

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145329.13 - phase: 2 /pseudo
(528 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
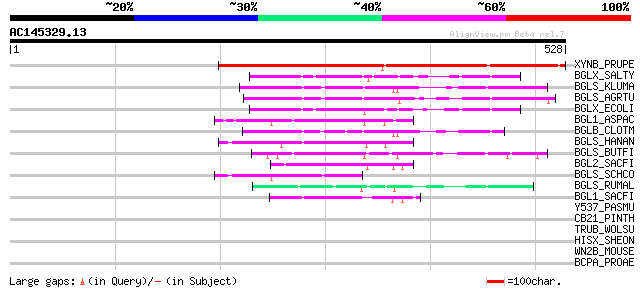
Score E
Sequences producing significant alignments: (bits) Value
XYNB_PRUPE (P83344) Putative beta-D-xylosidase (EC 3.2.1.-) (PpA... 430 e-120
BGLX_SALTY (Q56078) Periplasmic beta-glucosidase precursor (EC 3... 102 3e-21
BGLS_KLUMA (P07337) Beta-glucosidase precursor (EC 3.2.1.21) (Ge... 99 4e-20
BGLS_AGRTU (P27034) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)... 97 8e-20
BGLX_ECOLI (P33363) Periplasmic beta-glucosidase precursor (EC 3... 95 4e-19
BGL1_ASPAC (P48825) Beta-glucosidase 1 precursor (EC 3.2.1.21) (... 80 1e-14
BGLB_CLOTM (P14002) Thermostable beta-glucosidase B (EC 3.2.1.21... 70 1e-11
BGLS_HANAN (P06835) Beta-glucosidase precursor (EC 3.2.1.21) (Ge... 66 2e-10
BGLS_BUTFI (P16084) Beta-glucosidase A (EC 3.2.1.21) (Gentiobias... 66 2e-10
BGL2_SACFI (P22507) Beta-glucosidase 2 precursor (EC 3.2.1.21) (... 64 1e-09
BGLS_SCHCO (P29091) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)... 63 2e-09
BGLS_RUMAL (P15885) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)... 59 2e-08
BGL1_SACFI (P22506) Beta-glucosidase 1 precursor (EC 3.2.1.21) (... 59 4e-08
Y537_PASMU (Q9CN98) Hypothetical UPF0190 protein PM0537 32 4.1
CB21_PINTH (P10049) Chlorophyll a-b binding protein type I, chlo... 32 5.3
TRUB_WOLSU (Q7M9J1) tRNA pseudouridine synthase B (EC 4.2.1.70) ... 31 6.9
HISX_SHEON (Q8EFB1) Histidinol dehydrogenase (EC 1.1.1.23) (HDH) 31 6.9
WN2B_MOUSE (O70283) Wnt-2b protein precursor (Wnt-13) 31 9.1
BCPA_PROAE (P11741) Bacteriochlorophyll A protein (BChl a protei... 31 9.1
>XYNB_PRUPE (P83344) Putative beta-D-xylosidase (EC 3.2.1.-)
(PpAz152) (Fragment)
Length = 461
Score = 430 bits (1106), Expect = e-120
Identities = 211/333 (63%), Positives = 263/333 (78%), Gaps = 7/333 (2%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGIGRY +TIHQ GC++V C ++ FG A AAR ADAT+LV+GLDQSIEAE VDR
Sbjct: 132 PLQGIGRYTRTIHQAGCTDVHCNGNQLFGAAEAAARQADATVLVMGLDQSIEAEFVDRAG 191
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPGHQQ+LVS+VA AS+GPTILVLMSGGP+D+TFAKNDP+++ I+W GYPGQAGG AI
Sbjct: 192 LLLPGHQQELVSRVARASRGPTILVLMSGGPIDVTFAKNDPRISAIIWVGYPGQAGGTAI 251
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR--PSKIGYPGRTYRFYKGPVVYPFG 376
A++LFGTA+PGGKLP+TWYPQ Y+ +L MT+MAMR P++ GYPGRTYRFY GPVV+PFG
Sbjct: 252 ANVLFGTANPGGKLPMTWYPQNYVTHLPMTDMAMRADPAR-GYPGRTYRFYIGPVVFPFG 310
Query: 377 HGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLS-IALHVDVKN 435
GL+YT F H L+ PT+VSVP+ + N+ + +K +RV+H C LS + +HVDVKN
Sbjct: 311 LGLSYTTFAHNLAHGPTLVSVPLTSLKATANSTMLSKTVRVSHPDCNALSPLDVHVDVKN 370
Query: 436 VGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDK 495
GS DGTHTLLVF++PP+G W K L+ F K+H+ +++RVR+ +HVCK LSVVD+
Sbjct: 371 TGSMDGTHTLLVFTSPPDG--KWASSKQLMGFHKIHIATGSEKRVRIAVHVCKHLSVVDR 428
Query: 496 SGIRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
GIRRIP+GEH L IGD+ H VSLQ LG IK
Sbjct: 429 FGIRRIPLGEHKLQIGDLSHHVSLQTN-LGEIK 460
>BGLX_SALTY (Q56078) Periplasmic beta-glucosidase precursor (EC
3.2.1.21) (Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase) (T-cell inhibitor)
Length = 765
Score = 102 bits (253), Expect = 3e-21
Identities = 82/265 (30%), Positives = 126/265 (46%), Gaps = 39/265 (14%)
Query: 229 ALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGG 288
A+ AA+ AD + V+G Q + E RT++ +P Q+DL++ + A K P +LVLM+G
Sbjct: 495 AVQAAKQADVVVAVVGESQGMAHEASSRTNITIPQSQRDLITALKATGK-PLVLVLMNGR 553
Query: 289 PVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQE------YL 342
P + K D + IL + G GG AIAD+LFG +P GKLP++ +P+ Y
Sbjct: 554 P--LALVKEDQQADAILETWFAGTEGGNAIADVLFGDYNPSGKLPIS-FPRSVGQIPVYY 610
Query: 343 KNLAMTNMAMRPSKIG-YPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHG 401
+L T P K Y R + GP +YPFG+GL+YT F + + +S P
Sbjct: 611 SHL-NTGRPYNPEKPNKYTSRYFDEANGP-LYPFGYGLSYTTF----TVSDVTLSSP--- 661
Query: 402 HRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQ 461
T R GK++ + V+V N G R+G + ++ P
Sbjct: 662 ----------------TMQRDGKVTAS--VEVTNTGKREGATVIQMYLQDVTASMS-RPV 702
Query: 462 KSLVAFEKVHVPAKTKQRVRVNIHV 486
K L FEK+ + ++ V I +
Sbjct: 703 KQLKGFEKITLKPGERKTVSFPIDI 727
>BGLS_KLUMA (P07337) Beta-glucosidase precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 845
Score = 98.6 bits (244), Expect = 4e-20
Identities = 86/301 (28%), Positives = 138/301 (45%), Gaps = 43/301 (14%)
Query: 219 ACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKG 278
A DD++ A + A D +L+IGL+ E E DR ++ LP +LV V A+
Sbjct: 557 AIDDDEEIRNAAELAAKHDKAVLIIGLNGEWETEGYDRENMDLPKRTNELVRAVLKANPN 616
Query: 279 PTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYP 338
T++V SG PV+ + + + ++ A Y G G AIAD+L+G P GKL ++W P
Sbjct: 617 -TVIVNQSGTPVEFPWLE---EANALVQAWYGGNELGNAIADVLYGDVVPNGKLSLSW-P 671
Query: 339 QEYLKNLAMTNMAMRPSKIGYPGRT---YRFY---KGPVVYPFGHGLTYTHFVHELSSAP 392
+ N A N ++ Y YR+Y + V +PFG+GL+YT F ++S
Sbjct: 672 FKLQDNPAFLNFKTEFGRVVYGEDIFVGYRYYEKLQRKVAFPFGYGLSYTTFELDISD-- 729
Query: 393 TVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSR-DGTHTLLVFSAP 451
+VT + I + VDVKN G + G+ + V+ +
Sbjct: 730 ----------------------FKVTDDK-----IDISVDVKNTGDKFAGSEVVQVYFSA 762
Query: 452 PNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKS-GIRRIPMGEHSLHI 510
N P K L FEKVH+ K+ V + + + +S ++ G + GE+ + +
Sbjct: 763 LN-SKVSRPVKELKGFEKVHLEPGEKKTVNIELELKDAISYFNEELGKWHVEAGEYLVSV 821
Query: 511 G 511
G
Sbjct: 822 G 822
>BGLS_AGRTU (P27034) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)
(Cellobiase) (Beta-D-glucoside glucohydrolase)
Length = 818
Score = 97.4 bits (241), Expect = 8e-20
Identities = 85/311 (27%), Positives = 141/311 (45%), Gaps = 46/311 (14%)
Query: 223 DKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTIL 282
D A++ AR +D +L++G + + E +D + LPG Q++L+ VA + ++
Sbjct: 530 DAGIAEAVETARKSDIVLLLVGREGEWDTEGLDLPDMRLPGRQEELIEAVAETNPN-VVV 588
Query: 283 VLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYL 342
VL +GGP+++ + KV +L YPGQ G A+AD+LFG P G+LP T +P+
Sbjct: 589 VLQTGGPIEMPWLG---KVRAVLQMWYPGQELGNALADVLFGDVEPAGRLPQT-FPKALT 644
Query: 343 KNLAMT-NMAMRPSKIGYPGRTYRFYKG---------PVVYPFGHGLTYTHFVHELSSAP 392
N A+T + ++ P + G+ + G ++PFG GL YT F AP
Sbjct: 645 DNSAITDDPSIYPGQDGHVRYAEGIFVGYRHHDTREIEPLFPFGFGLGYTRFTW---GAP 701
Query: 393 TVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPP 452
+ + T + + VT VDV N+G R G+ + ++ P
Sbjct: 702 QL-----------SGTEMGADGLTVT------------VDVTNIGDRAGSDVVQLYVHSP 738
Query: 453 NGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIG- 511
N P K L AF K+ + + I L ++G R G++ L +
Sbjct: 739 NARVE-RPFKELRAFAKLKLAPGATGTAVLKIAPRDLAYFDVEAGRFRADAGKYELIVAA 797
Query: 512 ---DVKHSVSL 519
D++ SVS+
Sbjct: 798 SAIDIRASVSI 808
>BGLX_ECOLI (P33363) Periplasmic beta-glucosidase precursor (EC
3.2.1.21) (Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 765
Score = 95.1 bits (235), Expect = 4e-19
Identities = 82/265 (30%), Positives = 120/265 (44%), Gaps = 39/265 (14%)
Query: 229 ALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGG 288
A+ A+ +D + V+G Q + E RT + +P Q+DL++ + A K P +LVLM+G
Sbjct: 495 AVQTAKQSDVVVAVVGEAQGMAHEASSRTDITIPQSQRDLIAALKATGK-PLVLVLMNGR 553
Query: 289 PVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTW------YPQEYL 342
P + K D + IL + G GG AIAD+LFG +P GKLP+++ P Y
Sbjct: 554 P--LALVKEDQQADAILETWFAGTEGGNAIADVLFGDYNPSGKLPMSFPRSVGQIPVYYS 611
Query: 343 K-NLAMTNMAMRPSKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHG 401
N A +P+K Y R + G +YPFG+GL+YT F TV V
Sbjct: 612 HLNTGRPYNADKPNK--YTSRYFDEANG-ALYPFGYGLSYTTF--------TVSDV---- 656
Query: 402 HRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQ 461
K T R GK++ + V V N G R+G + ++ P
Sbjct: 657 -----------KLSAPTMKRDGKVTAS--VQVTNTGKREGATVVQMYLQDVTASMS-RPV 702
Query: 462 KSLVAFEKVHVPAKTKQRVRVNIHV 486
K L FEK+ + Q V I +
Sbjct: 703 KQLKGFEKITLKPGETQTVSFPIDI 727
>BGL1_ASPAC (P48825) Beta-glucosidase 1 precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 860
Score = 80.1 bits (196), Expect = 1e-14
Identities = 66/205 (32%), Positives = 96/205 (46%), Gaps = 22/205 (10%)
Query: 196 LCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQ-----SIE 250
L P Q I A+ + +G S A D+ A+ A +++ + D S++
Sbjct: 459 LVTPEQAI--QAEVLKHKG-SVYAITDNWALSQVETLAKQASVSLVFVNSDAGEGYISVD 515
Query: 251 AETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYP 310
DR +L L + +L+ K AA + TI+V+ S GPV + + P V ILWAG P
Sbjct: 516 GNEGDRNNLTLWKNGDNLI-KAAANNCNNTIVVIHSVGPVLVDEWYDHPNVTAILWAGLP 574
Query: 311 GQAGGAAIADILFGTASPGGKLPVTW------YPQEYLKNLAMTNMAMRPS-----KIGY 359
GQ G ++AD+L+G +PG K P TW Y ++ L N A + I Y
Sbjct: 575 GQESGNSLADVLYGRVNPGAKSPFTWGKTREAYGDYLVRELNNGNGAPQDDFSEGVFIDY 634
Query: 360 PGRTYRFYKGPVVYPFGHGLTYTHF 384
G R +Y FGHGL+YT F
Sbjct: 635 RGFDKR--NETPIYEFGHGLSYTTF 657
>BGLB_CLOTM (P14002) Thermostable beta-glucosidase B (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 754
Score = 70.5 bits (171), Expect = 1e-11
Identities = 71/255 (27%), Positives = 111/255 (42%), Gaps = 38/255 (14%)
Query: 222 DDKQFGPALDAARHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTI 281
D++ A AA +D ++ GL E+E DRT + +P +Q L+ VA +
Sbjct: 390 DEELINEAKKAASSSDVAVVFAGLPDEYESEGFDRTHMSIPENQNRLIEAVAEVQSN-IV 448
Query: 282 LVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEY 341
+VL++G PV++ + KV +L A GQA G + + S GKL T +P +
Sbjct: 449 VVLLNGSPVEMPWI---DKVKSVLEAYLGGQALGGRWR-MCYSVKSIVGKLAET-FPVKL 503
Query: 342 LKNLAMTNMAMRPSKIGYPGRT---YRFY--KG-PVVYPFGHGLTYTHFVHELSSAPTVV 395
N + N ++ Y YR+Y KG ++PFGHGL+YT F + S
Sbjct: 504 SHNPSYLNFPGEDDRVEYKEGLFVGYRYYDTKGIEPLFPFGHGLSYTKFEYSDISV---- 559
Query: 396 SVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGG 455
+ ++S+ +I I + V VKNVG G + ++
Sbjct: 560 ----------DKKDVSDNSI-----------INVSVKVKNVGKMAGKEIVQLYVKDVKSS 598
Query: 456 NHWVPQKSLVAFEKV 470
P+K L FEKV
Sbjct: 599 VR-RPEKELKGFEKV 612
>BGLS_HANAN (P06835) Beta-glucosidase precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 825
Score = 66.2 bits (160), Expect = 2e-10
Identities = 56/197 (28%), Positives = 92/197 (46%), Gaps = 15/197 (7%)
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRT- 257
P GIG A+ Q+ S D A+D+A +ADA I V E VD
Sbjct: 476 PADGIGARAQ---QEKISYEFIGDSWNQAAAMDSALYADAAIEVANSVAGEEIGDVDGNY 532
Query: 258 ----SLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
+L L + L+ +++ + TI+++ SG +D+ ++ V ++++ Y GQ
Sbjct: 533 GDLNNLTLWHNAVPLIKNISSINNN-TIVIVTSGQQIDLEPFIDNENVTAVIYSSYLGQD 591
Query: 314 GGAAIADILFGTASPGGKLPVTWYP--QEYLKNLAMTNMAMRPSK----IGYPGRTYRFY 367
G +A +LFG +P GKLP T +Y+ + ++ K I R + Y
Sbjct: 592 FGTVLAKVLFGDENPSGKLPFTIAKDVNDYIPVIEKVDVPDPVDKFTESIYVDYRYFDKY 651
Query: 368 KGPVVYPFGHGLTYTHF 384
PV Y FG+GL+Y++F
Sbjct: 652 NKPVRYEFGYGLSYSNF 668
>BGLS_BUTFI (P16084) Beta-glucosidase A (EC 3.2.1.21) (Gentiobiase)
(Cellobiase) (Beta-D-glucoside glucohydrolase)
Length = 830
Score = 66.2 bits (160), Expect = 2e-10
Identities = 83/324 (25%), Positives = 141/324 (42%), Gaps = 66/324 (20%)
Query: 231 DAARHADATILVI-------------GLDQSIEAET----------VDRTSLLLPGHQQD 267
DAA +AD I+ I G+D+ I+ E D L ++
Sbjct: 136 DAAAYADTAIIAISRFSGEGWDRKVAGVDREIKCEAKDLVEQGNKIFDHGDFYLTNAEKK 195
Query: 268 LVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTAS 327
+V K+ + I+V+ GG VD T+ K D +++ +L A G GG A A IL G +
Sbjct: 196 MV-KMVKENFSSVIVVMNVGGVVDTTWFKKDDQISSVLMAWQGGIEGGLAAARILLGKVN 254
Query: 328 PGGKLPVTW------YPQEYLKNLAMTNMAMRPSKIGYPGRTYRFY------KGPVVYPF 375
P GKL T+ YP + + + ++ Y G YR++ K V YPF
Sbjct: 255 PSGKLSDTFAARLEDYPS--TEGFHEDDDYVDYTEDIYVG--YRYFETIPGAKEKVNYPF 310
Query: 376 GHGLTYTHF-VHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVK 434
G+GL+YT F + + + P V S + +++++++ +I V V
Sbjct: 311 GYGLSYTTFLLEDYKAEPFVASAADEVGK--SDSDLAD-------------AIVASVTVT 355
Query: 435 NVGSRDGTHTL-LVFSAPPNGGNHWVPQKSLVAFEKVHV--PAKTKQRVRVNIHVCKLLS 491
N+G G + L +SAP G P K L + K + P ++ QRV + +++ + S
Sbjct: 356 NIGKIPGKEVVQLYYSAPQ--GKLGKPAKVLGGYAKTRLLQPGES-QRVTIALYMEDMAS 412
Query: 492 VVDKSGIRR----IPMGEHSLHIG 511
D +++ + GE+ +G
Sbjct: 413 YDDLGKVKKAAWLLEKGEYHFFLG 436
>BGL2_SACFI (P22507) Beta-glucosidase 2 precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 880
Score = 63.5 bits (153), Expect = 1e-09
Identities = 44/145 (30%), Positives = 68/145 (46%), Gaps = 10/145 (6%)
Query: 249 IEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAG 308
I+ D+ ++ L H D + K A + T++V+ S G VD+ + P V I+WAG
Sbjct: 539 IDGNRGDKNNVTL-WHNSDNLIKAVAENCANTVVVITSTGQVDVESFADHPNVTAIVWAG 597
Query: 309 YPGQAGGAAIADILFGTASPGGKLPVTWYPQ--EYLKNLAMTNMAMRPSKIGYPGRT--- 363
G G AIA+ILFG A+P G LP T +Y+ + P
Sbjct: 598 PLGDRSGTAIANILFGNANPSGHLPFTVAKSNDDYIPIVTYNPPNGEPEDNTLAEHDLLV 657
Query: 364 -YRFYKGPVV---YPFGHGLTYTHF 384
YR+++ + Y FG+GL+Y +
Sbjct: 658 DYRYFEEKNIEPRYAFGYGLSYNEY 682
>BGLS_SCHCO (P29091) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)
(Cellobiase) (Beta-D-glucoside glucohydrolase)
(Fragment)
Length = 192
Score = 62.8 bits (151), Expect = 2e-09
Identities = 44/145 (30%), Positives = 66/145 (45%), Gaps = 9/145 (6%)
Query: 196 LCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQ-----SIE 250
L PL I A+ + G + + D A A AD I+ I D ++E
Sbjct: 5 LITPLDAITARAQ---EDGTTVTSSLSDSDTARAAQIAAAADVAIVFISSDSGEGYLTVE 61
Query: 251 AETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYP 310
DR LL LV VA A++ + V G + + ++ P V ++W+G P
Sbjct: 62 GNAGDRNDLLAWHDGDALVQAVADANENTIVAVNTVGAIITEAWIEH-PNVKAVVWSGLP 120
Query: 311 GQAGGAAIADILFGTASPGGKLPVT 335
GQ G ++ADIL+G +P G+LP T
Sbjct: 121 GQEAGNSVADILYGAYNPSGRLPYT 145
>BGLS_RUMAL (P15885) Beta-glucosidase (EC 3.2.1.21) (Gentiobiase)
(Cellobiase) (Beta-D-glucoside glucohydrolase)
Length = 947
Score = 59.3 bits (142), Expect = 2e-08
Identities = 72/277 (25%), Positives = 111/277 (39%), Gaps = 47/277 (16%)
Query: 232 AARHADATILVIGLDQSIEAETVDRT-SLLLPGHQQDLVSKVAAASKGPTILVLMSGGPV 290
AA +D I +IG E + + S LL ++ ++ KV + +++L G +
Sbjct: 121 AAESSDTAICIIGRTAGEEQDNSCKAGSYLLTDGEKAILRKVRD-NFSKMVILLNVGNII 179
Query: 291 DITFAKN-DPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLP------VTWYPQEYLK 343
D+ F P +W G G GG A +L G SP GKLP +T YP + K
Sbjct: 180 DMGFIDEFSPDAVMYVWQG--GMTGGTGTARVLLGEVSPCGKLPDTIAYDITDYPSD--K 235
Query: 344 NLAMTNMAMRPSKIGYPGRTY--RFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHG 401
N ++ + I + G Y F K V +PFG+GL+YT F + G
Sbjct: 236 NFHNRDVDIYAEDI-FVGYRYFDTFAKDRVRFPFGYGLSYTQF-----------EISAEG 283
Query: 402 HRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQ 461
+ + I+ K VKN+GS G + V+ PN
Sbjct: 284 RKTDDGVVITAK-------------------VKNIGSAAGKEVVQVYLEAPN-CKLGKAA 323
Query: 462 KSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGI 498
+ L FEK V A +++ + ++ D SGI
Sbjct: 324 RVLCGFEKTKVLAPNEEQTLTIEVTERDIASYDDSGI 360
>BGL1_SACFI (P22506) Beta-glucosidase 1 precursor (EC 3.2.1.21)
(Gentiobiase) (Cellobiase) (Beta-D-glucoside
glucohydrolase)
Length = 876
Score = 58.5 bits (140), Expect = 4e-08
Identities = 47/162 (29%), Positives = 74/162 (45%), Gaps = 30/162 (18%)
Query: 248 SIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWA 307
+++ DR +L L + L+ VA T++V+ S G ++ + P V I+WA
Sbjct: 534 TVDGNQGDRKNLTLWNNGDKLIETVAENCAN-TVVVVTSTGQINFEGFADHPNVTAIVWA 592
Query: 308 GYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPGR----- 362
G G G AIA+ILFG A+P G LP T +A T+ P + P
Sbjct: 593 GPLGDRSGTAIANILFGKANPSGHLPFT---------IAKTDDDYIPIETYSPSSGEPED 643
Query: 363 ----------TYRFYKGPVV---YPFGHGLTYTHFVHELSSA 391
YR+++ + Y FG+GL+Y + E+S+A
Sbjct: 644 NHLVENDLLVDYRYFEEKNIEPRYAFGYGLSYNEY--EVSNA 683
>Y537_PASMU (Q9CN98) Hypothetical UPF0190 protein PM0537
Length = 343
Score = 32.0 bits (71), Expect = 4.1
Identities = 15/42 (35%), Positives = 25/42 (58%), Gaps = 4/42 (9%)
Query: 360 PGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHG 401
PG+ F+ G ++YP+ GLT +H L T++SV ++G
Sbjct: 198 PGQKNHFFGGGLIYPYVEGLTMAEAMHPL----TLLSVGLYG 235
>CB21_PINTH (P10049) Chlorophyll a-b binding protein type I,
chloroplast precursor (CAB) (LHCP)
Length = 266
Score = 31.6 bits (70), Expect = 5.3
Identities = 29/114 (25%), Positives = 49/114 (42%), Gaps = 7/114 (6%)
Query: 248 SIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWA 307
+I++ ++ +LL P Q +LV KV A T+ + P I + + PK G
Sbjct: 6 AIQSSSLAGQTLLRP-QQNELVKKVGTAQARITMRRTVRSAPESIWYGPDRPKYLGPFSE 64
Query: 308 GYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPSKIGYPG 361
G P G D + TA+ V+ P+ + KN + + R + +G G
Sbjct: 65 GTPSYLTGEFPGDYGWDTAA------VSADPETFAKNRELEVIHCRWAMLGALG 112
>TRUB_WOLSU (Q7M9J1) tRNA pseudouridine synthase B (EC 4.2.1.70)
(tRNA pseudouridine 55 synthase) (Psi55 synthase)
(Pseudouridylate synthase) (Uracil hydrolyase)
Length = 291
Score = 31.2 bits (69), Expect = 6.9
Identities = 19/57 (33%), Positives = 25/57 (43%), Gaps = 3/57 (5%)
Query: 356 KIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISN 412
K GY G F KG +V FGH YT L+ P V + H ++ +I N
Sbjct: 48 KAGYSGTLDPFAKGVLVVAFGH---YTRLFDHLNQEPKVYRATLWLGLHSDSLDIEN 101
>HISX_SHEON (Q8EFB1) Histidinol dehydrogenase (EC 1.1.1.23) (HDH)
Length = 438
Score = 31.2 bits (69), Expect = 6.9
Identities = 26/89 (29%), Positives = 41/89 (45%), Gaps = 12/89 (13%)
Query: 244 GLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTIL-----VLMSGGPVD--ITFAK 296
G+ + +E +++ L +PG L+S V + TI VL+S P++ I +A
Sbjct: 119 GVRCELRSEPIEKVGLYIPGGSAPLISTVLMLALPATIAGCEQRVLVSPPPINDAIVYAA 178
Query: 297 NDPKVAGILWAGYPGQAGGAAIADILFGT 325
N + I G G AIA + FGT
Sbjct: 179 NVCGITEIYQVG-----GAQAIAALAFGT 202
>WN2B_MOUSE (O70283) Wnt-2b protein precursor (Wnt-13)
Length = 389
Score = 30.8 bits (68), Expect = 9.1
Identities = 27/104 (25%), Positives = 39/104 (36%), Gaps = 2/104 (1%)
Query: 179 CGCHWAQFRCHSYHDWKLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADA 238
C CH C W+ + G Y + + A +D F A RHA
Sbjct: 235 CKCHGVSGSCTLRTCWRALSDFRRTGDYLRRRYDGAVQVTATQDGANFTAARQGYRHATR 294
Query: 239 TILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTIL 282
T LV D S + +D+ S L G + SK + + G I+
Sbjct: 295 TDLVY-FDNSPDYCVLDKASGSL-GTAGRVCSKTSKGTDGCEIM 336
>BCPA_PROAE (P11741) Bacteriochlorophyll A protein (BChl a protein)
(BCP)
Length = 366
Score = 30.8 bits (68), Expect = 9.1
Identities = 18/66 (27%), Positives = 32/66 (48%), Gaps = 4/66 (6%)
Query: 461 QKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
QK +V F + +N+ V + +++ RRI +GE SL +GD HS S +
Sbjct: 60 QKGVVRFTTKIESVVDSVKNTLNVEV----DIANETKDRRIAVGEGSLSVGDFSHSFSFE 115
Query: 521 AEALGI 526
+ + +
Sbjct: 116 GQVVNM 121
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.337 0.146 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 61,925,876
Number of Sequences: 164201
Number of extensions: 2597361
Number of successful extensions: 8160
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 8122
Number of HSP's gapped (non-prelim): 25
length of query: 528
length of database: 59,974,054
effective HSP length: 115
effective length of query: 413
effective length of database: 41,090,939
effective search space: 16970557807
effective search space used: 16970557807
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC145329.13


