHKSSSSSSSSSSDSDSSSDSQSSSDSSDSESATEEKSKKRKK 232
Query: 95 KVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV--KNSKIGKDKQASPAVSNGTSALDSRP 152
K + +K K K +K +SS A + + + ++ P N S P
Sbjct: 233 KHRKNSRKHKKEKKKRKKSKKSASSESEAENLEAQPQSTVRPEEIPPIPENRFLMRKSPP 292
Query: 153 RQPIKNRSSNDRQSQLSKKKPKS---------LKKGPLDKVQGEGESSLYPFSLRKGESS 203
+ K R + +R+ + P S L K++G G R S
Sbjct: 293 KADEKERKNRERERERECNPPNSQPASYQRRLLVTRSGRKIKGRGPR-------RYRTPS 345
Query: 204 SLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
R R RFR R +Q +R ++ R+ S
Sbjct: 346 RSRSRDRFR-RSETPPHWRQEMQRAQRMRVSS 376
Score = 30.8 bits (68), Expect = 6.8
Identities = 32/178 (17%), Positives = 70/178 (38%), Gaps = 9/178 (5%)
Query: 53 SENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKN 112
+++ N ES N K+++ N ++ + EK+ K K ++ K K+K
Sbjct: 407 TDHRNVSESPNRKNEKEKKVKDHKSNSKERDIRRNSEKDDKYKNKVKKRAKSKSRSKSKE 466
Query: 113 AKVGSSSGVNASLVKNSKIGK-DKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKK 171
+ S ++SK + +++ + S G + + ++ + D++ SK+
Sbjct: 467 K--------SKSKERDSKHNRNEEKRMRSRSKGRDHENVKEKEKQSDSKGKDQERSRSKE 518
Query: 172 KPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVK 229
K K L+ + + + R E + + + R R R + RRV+
Sbjct: 519 KSKQLESKSNEHDHSKSKEKDRRAQSRSRECDITKGKHSYNSRTRERSRSRDRSRRVR 576
>MLH_TETTH (P40631) Micronuclear linker histone polyprotein (MIC LH)
[Contains: Micronuclear linker histone-alpha;
Micronuclear linker histone-beta; Micronuclear linker
histone-delta; Micronuclear linker histone-gamma]
Length = 633
Score = 41.6 bits (96), Expect = 0.004
Identities = 41/148 (27%), Positives = 64/148 (42%), Gaps = 8/148 (5%)
Query: 73 EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNK--NAKVGSSSGVNASLVKNSK 130
+G + + + S K+ K S+ + + KN N + SSS +S KN K
Sbjct: 216 QGRSQSSSSNRRKASSSKDQKGTRSSSRKASNSKGRKNSTSNKRNSSSSSKRSSSSKNKK 275
Query: 131 IGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGES 190
K + S G + SR R K SS +R+S SK + S KG K +S
Sbjct: 276 SSSSKNKKSSSSKGRKSSSSRGR---KASSSKNRKSSKSKDRKSSSSKG--RKSSSSSKS 330
Query: 191 SLYPFSLRKG-ESSSLRLRKRFRQRRRR 217
+ S +G +SSS + RK + + R+
Sbjct: 331 NKRKASSSRGRKSSSSKGRKSSKSQERK 358
Score = 38.9 bits (89), Expect = 0.025
Identities = 43/161 (26%), Positives = 67/161 (40%), Gaps = 18/161 (11%)
Query: 87 SKEKEVKIKVSTEQSR---------AQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQA 137
+++K+ K K ST +SR KG K+ ++K + S + ++S + K +
Sbjct: 171 AQKKQTKRKNSTSKSRRSSSKGKSSVSKGRTKSTSSKRRADSSASQGRSQSSSSNRRKAS 230
Query: 138 SPAVSNGT--SALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPF 195
S GT S+ + + KN +SN R S S K+ S K + + SS
Sbjct: 231 SSKDQKGTRSSSRKASNSKGRKNSTSNKRNSSSSSKRSSSSKNKKSSSSKNKKSSS---- 286
Query: 196 SLRKG-ESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKS 235
KG +SSS R RK + R+ K K R S
Sbjct: 287 --SKGRKSSSSRGRKASSSKNRKSSKSKDRKSSSSKGRKSS 325
Score = 32.3 bits (72), Expect = 2.3
Identities = 32/123 (26%), Positives = 51/123 (41%), Gaps = 14/123 (11%)
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPV--KNKNAKVGSS 118
S + ++MKE N N + SK + K + + KG K KN+K S+
Sbjct: 388 SKKSRRNSMKEARTKKAN----NKSASKASKSGSKSKGKSASKSKGKSSSKGKNSKSRSA 443
Query: 119 SGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPR------QPIKNRSSNDRQSQLSKKK 172
S ++ +NS Q + + N +S +R R + + N SN + S KK
Sbjct: 444 SKPKSNAAQNS--NNTHQTADSSENASSTTQTRTRGRQREQKDMVNEKSNSKSSSKGKKN 501
Query: 173 PKS 175
KS
Sbjct: 502 SKS 504
Score = 32.0 bits (71), Expect = 3.0
Identities = 39/181 (21%), Positives = 71/181 (38%), Gaps = 7/181 (3%)
Query: 53 SENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKN 112
S + +++ S E EG S+ + + + S +++R +K NK+
Sbjct: 351 SSKSQERKNSHADTSKQMEDEGQKRRQSSSSAKRDESSKKSRRNSMKEARTKKA--NNKS 408
Query: 113 AKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKK 172
A S SG + S K++ K K +S N S S+P+ S+N Q+ S +
Sbjct: 409 ASKASKSG-SKSKGKSASKSKGKSSSKG-KNSKSRSASKPKSNAAQNSNNTHQTADSSEN 466
Query: 173 PKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRK---RFRQRRRRRVACKQNPRRVK 229
S + Q E + + S K S + K R + + + ++N + K
Sbjct: 467 ASSTTQTRTRGRQREQKDMVNEKSNSKSSSKGKKNSKSNTRSKSKSKSASKSRKNASKSK 526
Query: 230 K 230
K
Sbjct: 527 K 527
Score = 31.6 bits (70), Expect = 4.0
Identities = 38/214 (17%), Positives = 78/214 (35%), Gaps = 19/214 (8%)
Query: 17 QNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNSAMKEIEGSN 76
+N S +A N + H T ++ SEN + T T ++ + N
Sbjct: 436 KNSKSRSASKPKSNAAQNSNNTHQTADS--------SENASSTTQTRTRGRQREQKDMVN 487
Query: 77 DNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKI---GK 133
+ ++ + SK K+ + +S+++ KNA N SK K
Sbjct: 488 EK--SNSKSSSKGKKNSKSNTRSKSKSKSASKSRKNASKSKKDTTNHGRQTRSKSRSESK 545
Query: 134 DKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLY 193
K +P + + +P++ +R + +SQ +KK K D + E
Sbjct: 546 SKSEAPNKPSNKMEVIEQPKEESSDRKRRESRSQSAKKTSDKKSKNRSDSKKMTAEDP-- 603
Query: 194 PFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRR 227
+K + + +K+ ++ + K N ++
Sbjct: 604 ----KKNNAEDSKGKKKSKEGKTGAYGKKANKKQ 633
>NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130
kDa protein) (140 kDa nucleolar phosphoprotein)
(Nopp140) (Nucleolar and coiled-body phosphoprotein 1)
Length = 699
Score = 41.2 bits (95), Expect = 0.005
Identities = 42/178 (23%), Positives = 79/178 (43%), Gaps = 23/178 (12%)
Query: 73 EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQK-----GPVKNKNAKVGSSSGVNASLVK 127
+GS D+ S+ + S+E+E K S + + QK P K +AK G + N+S
Sbjct: 464 QGSRDSSSDSDSSSSEEEEEKTSKSAVKKKPQKVAGGAAPSKPASAKKGKAESSNSSSSD 523
Query: 128 NSKIGKDKQ----ASPAV----SNGTSALDSRPRQPIKNRSSNDRQSQLSK---KKPKSL 176
+S ++++ SP +NGTSAL ++ + KN + + + + K SL
Sbjct: 524 DSSEEEEEKLKGKGSPRPQAPKANGTSALTAQNGKAAKNSEEEEEEKKKAAVVVSKSGSL 583
Query: 177 KKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLK 234
KK ++ E E+ + + L+ F +R++ RRV++ ++
Sbjct: 584 KKRKQNEAAKEAETP-------QAKKIKLQTPNTFPKRKKGEKRASSPFRRVREEEIE 634
Score = 36.2 bits (82), Expect = 0.16
Identities = 43/196 (21%), Positives = 81/196 (40%), Gaps = 28/196 (14%)
Query: 72 IEGSNDNVDGSNLTVSKEKE-----VKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
+E S D+ D S+ + +EK+ V K +T+ A+K + ++ SS + +
Sbjct: 319 VESSEDSSDESDSSSEEEKKPPTKAVVSKATTKPPPAKKAAESSSDSSDSDSSEDDEAPS 378
Query: 127 KNSKIGKDKQASPAVSNGTSALD--SRPRQPI---------KNRSSNDRQSQLSKKKPKS 175
K + K+ PAV+ + A+ + P+QP+ K SS+ + S ++ K+
Sbjct: 379 KPAGTTKNSSNKPAVTTKSPAVKPAAAPKQPVGGGQKLLTRKADSSSSEEESSSSEEEKT 438
Query: 176 LKKGPLDKVQGEGESSL------YPFSLRKGESSSLRLRKRFRQRRRRRVACKQNPRRV- 228
K K + +++L P R S S + + + A K+ P++V
Sbjct: 439 KKMVATTKPKATAKAALSLPAKQAPQGSRDSSSDSDSSSSEEEEEKTSKSAVKKKPQKVA 498
Query: 229 -----KKPRLKSLGRA 239
KP G+A
Sbjct: 499 GGAAPSKPASAKKGKA 514
>PI4K_DICDI (P54677) Phosphatidylinositol 4-kinase (EC 2.7.1.67)
(PI4-kinase) (PtdIns-4-kinase) (PI4K-alpha)
Length = 1093
Score = 40.8 bits (94), Expect = 0.007
Identities = 40/178 (22%), Positives = 77/178 (42%), Gaps = 24/178 (13%)
Query: 25 SNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLESTATGNS-AMKEIEGSND------ 77
+N+ + ++N+ + N +P+ NS+N E+ GNS I G N+
Sbjct: 186 NNNNNNNINNNNSNNDDNNNNEILPNENSDNSINDENNQYGNSNNNNNISGENNNIKIDI 245
Query: 78 --------NVDGSNLTVSKE-KEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKN 128
N++ N T+ +E K IK E + N N +++ +N + + N
Sbjct: 246 NSQNKSDSNIETLNSTLCEETKTSPIKDDMENNNNNNNNNNNNNNNNNNNNNINNNNINN 305
Query: 129 SKIGKDKQASPAVSNGT-SALD-------SRPRQPIKNRSSNDRQSQLSKKKPKSLKK 178
+ I + + NG+ S LD S+P PI+N + +++++ KK + K+
Sbjct: 306 NNINNNNNINYGHINGSLSTLDGIGQPYISQPNDPIENITQILKRNRIIYKKVEEKKE 363
>HDA1_STRPU (P56518) Histone deacetylase 1 (HD1)
Length = 576
Score = 40.4 bits (93), Expect = 0.009
Identities = 42/185 (22%), Positives = 61/185 (32%), Gaps = 19/185 (10%)
Query: 20 VHDEPSNSGEDAVSNDLD-------------PHVTVNTETFVPDGNSENINQLESTATGN 66
+H PSN LD PH +P+ + + E A
Sbjct: 342 LHISPSNMTNQNTGEYLDKIKTRLYENMRMIPHAPGVQMQPIPEDAIPDDSDAEDEAENP 401
Query: 67 SAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLV 126
I + + + E E + ++ E R K K +K+ S G A
Sbjct: 402 DKRISIMAQDKRIQRDDEFSDSEDEGETRLPGEGGRRDHRSHKAKRSKIDDSPGKEADKE 461
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQG 186
S K+A PA +D+ P P K S KP +K P +K G
Sbjct: 462 AKSS-DASKEAKPAAEPQAVPMDTTPAPPPKKSEDKPEAS-----KPTEVKAKPAEKEPG 515
Query: 187 EGESS 191
EGE+S
Sbjct: 516 EGEAS 520
>SFR4_HUMAN (Q08170) Splicing factor, arginine/serine-rich 4
(Pre-mRNA splicing factor SRP75) (SRP001LB)
Length = 494
Score = 39.7 bits (91), Expect = 0.015
Identities = 33/143 (23%), Positives = 65/143 (45%), Gaps = 6/143 (4%)
Query: 97 STEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPI 156
S QSR++ K+++ S + S K+ KD QA + N + + R P
Sbjct: 234 SRSQSRSRSKKEKSRSPSKEKSRSRSHSAGKSRSKSKD-QAEEKIQNNDNVGKPKSRSPS 292
Query: 157 KNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRK----GESSSLRLRKRFR 212
+++S + +S+ +++ + K+G + + + + E SL R G S R R + +
Sbjct: 293 RHKSKSKSRSRSQERRVEEEKRGSVSRGRSQ-EKSLRQSRSRSRSKGGSRSRSRSRSKSK 351
Query: 213 QRRRRRVACKQNPRRVKKPRLKS 235
+R+ R ++ R + R KS
Sbjct: 352 DKRKGRKRSREESRSRSRSRSKS 374
>CYP1_BRUMA (Q27450) Peptidylprolyl isomerase 1 (EC 5.2.1.8)
(Peptidylprolyl cis-trans isomerase 1) (PPIase 1)
(Cyclophilin) (BmCYP-1)
Length = 843
Score = 39.7 bits (91), Expect = 0.015
Identities = 63/247 (25%), Positives = 100/247 (39%), Gaps = 53/247 (21%)
Query: 2 DPVNSLPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLES 61
D S + LDD+ + + S + D TV +T V D NS+N S
Sbjct: 594 DGRRSTSREKLDDLKRKETSGQKSQA---------DSEQTVEAKTNVVDSNSDN-----S 639
Query: 62 TATGNSAMKEIEGSNDNVDGS-----NLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAKVG 116
+ N +KE+ +N + S +K +E+K +V+ E SR QKG K K K
Sbjct: 640 KMSVNGKLKEVSSTNKENEVSEQKDLKAESTKSEEIKQQVN-EVSRKQKGGEKPKEHK-- 696
Query: 117 SSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLSKKKPKSL 176
+N + ++ S SNG R R + S DR + K +S
Sbjct: 697 ----------RNERSRSRRRRSR--SNG-----RRRRSSSRRSRSRDR-----RHKSRSR 734
Query: 177 KKGPLDKVQGEGES------SLYPFSLRKGESSSLRLRKRF--RQRRRRRVACKQNPRRV 228
+G + + +G S LY +R+ E R R+ R+R R R A + + R
Sbjct: 735 SRGYVRRFEGWSRSRRPTRRELYDERMRR-ERERRRSFDRYSDRRRTRSRSARRDSDRHS 793
Query: 229 KKPRLKS 235
++ R +S
Sbjct: 794 RRSRKRS 800
>YKD1_CAEEL (Q03560) Hypothetical protein B0464.2 in chromosome III
Length = 1150
Score = 38.5 bits (88), Expect = 0.032
Identities = 39/172 (22%), Positives = 68/172 (38%), Gaps = 13/172 (7%)
Query: 28 GEDAVSNDLDPHVTVNTETFVPDGNS--ENINQLESTATGNSAMKEIEGSNDNVDGSNLT 85
G ++D D V +++ DG E+ + E +A K + D
Sbjct: 941 GRKRRNDDGDEFVNDSSDAGNYDGEEGGEDGERRERRKKDKAAKKASRKKRERRDSGGPD 1000
Query: 86 VSKEKEVKIKVSTEQSRAQKGPVKNK-NAKVGSSSGVNASLVKNSKIGKDKQASPAVSNG 144
++ E K K E+ R + + K +AK+ S + + +S+ D + PA +
Sbjct: 1001 SNRRDEKKRKRKEERDRKLQEKLSAKQSAKIKSRA-----FLSSSESSDDDKPKPAADSS 1055
Query: 145 TSALDSRPR-----QPIKNRSSNDRQSQLSKKKPKSLKKGPLDKVQGEGESS 191
+D RP P + S +DR++ KKK K++ V G G S
Sbjct: 1056 DDEVDPRPPVDEFDSPTRTDSDSDRETTTKKKKKKAVVDSDEGSVSGSGSGS 1107
>KIJ5_YEAST (P40494) Probable serine/threonine-protein kinase
YIL095W (EC 2.7.1.37)
Length = 810
Score = 38.5 bits (88), Expect = 0.032
Identities = 37/172 (21%), Positives = 69/172 (39%), Gaps = 19/172 (11%)
Query: 41 TVNTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSNLT-VSKEKEVKIKVSTE 99
T + T +PD + NINQ + + ++ S+D V + + K ++ + S
Sbjct: 653 TTSKPTLIPDNGNVNINQEKKESIQRRVHNLLKSSDDPVTYKSASGYGKYTDIGTETSNR 712
Query: 100 QSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNR 159
S + P+ + K GV +K GKDK P P +P+ R
Sbjct: 713 HSSVRITPITEEKFKKTLKDGVLD--IKTKSNGKDKSRPP----------RPPPKPLHLR 760
Query: 160 SSNDRQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRF 211
+ + S+ + K L P++++ E ++ ++ E+ RKRF
Sbjct: 761 TEIQKIRNFSRLQSKKL---PIERISSEATETIVDVNVDDLEAD---FRKRF 806
>THO2_HUMAN (Q8NI27) THO complex subunit 2 (Tho2)
Length = 1478
Score = 38.1 bits (87), Expect = 0.042
Identities = 40/168 (23%), Positives = 76/168 (44%), Gaps = 9/168 (5%)
Query: 61 STATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIK-VSTEQSRAQKGPVKNKNAKVGSSS 119
S+++ SA K E S + D S + + +K V+ S KG N N+ S+
Sbjct: 1094 SSSSIGSASKSDESSTEETDKSR----ERSQCGVKAVNKASSTTPKGNSSNGNSGSNSNK 1149
Query: 120 GV--NASLVKNSKIGKDKQASPAVSNGTSALDSRPRQ-PIKNRSSNDRQSQLSKKK-PKS 175
V N K + K+ +PA + L ++ P + R + D +++ +K++ PKS
Sbjct: 1150 AVKENDKEKGKEKEKEKKEKTPATTPEARVLGKDGKEKPKEERPNKDEKARETKERTPKS 1209
Query: 176 LKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQ 223
K+ K + + + + ++ ES S + R+R ++ R R K+
Sbjct: 1210 DKEKEKFKKEEKAKDEKFKTTVPNAESKSTQEREREKEPSRERDIAKE 1257
>YBM5_SCHPO (Q10337) Hypothetical protein C582.05c in chromosome II
Length = 878
Score = 37.4 bits (85), Expect = 0.072
Identities = 38/178 (21%), Positives = 77/178 (42%), Gaps = 17/178 (9%)
Query: 12 LDDVHQNGVHDEPSNSGEDAVSNDLDPHVTV-NTETFVPDGNSENINQLESTATGNSAMK 70
LDD + + + P N + + + L +T N+E + D + +++ ++ T K
Sbjct: 401 LDDSYMDFIFPCPLNVEKGSFEDTLKSSLTKGNSEVLLDDLSDPSVSSIKGNKTNEELEK 460
Query: 71 EIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKG----PVKNKNAKVGSSSGVNASLV 126
E + ++DN K + + S A KG ++N + + +GVN L
Sbjct: 461 EFKSTSDNFG---------KHIILTSSFSNQSADKGSSLAAEDDRNDEGSTITGVNRELQ 511
Query: 127 KNSKIGKDKQASPAVSNGTSALDSRP-RQPIKNRSSNDRQSQLSKKKPKSL--KKGPL 181
++ D ++S + + L P ++ +K SS+D L+ K ++ +K PL
Sbjct: 512 DEGRLEIDAKSSKTNTPPSPLLVGTPSKESLKEASSDDELPVLATKLVDNVIKEKSPL 569
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,086,593
Number of Sequences: 164201
Number of extensions: 1752499
Number of successful extensions: 8220
Number of sequences better than 10.0: 151
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 130
Number of HSP's that attempted gapping in prelim test: 7886
Number of HSP's gapped (non-prelim): 308
length of query: 419
length of database: 59,974,054
effective HSP length: 113
effective length of query: 306
effective length of database: 41,419,341
effective search space: 12674318346
effective search space used: 12674318346
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC145331.12


