

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145221.2 - phase: 0
(704 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
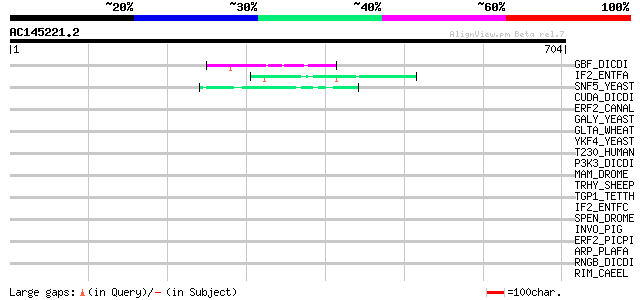
Score E
Sequences producing significant alignments: (bits) Value
GBF_DICDI (P36417) G-box binding factor (GBF) 47 2e-04
IF2_ENTFA (Q835U8) Translation initiation factor IF-2 45 7e-04
SNF5_YEAST (P18480) Transcription regulatory protein SNF5 (SWI/S... 45 9e-04
CUDA_DICDI (O00841) Putative transcriptional regulator cudA 44 0.002
ERF2_CANAL (O13354) Eukaryotic peptide chain release factor GTP-... 43 0.003
GALY_YEAST (P19659) Transcription regulatory protein GAL11 41 0.012
GLTA_WHEAT (P10385) Glutenin, low molecular weight subunit precu... 40 0.028
YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4... 39 0.036
T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein ... 38 0.10
P3K3_DICDI (P54675) Phosphatidylinositol 3-kinase 3 (EC 2.7.1.13... 38 0.10
MAM_DROME (P21519) Neurogenic protein mastermind 38 0.10
TRHY_SHEEP (P22793) Trichohyalin 37 0.23
TGP1_TETTH (O43952) G-quartet DNA binding protein TGP1 37 0.23
IF2_ENTFC (P18311) Translation initiation factor IF-2 37 0.23
SPEN_DROME (Q8SX83) Split ends protein 36 0.30
INVO_PIG (P18175) Involucrin 36 0.30
ERF2_PICPI (P23637) Eukaryotic peptide chain release factor GTP-... 36 0.30
ARP_PLAFA (P04931) Asparagine-rich protein (AG319) (ARP) (Fragment) 36 0.30
RNGB_DICDI (Q7M3S9) Ring finger protein B (Protein rngB) 36 0.40
RIM_CAEEL (Q22366) Rab-3 interacting molecule unc-10 (Rim) (Unco... 36 0.40
>GBF_DICDI (P36417) G-box binding factor (GBF)
Length = 708
Score = 46.6 bits (109), Expect = 2e-04
Identities = 39/170 (22%), Positives = 78/170 (44%), Gaps = 13/170 (7%)
Query: 250 QMEEMMKQMKEMPKLVKEQIQKEQRHQQV-----HFCEMCSGDHPTGYCPPQLEEEVNYV 304
Q ++ +Q ++ P+ +Q+Q++Q HQQ+ H +M H QL++ +
Sbjct: 135 QQQQHHQQQQQQPQH-HQQMQQQQHHQQMQQQQQHHQQMQQQQHHQQMQHHQLQQHQHQH 193
Query: 305 GNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQ 364
QQ+Q Q Q Q+ QQ+ ++ Q+ + QP + + + QQ +N QQ+ Q
Sbjct: 194 QQQQQQQQHQQQHHQQQQQQQ-QQHHQQQQHHQHSQPQQQHQ-HNQQQQHQHNQQQHQQQ 251
Query: 365 QDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKL 414
Q++ Q + +++N NN + ++ + N +H+L
Sbjct: 252 QNQIQMVPQ-----QPQSLSNSGNNNNNNNNNNNSNNNNNNNNNNNSHQL 296
Score = 39.7 bits (91), Expect = 0.028
Identities = 38/163 (23%), Positives = 66/163 (40%), Gaps = 12/163 (7%)
Query: 249 QQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQ 308
QQ ++ +QM++ + Q + Q+HQ H + H + Q +++ + QQ
Sbjct: 163 QQQQQHHQQMQQQQHHQQMQHHQLQQHQHQHQQQQQQQQHQQQHHQQQQQQQQQHHQQQQ 222
Query: 309 --RQGQFQGQYSGNANQQRFPNNNNFQRGNNF-----RQPWK-SNAGPSNQQPFNNNFQQ 360
+ Q Q Q+ N QQ N Q+ N +QP SN+G +N NNN
Sbjct: 223 HHQHSQPQQQHQHNQQQQHQHNQQQHQQQQNQIQMVPQQPQSLSNSGNNN----NNNNNN 278
Query: 361 YPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKK 403
+ + +Q LT++ + + S K K+K
Sbjct: 279 NNSNNNNNNNNNNNSHQLNNLTLSQNNTSGSNTPSPSTKGKRK 321
Score = 37.0 bits (84), Expect = 0.18
Identities = 35/162 (21%), Positives = 62/162 (37%), Gaps = 9/162 (5%)
Query: 249 QQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQ 308
QQ + +Q ++ + ++ Q++Q+ QQ H + H Q ++ + NQQ
Sbjct: 187 QQHQHQHQQQQQQQQHQQQHHQQQQQQQQQHHQQQQHHQHSQPQQQHQHNQQQQHQHNQQ 246
Query: 309 RQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRT 368
+ Q Q Q Q + +N+ NN +N SN NNN + T
Sbjct: 247 QHQQQQNQIQMVPQQPQSLSNSGNNNNNN------NNNNNSNNNNNNNNNNNSHQLNNLT 300
Query: 369 QKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVP 410
+T + + R H E+ +KK G+ +P
Sbjct: 301 LSQNNTSGS--NTPSPSTKGKRKHHETS-NSEKKDSSGQTIP 339
>IF2_ENTFA (Q835U8) Translation initiation factor IF-2
Length = 798
Score = 45.1 bits (105), Expect = 7e-04
Identities = 54/223 (24%), Positives = 84/223 (37%), Gaps = 26/223 (11%)
Query: 306 NQQRQGQFQGQYSGNA----NQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQY 361
NQ R GQ Q N NQ R NNN N R +N G + + N +
Sbjct: 109 NQNRHGQSNTQNRSNQTNTNNQNRNTQNNNGSTTNQNRTSQNNNGGNNQNRGGQNRNNNF 168
Query: 362 PPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHK-------L 414
Q+R + N F NQ NR +K+ K +++ VPA K L
Sbjct: 169 GGGQNRNNR-----NNFN-----NQNRNRFNKKGKKGKHQQESAKPAVPARKFRELPDVL 218
Query: 415 PYPKAQSKKDLEKHFKRF-LDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDE 473
Y + + D+ K R +I K+L + Q K E+L+ + + E
Sbjct: 219 EYTEGMNVADIAKKIHREPAEIIKKL---FMMGVMVNQNQALDKDTIELLAVDYGMEPQE 275
Query: 474 TIQLD-ANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALI 515
+Q+D A+ + +E R + +G V+ GK +
Sbjct: 276 KVQVDIADIDKFFEPEAVVEENLTTRPPVVTIMGHVDHGKTTL 318
>SNF5_YEAST (P18480) Transcription regulatory protein SNF5 (SWI/SNF
complex component SNF5) (Transcription factor TYE4)
Length = 905
Score = 44.7 bits (104), Expect = 9e-04
Identities = 48/201 (23%), Positives = 82/201 (39%), Gaps = 30/201 (14%)
Query: 242 AQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEV 301
AQT+L +Q +++ Q ++ +L + QIQ++Q+ Q H ++ Q +++
Sbjct: 179 AQTQL--EQQRQLLVQQQQQQQL-RNQIQRQQQQQFRHHVQIQQ---------QQQKQQQ 226
Query: 302 NYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQY 361
+QQ+Q Q Q Q QQ+ Q+ +Q + S Q P +
Sbjct: 227 QQQQHQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQGQIPQSQQVPQVRSMSGQ 286
Query: 362 PPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQS 421
PP ++ T+ Q QL N + K ++ D P KLPYP S
Sbjct: 287 PP-----TNVQPTIGQLPQLPKLN-----LPKYQTIQYDP--------PETKLPYPTYWS 328
Query: 422 KKDLEKHFKRFLDIFKRLEIN 442
K + + I +R +IN
Sbjct: 329 DKKADTDTLLYEQIIQRDKIN 349
>CUDA_DICDI (O00841) Putative transcriptional regulator cudA
Length = 791
Score = 43.5 bits (101), Expect = 0.002
Identities = 38/180 (21%), Positives = 62/180 (34%), Gaps = 23/180 (12%)
Query: 268 QIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFP 327
+I+ +Q+ QQ ++C D + + QQ+Q Q + N N
Sbjct: 337 RIEHQQKEQQRLLKKLCYHDKENNI--------IQLIQQQQQQQQLLNNVTNNINNNNNI 388
Query: 328 NNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQ 387
NNNN NN +N +N NNN + T T N
Sbjct: 389 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNN--------NNTTSTTTTTTTTTSSCNNNNN 440
Query: 388 NNRIHKE-------SDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLEKHFKRFLDIFKRLE 440
NN + E ++ + + + P SK + + FK F+ FK+L+
Sbjct: 441 NNNENTEHIVKIENTECNNNNNNIINNTENDENINKPILNSKDEFQSAFKEFIGAFKQLQ 500
Score = 36.2 bits (82), Expect = 0.30
Identities = 36/158 (22%), Positives = 62/158 (38%), Gaps = 26/158 (16%)
Query: 249 QQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHP-----------TGYCPPQL 297
Q ++ +QM++ P+ ++Q Q++Q++QQ G P T Y PQL
Sbjct: 616 QVQQQQQQQMQQQPQQQQQQPQQQQQNQQ-------QGQQPQQQQQQGQLDYTTYIDPQL 668
Query: 298 -------EEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSN 350
++ + +QQ+Q Q+ Q QQ+ + + Q +
Sbjct: 669 QMQQAAQQQYMQQTMDQQQQQQYYMQQYHLQQQQQQQQAQRYLMQQQYMQQQAQQQQQQH 728
Query: 351 QQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQN 388
QQ QQ QQ++ Q+ + NQ L N N
Sbjct: 729 QQVAIQQQQQQNQQQNQQQQNQSPQNQ-QSLDFQNANN 765
Score = 35.0 bits (79), Expect = 0.68
Identities = 35/150 (23%), Positives = 60/150 (39%), Gaps = 18/150 (12%)
Query: 242 AQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEV 301
+Q ++ QQ ++M +Q ++ + ++Q Q +Q+ QQ + T Y PQL+ +
Sbjct: 613 SQQQVQQQQQQQMQQQPQQQQQQPQQQQQNQQQGQQPQQQQQQGQLDYTTYIDPQLQMQ- 671
Query: 302 NYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQ- 360
Q Q Q+ Q QQ++ + Q + QQ QQ
Sbjct: 672 -----QAAQQQYMQQTMDQQQQQQY-----------YMQQYHLQQQQQQQQAQRYLMQQQ 715
Query: 361 YPPQQDRTQKIEDTLNQFMQLTMANQQNNR 390
Y QQ + Q+ + Q NQQ N+
Sbjct: 716 YMQQQAQQQQQQHQQVAIQQQQQQNQQQNQ 745
Score = 33.9 bits (76), Expect = 1.5
Identities = 32/166 (19%), Positives = 70/166 (41%), Gaps = 24/166 (14%)
Query: 245 KLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYV 304
+L QQ+++ + +++ + ++Q+Q++ + QQ PQ +++
Sbjct: 601 QLQQQQIQQQQQSQQQVQQQQQQQMQQQPQQQQQQ---------------PQQQQQNQQQ 645
Query: 305 GNQQRQGQFQGQ--YSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYP 362
G Q +Q Q QGQ Y+ + Q Q +Q + QQ Q +
Sbjct: 646 GQQPQQQQQQGQLDYTTYIDPQ-------LQMQQAAQQQYMQQTMDQQQQQQYYMQQYHL 698
Query: 363 PQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKN 408
QQ + Q+ + L Q + QQ + H++ +++ +++ +N
Sbjct: 699 QQQQQQQQAQRYLMQQQYMQQQAQQQQQQHQQVAIQQQQQQNQQQN 744
>ERF2_CANAL (O13354) Eukaryotic peptide chain release factor
GTP-binding subunit (ERF2) (Translation release factor
3) (ERF3) (ERF-3)
Length = 715
Score = 42.7 bits (99), Expect = 0.003
Identities = 38/165 (23%), Positives = 72/165 (43%), Gaps = 11/165 (6%)
Query: 266 KEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQG-----QYSGN 320
++Q Q++Q+ QQ ++ + + + P ++ QQ+Q Q+ G QY G
Sbjct: 14 QQQQQQQQQQQQNYY----NPNAAQSFVPQGGYQQFQQFQPQQQQQQYGGYNQYNQYQGG 69
Query: 321 ANQQRFPNNNNFQRGNNFRQPWKSN-AGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFM 379
QQ + N +Q+G N R ++ N Q +N N Q QQ +Q + Q
Sbjct: 70 Y-QQNYNNRGGYQQGYNNRGGYQQNYNNRGGYQGYNQNQQYGGYQQYNSQPQQQQQQQSQ 128
Query: 380 QLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKD 424
+++A+ Q + +++ + K K+ K + + A K D
Sbjct: 129 GMSLADFQKQKTEQQASLNKPAVKKTLKLAGSSGIKLANATKKVD 173
>GALY_YEAST (P19659) Transcription regulatory protein GAL11
Length = 1081
Score = 40.8 bits (94), Expect = 0.012
Identities = 42/171 (24%), Positives = 68/171 (39%), Gaps = 17/171 (9%)
Query: 243 QTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQR-HQQVHFCEMCSGDHPTGYCPPQLEEEV 301
Q + LT Q ++++ QMK P + K+ +Q+ ++ + + PQ E
Sbjct: 158 QRRQLTPQQQQLVNQMKVAP-IPKQLLQRIPNIPPNINTWQQVTALAQQKLLTPQDMEAA 216
Query: 302 N--YVGNQQ-------RQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQ 352
Y +QQ +Q Q Q Q N N P N N NN P + P N
Sbjct: 217 KEVYKIHQQLLFKARLQQQQAQAQAQANNNNNGLPQNGNIN--NNINIPQQQQMQPPNSS 274
Query: 353 PFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKK 403
NN Q QQ + + LNQ Q+ +Q + + + + K+ +K
Sbjct: 275 ANNNPLQ----QQSSQNTVPNVLNQINQIFSPEEQRSLLQEAIETCKNFEK 321
>GLTA_WHEAT (P10385) Glutenin, low molecular weight subunit
precursor
Length = 356
Score = 39.7 bits (91), Expect = 0.028
Identities = 34/144 (23%), Positives = 58/144 (39%), Gaps = 3/144 (2%)
Query: 243 QTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVN 302
Q +QQ + Q ++ P ++Q Q+ QQ F + P P + ++
Sbjct: 26 QAPPFSQQQQPPFSQQQQPPFSQQQQSPFSQQQQQPPFAQQ---QQPPFSQQPPISQQQQ 82
Query: 303 YVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYP 362
+QQ+Q QF Q +QQ+ P + Q+ +Q + Q PF QQ
Sbjct: 83 PPFSQQQQPQFSQQQQPPYSQQQQPPYSQQQQPPFSQQQQPPFSQQQQQPPFTQQQQQQQ 142
Query: 363 PQQDRTQKIEDTLNQFMQLTMANQ 386
QQ TQ+ + +Q ++ Q
Sbjct: 143 QQQPFTQQQQPPFSQQPPISQQQQ 166
>YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4
intergenic region
Length = 738
Score = 39.3 bits (90), Expect = 0.036
Identities = 37/137 (27%), Positives = 53/137 (38%), Gaps = 18/137 (13%)
Query: 241 LAQTKLLTQQMEEMMKQMKEMPKLVKE-QIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEE 299
L+Q + QQ ++ + + P+ + Q QK+ + M P GY P + +
Sbjct: 468 LSQQQQYAQQQQQHPQPQSQQPQSQQSPQSQKQGNNVAAQQYYMYQNQFP-GYSYPGMFD 526
Query: 300 EVNYVGNQQRQ--GQFQGQYSGNANQQRFPNNNNFQRGNN-------------FRQPWKS 344
Y QQ Q Q Q SGNANQ F Q G N P +
Sbjct: 527 SQGYAYGQQYQQLAQNNAQTSGNANQYNFQQGYG-QAGANTAAANLTSAAAAAAASPATA 585
Query: 345 NAGPSNQQPFNNNFQQY 361
+A P QQP+ +F Y
Sbjct: 586 HAQPQQQQPYGGSFMPY 602
Score = 31.6 bits (70), Expect = 7.5
Identities = 29/125 (23%), Positives = 51/125 (40%), Gaps = 16/125 (12%)
Query: 249 QQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEV--NYVGN 306
Q + +Q + P+ ++Q+Q++Q+ QQ P EE++ NY
Sbjct: 401 QPQQPQQQQQPQQPQQPQQQLQQQQQQQQ----------QPVQAQAQAQEEQLSQNYYTQ 450
Query: 307 QQRQGQFQGQYSGN----ANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYP 362
QQ+Q Q Q+ + QQ++ +QP + S +Q N QQY
Sbjct: 451 QQQQQYAQQQHQLQQQYLSQQQQYAQQQQQHPQPQSQQPQSQQSPQSQKQGNNVAAQQYY 510
Query: 363 PQQDR 367
Q++
Sbjct: 511 MYQNQ 515
>T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein
complex 230 kDa component (Trap230) (Activator-recruited
cofactor 240 KDa component) (ARC240) (CAG repeat protein
45) (OPA-containing protein) (Trinucleotide repeat
containing 11)
Length = 2212
Score = 37.7 bits (86), Expect = 0.10
Identities = 34/150 (22%), Positives = 65/150 (42%), Gaps = 24/150 (16%)
Query: 239 ANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLE 298
A + T +L +Q ++ +Q ++ + ++Q Q++Q+ QQ H +
Sbjct: 2075 AGVRSTAILPEQQQQQQQQQQQQQQ--QQQQQQQQQQQQYHI----------------RQ 2116
Query: 299 EEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNN-N 357
++ + QQ+Q Q Q Q QQ+ Q+ +Q + A P QP +
Sbjct: 2117 QQQQQILRQQQQQQQQQQ-----QQQQQQQQQQQQQQQQHQQQQQQQAAPPQPQPQSQPQ 2171
Query: 358 FQQYPPQQDRTQKIEDTLNQFMQLTMANQQ 387
FQ+ QQ + Q+ L + +Q ++N Q
Sbjct: 2172 FQRQGLQQTQQQQQTAALVRQLQQQLSNTQ 2201
>P3K3_DICDI (P54675) Phosphatidylinositol 3-kinase 3 (EC 2.7.1.137)
(PI3-kinase) (PtdIns-3-kinase) (PI3K) (Fragment)
Length = 1585
Score = 37.7 bits (86), Expect = 0.10
Identities = 22/86 (25%), Positives = 43/86 (49%), Gaps = 2/86 (2%)
Query: 320 NANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFM 379
N N NNNN NN +N +N++ NNN + + ++ E++L + +
Sbjct: 348 NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNEELINNNNNNNNDENYKIEETEESLKELL 407
Query: 380 QL-TMANQQNNRIHKE-SDVEKDKKK 403
+ + N++ +I KE ++++ KKK
Sbjct: 408 EKEKLENEEREKILKERNEIDNLKKK 433
>MAM_DROME (P21519) Neurogenic protein mastermind
Length = 1596
Score = 37.7 bits (86), Expect = 0.10
Identities = 48/181 (26%), Positives = 66/181 (35%), Gaps = 51/181 (28%)
Query: 238 DANLAQTKLLTQQMEEMMKQMKEMPKLVK-EQIQKEQRHQQVHFCEMCSGDHPTGYCPPQ 296
D Q + QQ ++ ++ PK+ K+Q+ QQV Q
Sbjct: 754 DLKRLQQQQAMQQQQQQQHHQQQQPKMGGVPNFNKQQQQQQVP--------------QQQ 799
Query: 297 LEEEVNYVGNQQRQGQFQGQYSGNANQQRFPN--------------NNNFQRGNNFRQPW 342
L+++ QQ+Q Q Q QYS +NQ PN N N Q GN +Q
Sbjct: 800 LQQQQQ---QQQQQQQQQQQYSPFSNQN--PNAAANFLNCPPRGGPNGNQQPGNLAQQQQ 854
Query: 343 KSNAGPSNQQPFNN---------------NFQQYPPQQDRTQKIEDTL--NQFMQLTMAN 385
+ AGP QQ N N P QQ + Q TL Q QL ++
Sbjct: 855 QPGAGPQQQQQRGNAGNGQQNNPNTGPGGNTPNAPQQQQQQQSTTTTLQMKQTQQLHISQ 914
Query: 386 Q 386
Q
Sbjct: 915 Q 915
>TRHY_SHEEP (P22793) Trichohyalin
Length = 1549
Score = 36.6 bits (83), Expect = 0.23
Identities = 31/117 (26%), Positives = 54/117 (45%), Gaps = 7/117 (5%)
Query: 213 LNDRAAKHNRSGT----QRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQ 268
LN++ AK R G +R+ D L + +L Q++EE+ ++ + K V+ +
Sbjct: 95 LNEKEAKCERKGNPLQDRRREDQRRFEPQDRQLEERRLKRQELEELAEEEELREKQVRRE 154
Query: 269 IQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQR 325
+ ++R Q+ + E P G +LEE +N +RQ Q + Q QQR
Sbjct: 155 QRLQRREQEEYGGEEELQQRPKG---RELEELLNREQRFERQEQRERQRLQVEQQQR 208
>TGP1_TETTH (O43952) G-quartet DNA binding protein TGP1
Length = 725
Score = 36.6 bits (83), Expect = 0.23
Identities = 30/115 (26%), Positives = 42/115 (36%), Gaps = 12/115 (10%)
Query: 268 QIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFP 327
Q + QR++ H + P Y ++ N Q+ Q Q Q N+Q
Sbjct: 497 QNNRNQRNRDKH--HNSQNNQPNKYKNTSVQNNNNNKNQQRSQSQNQRPPRNYDNRQGGE 554
Query: 328 NNNNFQRGNNFRQPWKSNAGPSNQQ----------PFNNNFQQYPPQQDRTQKIE 372
N NN QR N R + N N Q P NNN+ + QQ +E
Sbjct: 555 NRNNRQRNENNRNNFNGNGHRVNNQNNQRNRNSSYPRNNNYDHHHNQQTDISGLE 609
>IF2_ENTFC (P18311) Translation initiation factor IF-2
Length = 784
Score = 36.6 bits (83), Expect = 0.23
Identities = 55/225 (24%), Positives = 88/225 (38%), Gaps = 26/225 (11%)
Query: 306 NQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQ--PFNNNFQQYPP 363
N Q +GQ GQ + NQ N +N GNN + + +G N+Q N +
Sbjct: 90 NYQDRGQGSGQVNQGKNQSTNQNRSNNGGGNNQNRQGATQSGNQNRQGSTQGGNQNRQGS 149
Query: 364 QQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGK-NVPAHK-------LP 415
Q Q + N+ N NR +K+ +K K++ K VP K L
Sbjct: 150 NQGGNQNRNNNNNRG---KFNNNNRNRFNKKG--KKGKQQTSNKPAVPPRKFRELPEVLE 204
Query: 416 YPKAQSKKDLEKHFKRF-LDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDET 474
Y + + D+ K R +I K+L + Q K E+L+ + + E
Sbjct: 205 YTEGMNVADIAKKIHREPAEIIKKL---FMLGVMVNQNQALDKDTIELLATDYGMEPQEK 261
Query: 475 IQLDANCSAIIQRRLPKKEKDP----GRVTLPVAIGKVNVGKALI 515
IQ+D A I + +E +P R + +G V+ GK +
Sbjct: 262 IQVDI---ADIDKFFEPEELNPDTLVSRPPVVTIMGHVDHGKTTL 303
Score = 33.9 bits (76), Expect = 1.5
Identities = 28/90 (31%), Positives = 41/90 (45%), Gaps = 10/90 (11%)
Query: 305 GNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQ 364
GNQ RQG QG NQ R NNN + NN R + QQ +N PP+
Sbjct: 142 GNQNRQGSNQG-----GNQNRNNNNNRGKFNNNNRNRFNKKGKKGKQQ--TSNKPAVPPR 194
Query: 365 QDRTQKIEDTLNQFMQLTMANQQNNRIHKE 394
+ R ++ + L + +A+ +IH+E
Sbjct: 195 KFR--ELPEVLEYTEGMNVADIA-KKIHRE 221
Score = 32.3 bits (72), Expect = 4.4
Identities = 21/53 (39%), Positives = 24/53 (44%), Gaps = 14/53 (26%)
Query: 305 GNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNN 357
GNQ RQG QG GN N+Q N R NN +N+ FNNN
Sbjct: 131 GNQNRQGSTQG---GNQNRQGSNQGGNQNRNNN-----------NNRGKFNNN 169
>SPEN_DROME (Q8SX83) Split ends protein
Length = 5560
Score = 36.2 bits (82), Expect = 0.30
Identities = 41/198 (20%), Positives = 72/198 (35%), Gaps = 25/198 (12%)
Query: 180 LQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAAKHNRSGTQRKSGILELGSNDA 239
+QP+ A+ L +L P + V + + + Q + + +
Sbjct: 3782 VQPQQATQSQVASSPPLGSLPPHKNVHLNAHQNQQQPQVIAKMTAHQHQQHMQQFMHQQM 3841
Query: 240 NLAQTKLLTQQMEEMMKQMKEMPKLVKEQI----QKEQRHQQVHFCEMCSGD-HPTGYCP 294
Q + QQ+ +Q+ P+ Q Q++Q H Q H + HPT
Sbjct: 3842 IQRQQHMQQQQLHGQSQQITSAPQHQMHQQHQAQQQQQHHNQQHLNQQLHAQQHPT---- 3897
Query: 295 PQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPF 354
Q+Q Q Q Q++ Q + + Q+ N +Q S +QQ
Sbjct: 3898 -------------QKQHQAQQQFNQQIQQHQSQQQHQVQQQNQAQQQHLSQQQHQSQQQL 3944
Query: 355 NNNFQQYPPQQDRTQKIE 372
N QQ+ QQ + Q+I+
Sbjct: 3945 N---QQHQAQQQQLQQIQ 3959
Score = 35.8 bits (81), Expect = 0.40
Identities = 46/254 (18%), Positives = 91/254 (35%), Gaps = 34/254 (13%)
Query: 181 QPEPKLILDAAAHGSLMALS---------PREAVDIINKMSLNDRAAKHNRSGTQRKSGI 231
QP PK + + LM + P I +++ + A ++ + G
Sbjct: 3741 QPSPKSVAEVQTTPQLMTIPLQKMTPIQVPHHPTIISKVVTVQPQQATQSQVASSPPLGS 3800
Query: 232 LELGSN-DANLAQTKLLTQQMEEMM--KQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDH 288
L N N Q + Q + +M + + M + + +Q+ + Q+H Q + G
Sbjct: 3801 LPPHKNVHLNAHQNQQQPQVIAKMTAHQHQQHMQQFMHQQMIQRQQHMQQ---QQLHGQS 3857
Query: 289 PTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGP 348
PQ + + QQ+Q Q + + Q+ P Q
Sbjct: 3858 QQITSAPQHQMHQQHQAQQQQQHHNQQHLNQQLHAQQHPTQKQHQA-------------- 3903
Query: 349 SNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKN 408
QQ FN QQ+ QQ + ++ Q +Q +++++ ++ + +++ K
Sbjct: 3904 --QQQFNQQIQQHQSQQQHQVQQQNQAQQQHLSQQQHQSQQQLNQQHQAQQQQLQQIQKL 3961
Query: 409 VPAHKLPYPKAQSK 422
H P+ Q K
Sbjct: 3962 QQMHG---PQQQQK 3972
>INVO_PIG (P18175) Involucrin
Length = 347
Score = 36.2 bits (82), Expect = 0.30
Identities = 40/226 (17%), Positives = 93/226 (40%), Gaps = 11/226 (4%)
Query: 249 QQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQ 308
QQ ++ Q++E+ + ++Q Q+E + Q++H + E+E++ QQ
Sbjct: 91 QQQQQQESQVQEL-HVDQQQQQQESQEQELHVDQQQQQQE-------SQEQELHVDQQQQ 142
Query: 309 RQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAG-PSNQQPFNNNFQQYPPQQDR 367
++ Q Q + G+ QQ+ ++ +Q +QQ Q+ D+
Sbjct: 143 QESQVQELHVGHHQQQQESQEQELHVDHHQQQQESQEQELHVDQQQQQQESQEQELHVDQ 202
Query: 368 TQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLEK 427
Q+ +++ Q + + QQ +E V+ ++++ + H + + Q ++ E
Sbjct: 203 QQQQQESQEQELHVDHHQQQQESQVQELHVDHQQQQQESQEQELHVDQHQQQQESQEQEL 262
Query: 428 HFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDE 473
H + + E+ + EQ + K E L ++ +E +
Sbjct: 263 HVDQQQQELQVQEVQQQQQQQQEQQEDHQK--AEHLEQEEAQREQQ 306
>ERF2_PICPI (P23637) Eukaryotic peptide chain release factor
GTP-binding subunit (ERF2) (Translation release factor
3) (ERF3) (ERF-3) (Omnipotent suppressor protein 2)
Length = 741
Score = 36.2 bits (82), Expect = 0.30
Identities = 38/163 (23%), Positives = 66/163 (40%), Gaps = 35/163 (21%)
Query: 292 YCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRFPNN----NNFQRG------------ 335
Y P Q E++ G QQ+Q QG Y+ N+ + NN NN RG
Sbjct: 52 YIPGQQEQQFGQYG-QQQQNYNQGGYNNYNNRGGYSNNRGGYNNSNRGGYSNYNSYNTNS 110
Query: 336 -----NNFRQPWKSNAGPSNQQPFNNNFQQ-------YPPQQDRTQKIEDTLNQFMQLTM 383
+N+ + +N+ +N +NNN+ Q P QD+ Q+ Q+++
Sbjct: 111 NQGGYSNYNNNYANNSYNNNNN-YNNNYNQGYNNYNSQPQGQDQQQETGSG-----QMSL 164
Query: 384 ANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSKKDLE 426
+ Q + + + KK + N+ + + P KK+ E
Sbjct: 165 EDYQKQQKESLNKLNTKPKKVLKLNLNSSTVKAPIVTKKKEEE 207
Score = 31.2 bits (69), Expect = 9.8
Identities = 30/148 (20%), Positives = 56/148 (37%), Gaps = 31/148 (20%)
Query: 306 NQQRQGQFQG--QYSGNANQQRFPN------NNNFQRGNNFRQPWKSNAGPSNQQPFNNN 357
N +G + Y+ N+NQ + N NN++ NN+ + N QP +
Sbjct: 93 NNSNRGGYSNYNSYNTNSNQGGYSNYNNNYANNSYNNNNNYNNNYNQGYNNYNSQPQGQD 152
Query: 358 FQQ-----------YPPQQ-DRTQKIEDTLNQFMQLTM-----------ANQQNNRIHKE 394
QQ Y QQ + K+ + ++L + ++ +++E
Sbjct: 153 QQQETGSGQMSLEDYQKQQKESLNKLNTKPKKVLKLNLNSSTVKAPIVTKKKEEEPVNQE 212
Query: 395 SDVEKDKKKEVGKNVPAHKLPYPKAQSK 422
S E+ K+E+ PA + +SK
Sbjct: 213 SKTEEPAKEEIKNQEPAEAENKVEEESK 240
>ARP_PLAFA (P04931) Asparagine-rich protein (AG319) (ARP) (Fragment)
Length = 537
Score = 36.2 bits (82), Expect = 0.30
Identities = 22/92 (23%), Positives = 31/92 (32%), Gaps = 1/92 (1%)
Query: 302 NYVGNQQRQGQFQGQYSGNANQQRFPN-NNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQ 360
N N G F + N N+ N N NN Q NNNF
Sbjct: 197 NNNNNNNNNGVFSNYQNNNMNRNNSINIKRNLNNNNNINNNMNKMGSQDKNQNSNNNFYM 256
Query: 361 YPPQQDRTQKIEDTLNQFMQLTMANQQNNRIH 392
Q+R + + +N M M + NN ++
Sbjct: 257 NYNYQNRKNSMNNNMNNNMNNNMNHNMNNNMN 288
Score = 33.5 bits (75), Expect = 2.0
Identities = 24/97 (24%), Positives = 38/97 (38%), Gaps = 7/97 (7%)
Query: 313 FQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAG-PSNQQPFNNNFQQYPPQQDRTQKI 371
+ Y N N NNN GNN + N N NNN + +
Sbjct: 394 YNNNYLNNNNNMNSAVNNNSSNGNNMKNENSENKNVADNNDSLNNN-----KNNNNNINM 448
Query: 372 EDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKN 408
+++N L N+ NN+ + E D + D E+G++
Sbjct: 449 NESINNNNTLNNNNEYNNQNNNE-DEDDDDWGELGED 484
>RNGB_DICDI (Q7M3S9) Ring finger protein B (Protein rngB)
Length = 943
Score = 35.8 bits (81), Expect = 0.40
Identities = 43/170 (25%), Positives = 73/170 (42%), Gaps = 17/170 (10%)
Query: 243 QTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVN 302
Q LLT Q++ ++ Q E+ L KE + +++H ++ ++ D PT C E V
Sbjct: 619 QVSLLTNQIQSII-QKDELTSLKKEYSELKKKHSLLYSEDI--DDLPTETCLKLEEIHVK 675
Query: 303 YVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYP 362
+ + R + + QQ+ P NN + + P QQ QQ
Sbjct: 676 SL-EKLRVKKLPSSNQLSTLQQQIPQQPTTIICNNSQIIQQQPLPPLQQQQ-----QQQQ 729
Query: 363 PQQDRTQKIEDTLNQFMQLTM--------ANQQNNRIHKESDVEKDKKKE 404
QQ + Q+ + L QLTM + QQ N+I K+ EK ++++
Sbjct: 730 QQQQQQQQQQQPLEIQEQLTMLQLQLSQLSQQQQNQIDKQQKQEKLQQEQ 779
>RIM_CAEEL (Q22366) Rab-3 interacting molecule unc-10 (Rim)
(Uncoordinated protein 10)
Length = 1563
Score = 35.8 bits (81), Expect = 0.40
Identities = 31/123 (25%), Positives = 50/123 (40%), Gaps = 14/123 (11%)
Query: 294 PPQLEEEVNYVGNQQRQGQ---------FQGQYSGNANQQRFPNNNNFQRGNNFRQPWKS 344
PP NY NQQ +G Q + N N + PN N+ Q N + P ++
Sbjct: 167 PPDKMNTPNYQNNQQPRGMGMQPNHNQTQQNFMNQNQNSNQHPNQNHNQ--NQMQNPHQN 224
Query: 345 NAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQ-QNNRIHKESDVEKDKKK 403
N NN QQ + + Q + NQ Q+ NQ QN + ++ ++++
Sbjct: 225 QNHVQNNHQGANNHQQNNRRAMQQQPMSQ--NQANQINQMNQNQNQQQSHNQNMTQNQRN 282
Query: 404 EVG 406
+ G
Sbjct: 283 QTG 285
Score = 35.4 bits (80), Expect = 0.52
Identities = 37/197 (18%), Positives = 70/197 (34%), Gaps = 19/197 (9%)
Query: 209 NKMSLNDRAAKHNRSGTQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQ 268
N + N + A +++ +R + N AN Q + Q + + M + + Q
Sbjct: 226 NHVQNNHQGANNHQQNNRRAMQQQPMSQNQAN--QINQMNQNQNQQQSHNQNMTQNQRNQ 283
Query: 269 I--QKEQRHQQVHFCEMCSGDHPTGYCPPQLEEEVNYVGNQQRQGQFQGQYSGNANQQRF 326
Q +QR + P+ Y + V Q Q Q+ N+QR
Sbjct: 284 TGPQNQQRTNDSRTMKQTPQQQPSQY-----QNNVGAAHQHHNQHGQQEQHHQQMNEQRT 338
Query: 327 PNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQ 386
N N R+ G N+QP + +Q P +D +NQ N
Sbjct: 339 DN-------NRMRENTNGQGGMFNRQP---SLEQTTPMNKYNHVEDDGMNQRPTFYTGNS 388
Query: 387 QNNRIHKESDVEKDKKK 403
+N++ + +++ ++
Sbjct: 389 ENDQRQFDGQMQQGSQQ 405
Score = 33.9 bits (76), Expect = 1.5
Identities = 24/110 (21%), Positives = 45/110 (40%), Gaps = 3/110 (2%)
Query: 302 NYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQR---GNNFRQPWKSNAGPSNQQPFNNNF 358
N+ N +R Q Q ANQ N N Q+ N Q ++ GP NQQ N++
Sbjct: 237 NHQQNNRRAMQQQPMSQNQANQINQMNQNQNQQQSHNQNMTQNQRNQTGPQNQQRTNDSR 296
Query: 359 QQYPPQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKN 408
Q + + ++ + Q + Q + H++ + ++ + +N
Sbjct: 297 TMKQTPQQQPSQYQNNVGAAHQHHNQHGQQEQHHQQMNEQRTDNNRMREN 346
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 85,425,312
Number of Sequences: 164201
Number of extensions: 3845783
Number of successful extensions: 14886
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 16
Number of HSP's successfully gapped in prelim test: 119
Number of HSP's that attempted gapping in prelim test: 14356
Number of HSP's gapped (non-prelim): 424
length of query: 704
length of database: 59,974,054
effective HSP length: 117
effective length of query: 587
effective length of database: 40,762,537
effective search space: 23927609219
effective search space used: 23927609219
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 69 (31.2 bits)
Medicago: description of AC145221.2


