

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.1 - phase: 0
(520 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
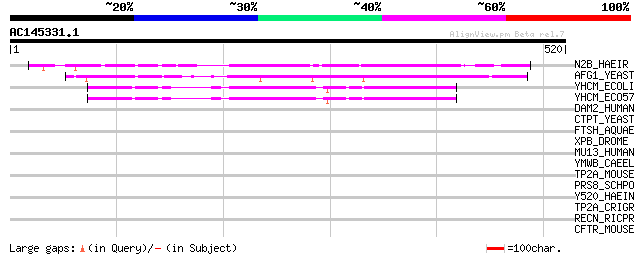
Score E
Sequences producing significant alignments: (bits) Value
N2B_HAEIR (P46441) Putative ATPase N2B (HFN2B) 191 5e-48
AFG1_YEAST (P32317) AFG1 protein 149 1e-35
YHCM_ECOLI (P64612) Hypothetical protein yhcM 133 1e-30
YHCM_ECO57 (P64613) Hypothetical protein yhcM 133 1e-30
DAM2_HUMAN (Q86T65) Disheveled associated activator of morphogen... 34 1.1
CTPT_YEAST (P13259) Choline-phosphate cytidylyltransferase (EC 2... 34 1.1
FTSH_AQUAE (O67077) Cell division protein ftsH homolog (EC 3.4.2... 33 1.4
XPB_DROME (Q02870) DNA excision repair protein haywire (ERCC-3 h... 33 2.3
MU13_HUMAN (Q9H3R2) Mucin 13 precursor (Down-regulated in colon ... 32 3.1
YMWB_CAEEL (Q09385) Hypothetical protein K04G7.11 in chromosome III 32 4.0
TP2A_MOUSE (Q01320) DNA topoisomerase II, alpha isozyme (EC 5.99... 31 6.8
PRS8_SCHPO (P41836) 26S protease regulatory subunit 8 homolog (P... 31 6.8
Y520_HAEIN (P44743) Hypothetical protein HI0520 31 8.9
TP2A_CRIGR (P41515) DNA topoisomerase II, alpha isozyme (EC 5.99... 31 8.9
RECN_RICPR (Q9ZDY2) DNA repair protein recN (Recombination prote... 31 8.9
CFTR_MOUSE (P26361) Cystic fibrosis transmembrane conductance re... 31 8.9
>N2B_HAEIR (P46441) Putative ATPase N2B (HFN2B)
Length = 464
Score = 191 bits (484), Expect = 5e-48
Identities = 144/476 (30%), Positives = 230/476 (48%), Gaps = 70/476 (14%)
Query: 18 ERRRILMDEVEKQ--QNDKDWWKRLNNKITERWTNSRKRPENVDP--GVGKWVSYLKREK 73
E + +L D+V+K+ Q +D + L N + +P V+ G G + ++K+E+
Sbjct: 46 ESKELLPDKVQKKTTQELEDLYNTLKNY--------QPKPVRVETSSGGGFFGRFMKKEQ 97
Query: 74 KLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRIN 133
+ TAP KG+YIYG+VG GKT LMDMFYS + I K ++R HF+ M +++
Sbjct: 98 SAPKIELLNTTAP---KGMYIYGSVGGGKTTLMDMFYSCCDDIPK-KQRVHFNSFMSKVH 153
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFD-TKAKEWLAAEERYKKEVQMKHILPDVADK 192
+HK + E P +S LPFD T + A E +
Sbjct: 154 GLIHKV---KQERGPQDRAFNSE-KPLPFDPTLPVAEMIANESW---------------- 193
Query: 193 FFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEAN 252
++CFDE Q D+ + L + + L + G + +ATSNR P DL +
Sbjct: 194 -------------LICFDEFQVTDIADAMILKSLFTHLFNEGIVCIATSNRHPNDLYKNG 240
Query: 253 MVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRVNYLWPIERETINKFEKKWQD 312
+ F + L C + S +DYRR IAQ + NY + + ++ E+ ++
Sbjct: 241 LQRSNFIPFIGVLLNRCNVAAMDSGVDYRR-IAQSGDT--NYFVTTQTDAKSQMERMFKI 297
Query: 313 ATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYHTVF 372
+ + TI+ FGR L +C V F+ LC RPLG +DYI + + +HTV
Sbjct: 298 LCSQENDIIRPRTIT-HFGRDLTFQRTCGQVLDSNFEELCNRPLGGSDYIQIGQFFHTVL 356
Query: 373 ISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGTEEGTLFDLESFQF 432
I D+P +++ ++ + RRFITLID LY++ + A +D+LF T++ DL
Sbjct: 357 IHDVPQLTLLLKSQMRRFITLIDTLYDNRVRVVISAEVPLDQLFSFTDKPK--DL----- 409
Query: 433 ETEAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQTQLY 488
A+ ++ D L G+ + S+ +G+EE+F F R +SRL EMQ + Y
Sbjct: 410 ---ADEQRMLMDDLKLGDTDTS------ASVFTGEEEMFAFDRTISRLYEMQKKEY 456
>AFG1_YEAST (P32317) AFG1 protein
Length = 509
Score = 149 bits (377), Expect = 1e-35
Identities = 128/461 (27%), Positives = 210/461 (44%), Gaps = 66/461 (14%)
Query: 53 KRPENVDPGVGKWVSYLK------REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLM 106
K P VD VG W++ LK + K + + V P+G+Y+YG+VG GKTMLM
Sbjct: 78 KTPNAVDQ-VGGWLNGLKSVFSRGKPKNIGAYVDVSKIGNSIPRGVYLYGDVGCGKTMLM 136
Query: 107 DMFYSATEGIVKHRRRYHFHEAMLRINEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKA 166
D+FY+ + ++R HFH+ M +++ H+ ++Q K L I +PF
Sbjct: 137 DLFYTTIPNHLT-KKRIHFHQFMQYVHKRSHEIVREQ-NLKELGDAKGKEIDTVPF---- 190
Query: 167 KEWLAAEERYKKEVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGI 226
LAAE +A+ +++LCFDE Q DV + L +
Sbjct: 191 ---LAAE---------------IANN-----------SHVLCFDEFQVTDVADAMILRRL 221
Query: 227 LSRLLSS--GTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRR-- 282
++ LLS G ++ ATSNR P +L + + F + ++ + + + S DYR+
Sbjct: 222 MTALLSDDYGVVLFATSNRHPDELYINGVQRQSFIPCIELIKHRTKVIFLNSPTDYRKIP 281
Query: 283 --------FIAQRSENRVNYLWPIERETINKFEKKWQDATGRFGGKVISNTISVMF---- 330
F + S + RET K + S+T+ F
Sbjct: 282 RPVSSVYYFPSDTSIKYASKECKTRRETHIKEWYNYFAQASHTDDSTDSHTVHKTFYDYP 341
Query: 331 ----GRTLEVPESCEG-VARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSMRIRD 385
GR +VP+ VA+FTF LCG PL A DY+ +A+N+ ++DIP +S+ +RD
Sbjct: 342 LTIWGREFKVPKCTPPRVAQFTFKQLCGEPLAAGDYLTLAKNFEAFIVTDIPYLSIYVRD 401
Query: 386 KARRFITLIDELYNHHSCLCCLASSSIDELFQGTEE-GTLFDLESFQFETEAEGSKLRRD 444
+ RRFIT +D +Y+ L ++ LF E+ F+L E ++ + + +
Sbjct: 402 EVRRFITFLDAVYDSGGKLATTGAADFSSLFVEPEQILNDFELRPTTKEPDSVDTGMVDE 461
Query: 445 VLAEGNVGSGGTPVGITSILSGQEELFTFQRAVSRLIEMQT 485
++ + G + + + EE F F RA+SRL +M +
Sbjct: 462 MVEKH--GFSKEIAKKSQMFALDEERFAFARALSRLSQMSS 500
>YHCM_ECOLI (P64612) Hypothetical protein yhcM
Length = 375
Score = 133 bits (334), Expect = 1e-30
Identities = 100/349 (28%), Positives = 159/349 (44%), Gaps = 62/349 (17%)
Query: 74 KLDSLVGRRPTAPPAP-KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRI 132
++ L G+R P +GLY++G VG GKT LMD+FY + G + ++R HFH MLR+
Sbjct: 55 RVGKLWGKREDTKHTPVRGLYMWGGVGRGKTWLMDLFYQSLPG--ERKQRLHFHRFMLRV 112
Query: 133 NEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADK 192
+E + Q + PL+ +AD+
Sbjct: 113 HEELTAL---QGQTDPLEI-------------------------------------IADR 132
Query: 193 FFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEAN 252
F + + +LCFDE D+ + L G++ L + G +VATSN P +L
Sbjct: 133 FKAETD-------VLCFDEFFVSDITDAMLLGGLMKALFARGITLVATSNIPPDELYRNG 185
Query: 253 MVPEFFQNLLSNLEEHCEKVLVGSEIDYR-RFIAQRSENRVNYLW--PIERETINKFEKK 309
+ F + +++HC+ + V + +DYR R + Q +LW P+ ET + +K
Sbjct: 186 LQRARFLPAIDAIKQHCDVMNVDAGVDYRLRTLTQA------HLWLSPLHDETRAQMDKL 239
Query: 310 WQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYH 369
W G G + S T+ + R L +F LC DYIA++ +H
Sbjct: 240 WLALAG--GKRENSPTLEINH-RPLATMGVENQTLAVSFTTLCVDARSQHDYIALSRLFH 296
Query: 370 TVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQG 418
TV + D+P+M+ + +ARRFI L+DE Y H L A + E++QG
Sbjct: 297 TVMLFDVPVMTRLMESEARRFIALVDEFYERHVKLVVSAEVPLYEIYQG 345
>YHCM_ECO57 (P64613) Hypothetical protein yhcM
Length = 375
Score = 133 bits (334), Expect = 1e-30
Identities = 100/349 (28%), Positives = 159/349 (44%), Gaps = 62/349 (17%)
Query: 74 KLDSLVGRRPTAPPAP-KGLYIYGNVGSGKTMLMDMFYSATEGIVKHRRRYHFHEAMLRI 132
++ L G+R P +GLY++G VG GKT LMD+FY + G + ++R HFH MLR+
Sbjct: 55 RVGKLWGKREDTKHTPVRGLYMWGGVGRGKTWLMDLFYQSLPG--ERKQRLHFHRFMLRV 112
Query: 133 NEHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHILPDVADK 192
+E + Q + PL+ +AD+
Sbjct: 113 HEELTAL---QGQTDPLEI-------------------------------------IADR 132
Query: 193 FFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEAN 252
F + + +LCFDE D+ + L G++ L + G +VATSN P +L
Sbjct: 133 FKAETD-------VLCFDEFFVSDITDAMLLGGLMKALFARGITLVATSNIPPDELYRNG 185
Query: 253 MVPEFFQNLLSNLEEHCEKVLVGSEIDYR-RFIAQRSENRVNYLW--PIERETINKFEKK 309
+ F + +++HC+ + V + +DYR R + Q +LW P+ ET + +K
Sbjct: 186 LQRARFLPAIDAIKQHCDVMNVDAGVDYRLRTLTQA------HLWLSPLHDETRAQMDKL 239
Query: 310 WQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYLCGRPLGAADYIAVAENYH 369
W G G + S T+ + R L +F LC DYIA++ +H
Sbjct: 240 WLALAG--GKRENSPTLEINH-RPLATMGVENQTLAVSFTTLCVDARSQHDYIALSRLFH 296
Query: 370 TVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQG 418
TV + D+P+M+ + +ARRFI L+DE Y H L A + E++QG
Sbjct: 297 TVMLFDVPVMTRLMESEARRFIALVDEFYERHVKLVVSAEVPLYEIYQG 345
>DAM2_HUMAN (Q86T65) Disheveled associated activator of morphogenesis
2
Length = 1068
Score = 33.9 bits (76), Expect = 1.1
Identities = 12/37 (32%), Positives = 23/37 (61%)
Query: 3 EYHVNLSNWEKKRENERRRILMDEVEKQQNDKDWWKR 39
E +L +++E E RR M+ + K+Q +++WW+R
Sbjct: 970 EARQDLEAMRRRKEEEERRARMEAMLKEQREREWWQR 1006
>CTPT_YEAST (P13259) Choline-phosphate cytidylyltransferase (EC
2.7.7.15) (Phosphorylcholine transferase)
(CTP:phosphocholine cytidylyltransferase) (CT) (CCT)
Length = 424
Score = 33.9 bits (76), Expect = 1.1
Identities = 29/94 (30%), Positives = 48/94 (50%), Gaps = 6/94 (6%)
Query: 7 NLSNWEKKRENERRRILMDEVEKQQN-DKDWWKRLN-NKITERWTNSRKR-PENVDPGVG 63
+LSN KK +N+R+R +E E+Q N DKD K + NK T+ R+R + +
Sbjct: 19 SLSNLFKKNKNKRQR---EETEEQDNEDKDESKNQDENKDTQLTPRKRRRLTKEFEEKEA 75
Query: 64 KWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGN 97
++ + L +E + G R PP + + IY +
Sbjct: 76 RYTNELPKELRKYRPKGFRFNLPPTDRPIRIYAD 109
>FTSH_AQUAE (O67077) Cell division protein ftsH homolog (EC
3.4.24.-)
Length = 634
Score = 33.5 bits (75), Expect = 1.4
Identities = 20/50 (40%), Positives = 25/50 (50%), Gaps = 6/50 (12%)
Query: 56 ENVDPGVGKWVSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
E V V + + YLK K L GR PKG+ +YG G GKT+L
Sbjct: 161 EEVKEEVKEIIEYLKDPVKFQKLGGR------PPKGVLLYGEPGVGKTLL 204
>XPB_DROME (Q02870) DNA excision repair protein haywire (ERCC-3
homolog protein)
Length = 802
Score = 32.7 bits (73), Expect = 2.3
Identities = 48/203 (23%), Positives = 83/203 (40%), Gaps = 29/203 (14%)
Query: 316 RFGGKVISNTISVMFGRTLEVPE-------SCEGVARFTFDYLCGRPLGAADYIAVAENY 368
+F KV + +S + + ++PE S G +R GR L A A+AE Y
Sbjct: 617 KFNSKVNTIFVSKVADTSFDLPEANVLIQISSHGGSRRQEAQRLGRILRAKKG-AIAEEY 675
Query: 369 HTVFISDIPMMSMRIRDKARRFITLIDELYNHHSCLCCLASSSIDELFQGT--EEGTLFD 426
+ F + + +M + +R L+++ Y++ + +L GT E+G L
Sbjct: 676 NAFFYTLVSQDTMEMSYSRKRQRFLVNQGYSYKVITHLKGMDTDSDLMYGTQEEQGQLLQ 735
Query: 427 LESFQFETEAEGSKLRRDVLAEGNVGSGG--TPVGITSILSGQEELFTFQRAVSRLIEMQ 484
L + + E KL + + GSGG VG S +SG ++ ++
Sbjct: 736 LVLSASDLDCEDEKLPGEPGYRPS-GSGGIVRRVGGLSSMSGGDDAIYYEHRKK------ 788
Query: 485 TQLYLDGVSNVHPYFQTQHKKFQ 507
+ +VHP F KKF+
Sbjct: 789 ------NIGSVHPLF----KKFR 801
>MU13_HUMAN (Q9H3R2) Mucin 13 precursor (Down-regulated in colon
cancer 1) (UNQ6194/PRO20221)
Length = 512
Score = 32.3 bits (72), Expect = 3.1
Identities = 17/54 (31%), Positives = 29/54 (53%), Gaps = 1/54 (1%)
Query: 208 CFDEIQTVDVFAIVALSGILSRLLSSGTIIVATSNRAPKDLNEANMVPEFFQNL 261
C D+ Q + + + ++GI+ + I+ A SN K + E N++ E FQNL
Sbjct: 415 CKDKFQLI-LTIVGTIAGIVILSMIIALIVTARSNNKTKHIEEENLIDEDFQNL 467
>YMWB_CAEEL (Q09385) Hypothetical protein K04G7.11 in chromosome III
Length = 216
Score = 32.0 bits (71), Expect = 4.0
Identities = 35/136 (25%), Positives = 53/136 (38%), Gaps = 24/136 (17%)
Query: 10 NWEKKRENERRRILMDEVEKQQNDK--DWWK----RLNNKITERWTNSRKRPENVDPGVG 63
N E K+E ++ ++ + K DK D+ + ++ +TE+ RKR +N D G
Sbjct: 38 NHEAKKERDQWQVKELQDRKAAEDKGLDYERVRSLEMSADVTEKLEQKRKRKKNPDQGFT 97
Query: 64 KWVSYLKRE---------------KKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTMLMDM 108
+ R+ KK+ VG P A I+GN T MD
Sbjct: 98 SYEDMTLRQHTRLTAALDPDLDSYKKMRECVGGEQFYPTA--DTLIHGN-HYPTTAAMDK 154
Query: 109 FYSATEGIVKHRRRYH 124
G VK R +YH
Sbjct: 155 LTKDVHGQVKRREQYH 170
>TP2A_MOUSE (Q01320) DNA topoisomerase II, alpha isozyme (EC
5.99.1.3)
Length = 1528
Score = 31.2 bits (69), Expect = 6.8
Identities = 16/43 (37%), Positives = 25/43 (57%)
Query: 144 MEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHIL 186
ME PLQ + +MN + K+ L+ E Y+K+ Q++HIL
Sbjct: 1 MELSPLQPVNENMLMNKKKNEDGKKRLSIERIYQKKTQLEHIL 43
>PRS8_SCHPO (P41836) 26S protease regulatory subunit 8 homolog
(Protein let1)
Length = 403
Score = 31.2 bits (69), Expect = 6.8
Identities = 19/58 (32%), Positives = 29/58 (49%), Gaps = 5/58 (8%)
Query: 53 KRPENVDPGVGKWVSYLKREKKLDSLVGRRPT-----APPAPKGLYIYGNVGSGKTML 105
K P++ VG +K K++ L + P P PKG+ +YG G+GKT+L
Sbjct: 138 KIPDSTYEMVGGLEKQIKEIKEVIELPVKHPELFESLGIPQPKGILLYGPPGTGKTLL 195
>Y520_HAEIN (P44743) Hypothetical protein HI0520
Length = 262
Score = 30.8 bits (68), Expect = 8.9
Identities = 14/25 (56%), Positives = 15/25 (60%)
Query: 186 LPDVADKFFLDREGEEKGANILCFD 210
L DV DKF D +GE G LCFD
Sbjct: 120 LIDVTDKFLFDLKGEGIGLQTLCFD 144
>TP2A_CRIGR (P41515) DNA topoisomerase II, alpha isozyme (EC
5.99.1.3)
Length = 1526
Score = 30.8 bits (68), Expect = 8.9
Identities = 17/43 (39%), Positives = 25/43 (57%)
Query: 144 MEEKPLQSGISSWIMNLPFDTKAKEWLAAEERYKKEVQMKHIL 186
ME PLQ + MN + AK+ L+ E Y+K+ Q++HIL
Sbjct: 1 MELSPLQPVNENMQMNKKKNEDAKKRLSIERIYQKKTQLEHIL 43
>RECN_RICPR (Q9ZDY2) DNA repair protein recN (Recombination protein
N)
Length = 554
Score = 30.8 bits (68), Expect = 8.9
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 6/55 (10%)
Query: 90 KGL-YIYGNVGSGKTMLMDMF-----YSATEGIVKHRRRYHFHEAMLRINEHMHK 138
KGL I G G+GK++L+D Y + I+KH + Y + +NE + K
Sbjct: 22 KGLCVITGETGAGKSILLDAILFCLGYKTSNNIIKHGKDYTVVNIIFSLNEKIKK 76
>CFTR_MOUSE (P26361) Cystic fibrosis transmembrane conductance
regulator (CFTR) (cAMP-dependent chloride channel)
Length = 1476
Score = 30.8 bits (68), Expect = 8.9
Identities = 17/37 (45%), Positives = 22/37 (58%), Gaps = 3/37 (8%)
Query: 92 LYIYGNVGSGKTMLMDMFYS---ATEGIVKHRRRYHF 125
L I G+ GSGKT L+ + A+EGI+KH R F
Sbjct: 454 LAITGSTGSGKTSLLMLILGELEASEGIIKHSGRVSF 490
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 62,277,584
Number of Sequences: 164201
Number of extensions: 2742509
Number of successful extensions: 8493
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 8467
Number of HSP's gapped (non-prelim): 23
length of query: 520
length of database: 59,974,054
effective HSP length: 115
effective length of query: 405
effective length of database: 41,090,939
effective search space: 16641830295
effective search space used: 16641830295
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145331.1


