

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.10 + phase: 0
(235 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
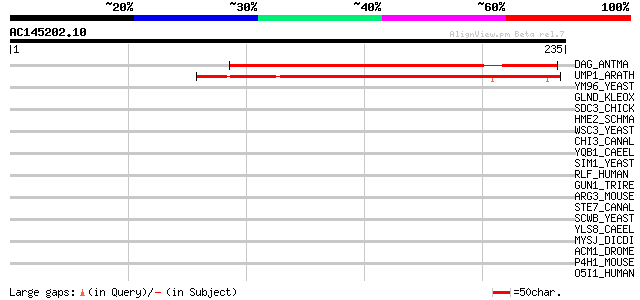
Score E
Sequences producing significant alignments: (bits) Value
DAG_ANTMA (Q38732) DAG protein, chloroplast precursor 139 4e-33
UMP1_ARATH (Q9LKA5) Unknown mitochondrial protein At3g15000 137 3e-32
YM96_YEAST (Q04893) Hypothetical 113.1 kDa protein in PRE5-FET4 ... 37 0.053
GLND_KLEOX (P41393) [Protein-PII] uridylyltransferase (EC 2.7.7.... 35 0.12
SDC3_CHICK (P26261) Syndecan-3 precursor 35 0.20
HME2_SCHMA (Q26601) Homeobox protein engrailed-like SMOX-2 32 1.00
WSC3_YEAST (Q12215) Cell wall integrity and stress response comp... 32 1.3
CHI3_CANAL (P40954) Chitinase 3 precursor (EC 3.2.1.14) 32 1.7
YQB1_CAEEL (Q09255) Hypothetical protein C30G12.1 in chromosome II 31 2.2
SIM1_YEAST (P40472) SIM1 protein precursor 31 2.2
RLF_HUMAN (Q13129) Zinc finger protein Rlf (Rearranged L-myc fus... 31 2.2
GUN1_TRIRE (P07981) Endoglucanase EG-1 precursor (EC 3.2.1.4) (E... 31 2.2
ARG3_MOUSE (Q9D8S3) ADP-ribosylation factor GTPase-activating pr... 31 2.2
STE7_CANAL (P46599) Serine/threonine-protein kinase STE7 homolog... 31 2.9
SCWB_YEAST (P53189) Probable family 17 glucosidase SCW11 precurs... 31 2.9
YLS8_CAEEL (P34393) Hypothetical protein F09G8.8 in chromosome I... 30 3.8
MYSJ_DICDI (P54697) Myosin IJ heavy chain 30 3.8
ACM1_DROME (P16395) Muscarinic acetylcholine receptor DM1 30 3.8
P4H1_MOUSE (Q60715) Prolyl 4-hydroxylase alpha-1 subunit precurs... 30 4.9
O5I1_HUMAN (Q13606) Olfactory receptor 5I1 (Olfactory receptor-l... 30 4.9
>DAG_ANTMA (Q38732) DAG protein, chloroplast precursor
Length = 230
Score = 139 bits (351), Expect = 4e-33
Identities = 69/139 (49%), Positives = 90/139 (64%), Gaps = 7/139 (5%)
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
+ PGCDYNHWLI+++ P T++QMID Y+ TLA VLGS EEAKK +Y S Y GF
Sbjct: 80 MLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLNTLATVLGSMEEAKKNMYAFSTTTYTGF 139
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQ 213
C + EETS K +G+PGVL+VLPDSY+D +++DYG + +VNGEI+ P Q
Sbjct: 140 QCTVTEETSEKFKGLPGVLWVLPDSYIDVKNKDYGGDKYVNGEIIPCQ-------YPTYQ 192
Query: 214 RGDSRPRYHDRTKYVRRRD 232
SR + YVR+RD
Sbjct: 193 PKQSRSSKYKSKAYVRQRD 211
>UMP1_ARATH (Q9LKA5) Unknown mitochondrial protein At3g15000
Length = 395
Score = 137 bits (344), Expect = 3e-32
Identities = 69/161 (42%), Positives = 107/161 (65%), Gaps = 9/161 (5%)
Query: 80 NTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAK 139
N ++S+RPP E L GCD+ HWL++++ P GE T+ ++ID Y+KTLAQ++GSE+EA+
Sbjct: 78 NPNWSNRPPKETI-LLDGCDFEHWLVVVEPPQGE-PTRDEIIDSYIKTLAQIVGSEDEAR 135
Query: 140 KKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQ 199
KIY+VS Y+ FG + E+ S+KL+ + V +VLPDSY+D ++DYG E F++G+ V
Sbjct: 136 MKIYSVSTRCYYAFGALVSEDLSHKLKELSNVRWVLPDSYLDVRNKDYGGEPFIDGKAVP 195
Query: 200 RSPE------RQRRVEPQAQRGDSRPRYHDRTK-YVRRRDN 233
P+ R + R + RPR +DR++ + RRR+N
Sbjct: 196 YDPKYHEEWIRNNARANERNRRNDRPRNNDRSRNFERRREN 236
>YM96_YEAST (Q04893) Hypothetical 113.1 kDa protein in PRE5-FET4
intergenic region
Length = 1140
Score = 36.6 bits (83), Expect = 0.053
Identities = 18/44 (40%), Positives = 27/44 (60%)
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSL 59
+TTTS++TST+T TTT + + TSP I+ S T+ S+
Sbjct: 19 TTTTSSTTSTTTPTTTSTTSTTSTKVTTSPEIIVSSSSTLVSSV 62
>GLND_KLEOX (P41393) [Protein-PII] uridylyltransferase (EC 2.7.7.59)
(PII uridylyl-transferase) (Uridylyl removing enzyme)
(UTase)
Length = 887
Score = 35.4 bits (80), Expect = 0.12
Identities = 21/56 (37%), Positives = 29/56 (51%), Gaps = 2/56 (3%)
Query: 13 RLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSL--HFGGIHR 66
R+F T S+ T Y+TTV LT P IP ++T+F S+ H GG+ R
Sbjct: 375 RMFYTMVRNSSITGIYSTTVRHLRHARRHLTQPLCYIPEARTLFLSMLRHQGGLSR 430
>SDC3_CHICK (P26261) Syndecan-3 precursor
Length = 405
Score = 34.7 bits (78), Expect = 0.20
Identities = 17/49 (34%), Positives = 29/49 (58%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIP 50
TT + +S STTT+T+ +T+T TTT++ ++ T+ RF+ P
Sbjct: 135 TTASTTASDSPSTTSTTTTTAATTTTTTTTISTTVATSKPTTTQRFLPP 183
>HME2_SCHMA (Q26601) Homeobox protein engrailed-like SMOX-2
Length = 524
Score = 32.3 bits (72), Expect = 1.00
Identities = 18/44 (40%), Positives = 25/44 (55%)
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSL 59
+TTT+T+T+T+T TTT + SS P II +T SL
Sbjct: 104 TTTTTTTTTTTTSTTTTTSGIIPLSSTIIPSTIISPVKTSLSSL 147
>WSC3_YEAST (Q12215) Cell wall integrity and stress response
component 3 precursor
Length = 556
Score = 32.0 bits (71), Expect = 1.3
Identities = 19/43 (44%), Positives = 26/43 (60%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
TT + SS S+TTS+STS++T +TT + S SS TS
Sbjct: 199 TTSSTTSSTTSSTTSSTTSSSTSSTTSSTTSSTTSSTTSSTTS 241
Score = 31.2 bits (69), Expect = 2.2
Identities = 27/82 (32%), Positives = 40/82 (47%), Gaps = 1/82 (1%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTI-FQSLH 60
TT + SS S+TTS++TS++T +TT + S SS TS + S +I S
Sbjct: 215 TTSSSTSSTTSSTTSSTTSSTTSSTTSSTTSSTTSSTTSSTTSIFSVTSSSSSITLSSSE 274
Query: 61 FGGIHRAGYCNYSPLVPGSNTS 82
+ S LVP S++S
Sbjct: 275 HTTVDSRTSSPSSTLVPVSSSS 296
Score = 30.8 bits (68), Expect = 2.9
Identities = 18/43 (41%), Positives = 26/43 (59%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
TT + SS S+TTS++TS++T +TT + S SS TS
Sbjct: 211 TTSSTTSSSTSSTTSSTTSSTTSSTTSSTTSSTTSSTTSSTTS 253
Score = 30.4 bits (67), Expect = 3.8
Identities = 17/44 (38%), Positives = 27/44 (60%)
Query: 1 MTTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
++T + SS S+TTS++TS++T +TT + S SS TS
Sbjct: 186 VSTTSTTSSSTSSTTSSTTSSTTSSTTSSTTSSSTSSTTSSTTS 229
Score = 30.4 bits (67), Expect = 3.8
Identities = 18/43 (41%), Positives = 26/43 (59%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
+T + SS S+TTS++TS+ST +TT + S SS TS
Sbjct: 195 STSSTTSSTTSSTTSSTTSSTTSSSTSSTTSSTTSSTTSSTTS 237
Score = 30.4 bits (67), Expect = 3.8
Identities = 18/43 (41%), Positives = 26/43 (59%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
TT + SS S+TTS++TS++T +TT + S SS TS
Sbjct: 207 TTSSTTSSTTSSSTSSTTSSTTSSTTSSTTSSTTSSTTSSTTS 249
>CHI3_CANAL (P40954) Chitinase 3 precursor (EC 3.2.1.14)
Length = 567
Score = 31.6 bits (70), Expect = 1.7
Identities = 20/57 (35%), Positives = 29/57 (50%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQS 58
TT +SS STT+STS+S+++ +T+ + S SS S P S T S
Sbjct: 349 TTSTTSSSISSTTSSTTSSTSSSSTSSSTSSTTSSSTTSSQISTTSTAPTSSTSLSS 405
>YQB1_CAEEL (Q09255) Hypothetical protein C30G12.1 in chromosome II
Length = 469
Score = 31.2 bits (69), Expect = 2.2
Identities = 18/48 (37%), Positives = 28/48 (57%), Gaps = 4/48 (8%)
Query: 14 LFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFI----IPMSQTIFQ 57
+F+T T+ +T+T TTT + + P++ T+PR I IP S T Q
Sbjct: 284 VFTTKTNPQITTTTMTTTTRTSTAIPTTTTTPRQISLQTIPFSTTTRQ 331
>SIM1_YEAST (P40472) SIM1 protein precursor
Length = 475
Score = 31.2 bits (69), Expect = 2.2
Identities = 18/43 (41%), Positives = 27/43 (61%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
T + +S+ K ST+ STSTSTST +T+ + + S SS +S
Sbjct: 156 TQSSASSATKSSTSSTSPSTSTSTSTSSTSSSSSSSSSSSSSS 198
Score = 30.4 bits (67), Expect = 3.8
Identities = 19/43 (44%), Positives = 26/43 (60%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
T+ A SS S+T+STS STST T+T + + S SS +S
Sbjct: 152 TSSATQSSASSATKSSTSSTSPSTSTSTSTSSTSSSSSSSSSS 194
>RLF_HUMAN (Q13129) Zinc finger protein Rlf (Rearranged L-myc fusion
gene protein) (Zn-15 related protein)
Length = 1914
Score = 31.2 bits (69), Expect = 2.2
Identities = 24/87 (27%), Positives = 41/87 (46%), Gaps = 9/87 (10%)
Query: 124 YVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPE 183
Y+ L +L +E+ A K+I V C+ C L+ EG FVL +Y+ +
Sbjct: 254 YITCLCSMLPNED-AIKEIAKVDCKEVLDIICNLES------EGQDNTAFVLCTTYLTQQ 306
Query: 184 HQDYGAELFVNGEIVQRSPERQRRVEP 210
Q A ++ + E+ + QRR++P
Sbjct: 307 LQT--ASVYCSWELTLFWSKLQRRIDP 331
>GUN1_TRIRE (P07981) Endoglucanase EG-1 precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Cellulase)
Length = 459
Score = 31.2 bits (69), Expect = 2.2
Identities = 14/54 (25%), Positives = 22/54 (39%)
Query: 50 PMSQTIFQSLHFGGIHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHW 103
P + +F ++ +G I P P S+T+FS + P C HW
Sbjct: 375 PNTHVVFSNIRWGDIGSTTNSTAPPPPPASSTTFSTTRRSSTTSSSPSCTQTHW 428
>ARG3_MOUSE (Q9D8S3) ADP-ribosylation factor GTPase-activating
protein 3 (ARF GAP 3)
Length = 525
Score = 31.2 bits (69), Expect = 2.2
Identities = 17/38 (44%), Positives = 23/38 (59%), Gaps = 3/38 (7%)
Query: 132 LGSEEEAKKKIYNV---SCERYFGFGCELDEETSNKLE 166
+GS +EA+KK NV S + YFG + D ET +LE
Sbjct: 420 IGSTDEAQKKFGNVKAISSDMYFGIQAQTDFETRARLE 457
>STE7_CANAL (P46599) Serine/threonine-protein kinase STE7 homolog
(EC 2.7.1.37)
Length = 589
Score = 30.8 bits (68), Expect = 2.9
Identities = 19/54 (35%), Positives = 32/54 (59%), Gaps = 3/54 (5%)
Query: 19 TSTSTSTSTYTTTVAGALSKPS---SLTSPRFIIPMSQTIFQSLHFGGIHRAGY 69
T++S++TST T+ + G+ S S + SP + +SQT+ + GGI +GY
Sbjct: 93 TASSSATSTPTSNITGSSSASSIQFAQKSPGSGVIVSQTLSRPSSAGGIPSSGY 146
>SCWB_YEAST (P53189) Probable family 17 glucosidase SCW11 precursor
(EC 3.2.1.-) (Soluble cell wall protein 11)
Length = 542
Score = 30.8 bits (68), Expect = 2.9
Identities = 18/43 (41%), Positives = 26/43 (59%)
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTS 44
T+ +IAS +T TS+STS+ST ++T + S SS TS
Sbjct: 203 TSTSIASQESTESTNTPTSSSTSSSTSSSTSSSTSSSTSSSTS 245
>YLS8_CAEEL (P34393) Hypothetical protein F09G8.8 in chromosome III
precursor
Length = 639
Score = 30.4 bits (67), Expect = 3.8
Identities = 13/30 (43%), Positives = 19/30 (63%)
Query: 16 STTTSTSTSTSTYTTTVAGALSKPSSLTSP 45
STTT+ + +T+T TTV G KP++ P
Sbjct: 234 STTTTRAPTTTTTVTTVKGVTGKPATTAKP 263
>MYSJ_DICDI (P54697) Myosin IJ heavy chain
Length = 2245
Score = 30.4 bits (67), Expect = 3.8
Identities = 21/73 (28%), Positives = 37/73 (49%), Gaps = 9/73 (12%)
Query: 118 QQMIDCYVKTLAQVLGSEEEAKKKIYNVSCE-----RYFGFGCELDEETSNKLEGIPGVL 172
Q+ + +K ++ LGSEEEAKK+I + E +L E+++ K++ + G L
Sbjct: 1213 QEQSEQLIKLSSEKLGSEEEAKKQINQLELELTDHKSKLQIQLQLTEQSNEKIKKLKGKL 1272
Query: 173 FVLPDSYVDPEHQ 185
+ Y D + Q
Sbjct: 1273 ----EEYQDEKKQ 1281
>ACM1_DROME (P16395) Muscarinic acetylcholine receptor DM1
Length = 805
Score = 30.4 bits (67), Expect = 3.8
Identities = 16/30 (53%), Positives = 20/30 (66%)
Query: 10 FKRRLFSTTTSTSTSTSTYTTTVAGALSKP 39
FK STTT+T+T+TST TTT + A P
Sbjct: 22 FKTVTTSTTTTTTTTTSTTTTTASPAGYSP 51
>P4H1_MOUSE (Q60715) Prolyl 4-hydroxylase alpha-1 subunit precursor
(EC 1.14.11.2) (4-PH alpha-1)
(Procollagen-proline,2-oxoglutarate-4-dioxygenase
alpha-1 subunit)
Length = 534
Score = 30.0 bits (66), Expect = 4.9
Identities = 21/70 (30%), Positives = 36/70 (51%), Gaps = 14/70 (20%)
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVD--PEHQDYGAELFVNGEIVQRSPERQRRVEPQ 211
G + D++T+ K +GI VD PE Q Y E+ GE ++ +P RQ+R+ +
Sbjct: 264 GDQSDQKTAPKKKGIA----------VDYLPERQKY--EMLCRGEGIKMTPRRQKRLFCR 311
Query: 212 AQRGDSRPRY 221
G+ P++
Sbjct: 312 YHDGNRNPKF 321
>O5I1_HUMAN (Q13606) Olfactory receptor 5I1 (Olfactory receptor-like
protein OLF1)
Length = 314
Score = 30.0 bits (66), Expect = 4.9
Identities = 20/50 (40%), Positives = 27/50 (54%), Gaps = 2/50 (4%)
Query: 9 SFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRF--IIPMSQTIF 56
S +++ FST S TS + Y T+ S+PS L SP II + TIF
Sbjct: 234 SGRKKTFSTCASHLTSVTIYQGTLLFIYSRPSYLYSPNTDKIISVFYTIF 283
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,154,774
Number of Sequences: 164201
Number of extensions: 1320201
Number of successful extensions: 5348
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 4944
Number of HSP's gapped (non-prelim): 230
length of query: 235
length of database: 59,974,054
effective HSP length: 107
effective length of query: 128
effective length of database: 42,404,547
effective search space: 5427782016
effective search space used: 5427782016
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC145202.10


