

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.28 - phase: 0
(1155 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
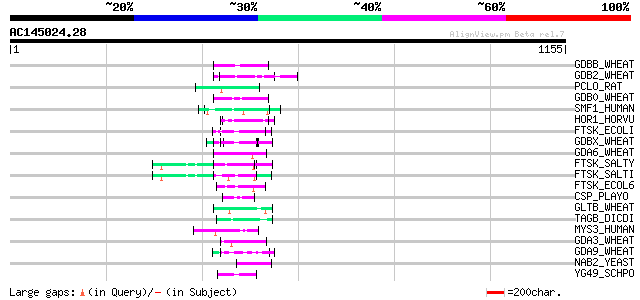
Score E
Sequences producing significant alignments: (bits) Value
GDBB_WHEAT (P06659) Gamma-gliadin B precursor 65 1e-09
GDB2_WHEAT (P08453) Gamma-gliadin precursor 59 6e-08
PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic cytom... 56 5e-07
GDB0_WHEAT (P08079) Gamma-gliadin precursor (Fragment) 56 5e-07
SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated, actin-de... 55 8e-07
HOR1_HORVU (P06470) B1-hordein precursor 54 2e-06
FTSK_ECOLI (P46889) DNA translocase ftsK 54 3e-06
GDBX_WHEAT (P21292) Gamma-gliadin precursor 52 1e-05
GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor (P... 52 1e-05
FTSK_SALTY (Q8ZQD5) DNA translocase ftsK 52 1e-05
FTSK_SALTI (Q8Z814) DNA translocase ftsK 51 2e-05
FTSK_ECOL6 (Q8FJC7) DNA translocase ftsK 50 3e-05
CSP_PLAYO (P06914) Circumsporozoite protein precursor (CS) 50 3e-05
GLTB_WHEAT (P10386) Glutenin, low molecular weight subunit 1D1 p... 50 4e-05
TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC... 49 8e-05
MYS3_HUMAN (Q92794) MYST histone acetyltransferase 3 (Runt-relat... 49 8e-05
GDA3_WHEAT (P04723) Alpha/beta-gliadin A-III precursor (Prolamin) 49 8e-05
GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin) 49 1e-04
NAB2_YEAST (P32505) Nuclear polyadenylated RNA-binding protein NAB2 48 1e-04
YG49_SCHPO (O60184) Protein C23E6.09 in chromosome II 48 2e-04
>GDBB_WHEAT (P06659) Gamma-gliadin B precursor
Length = 291
Score = 65.1 bits (157), Expect = 1e-09
Identities = 41/118 (34%), Positives = 52/118 (43%), Gaps = 13/118 (11%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P +S P PQ T P P Q P+PQ P + QP P PQQ
Sbjct: 40 PHQPFSQQPQQIFPQPQQTFPHQPQQQFPQPQQPQQQFLQPRQPF---------PQQPQQ 90
Query: 485 PYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLP----MTYGALLPSLLAQNLVQTIPP 538
PY Q+P P Q +PQ P+ Q Q QQ P P ++ PSL+ Q+L Q + P
Sbjct: 91 PYPQQPQQPFPQTQQPQQPFPQSKQPQQPFPQPQQPQQSFPQQQPSLIQQSLQQQLNP 148
Score = 40.4 bits (93), Expect = 0.028
Identities = 39/122 (31%), Positives = 47/122 (37%), Gaps = 23/122 (18%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQ------Q 478
PQP F + P+ PQ P Q P PQ P +P QP Q Q Q
Sbjct: 68 PQPQQPQQQF---LQPRQPFPQQP--QQPYPQQPQQPFPQTQQPQQPFPQSKQPQQPFPQ 122
Query: 479 PYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLP-MTYGALLPSLLAQNLVQTIP 537
P PQQ + PQQQP QQ+ +QQ +P P L +L I
Sbjct: 123 PQQPQQSF-------PQQQP----SLIQQSLQQQLNPCKNFLLQQCKPVSLVSSLWSIIL 171
Query: 538 PP 539
PP
Sbjct: 172 PP 173
Score = 32.3 bits (72), Expect = 7.6
Identities = 31/99 (31%), Positives = 36/99 (36%), Gaps = 7/99 (7%)
Query: 453 PRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQ 512
P Q P P QP Q QQ + QP Q P+ P QQ P Q QQ Q
Sbjct: 25 PSGQVQWPQQQPFLQPHQPFSQQPQQIF--PQPQQTFPHQPQQQFP--QPQQPQQQFLQP 80
Query: 513 FDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRP 551
P P P Q QT P + P P+ +P
Sbjct: 81 RQPFPQQPQQPYPQQPQQPFPQTQQPQQ---PFPQSKQP 116
>GDB2_WHEAT (P08453) Gamma-gliadin precursor
Length = 327
Score = 59.3 bits (142), Expect = 6e-08
Identities = 44/134 (32%), Positives = 55/134 (40%), Gaps = 24/134 (17%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNP--NYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPY-- 480
PQ + + P PQ PQ P Q P PQ P +P QP Q QQP+
Sbjct: 55 PQQTFPHQP--QQQVPQPQQPQQPFLQPQQPFPQQPQQPFPQTQQPQQPFPQQPQQPFPQ 112
Query: 481 --YPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPP 538
PQQP+ Q+P P Q +PQ P+ Q Q QQ P P Q +P
Sbjct: 113 TQQPQQPFPQQPQQPFPQTQQPQQPFPQLQQPQQPFPQPQ---------------QQLPQ 157
Query: 539 PRIP-DPLPRWYRP 551
P+ P P+ RP
Sbjct: 158 PQQPQQSFPQQQRP 171
Score = 54.3 bits (129), Expect = 2e-06
Identities = 47/165 (28%), Positives = 67/165 (40%), Gaps = 21/165 (12%)
Query: 438 ITPQMTAP--QNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYY-PQQPYQQRPYNPP 494
+ PQ+ P Q P P+PQ P+ P P Q QQP+ PQQP+ Q+P P
Sbjct: 36 LVPQLQQPLSQQPQQTFPQPQQTFPHQPQQQVPQ---PQQPQQPFLQPQQPFPQQPQQPF 92
Query: 495 QQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP-DPLPRWYRPDL 553
Q +PQ P+ QQ Q+ P P T P Q Q P + P P P+ +P
Sbjct: 93 PQTQQPQQPFPQQPQQ----PFPQTQQPQQP--FPQQPQQPFPQTQQPQQPFPQLQQPQ- 145
Query: 554 HCIYHQGAPGHDVERCFALKKEVQKLINSKELTFTDPDAVAQNNP 598
P ++ ++ Q+ ++ F P Q NP
Sbjct: 146 -------QPFPQPQQQLPQPQQPQQSFPQQQRPFIQPSLQQQLNP 183
Score = 35.8 bits (81), Expect = 0.69
Identities = 33/82 (40%), Positives = 35/82 (42%), Gaps = 15/82 (18%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQ--GPAPYYPPLYQPLYNLQQLSQQPYYP 482
PQ PF PQ PQ Q P PQ P +P QP QQ QP P
Sbjct: 111 PQTQQPQQPFPQQ--PQQPFPQTQQPQQPFPQLQQPQQPFP---QP----QQQLPQPQQP 161
Query: 483 QQ--PYQQRPYNPP--QQQPRP 500
QQ P QQRP+ P QQQ P
Sbjct: 162 QQSFPQQQRPFIQPSLQQQLNP 183
>PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic
cytomatrix protein)
Length = 5085
Score = 56.2 bits (134), Expect = 5e-07
Identities = 44/141 (31%), Positives = 49/141 (34%), Gaps = 8/141 (5%)
Query: 387 SGSSSGGSSGVSSSGNKKFGNGYPKRNAQEVGMVAHGGPQPVYSNHPFVANIT------- 439
S G S + G+ KF G K AQ+ G H QP V T
Sbjct: 321 SSQQPGPKSLAQTPGHGKFPLGPVKSPAQQPGTAKHPAQQPGPQTAAKVPGPTKTPAQQS 380
Query: 440 -PQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQP 498
P T Q P P PQ P P P QP+ Q Q QP Q P P QQP
Sbjct: 381 GPGKTPAQQPGPTKPSPQQPIPAKPQPQQPVATKTQPQQSAPAKPQPQQPAPAKPQPQQP 440
Query: 499 RPQAPYNQQNQKQQFDPLPMT 519
P P Q + P P T
Sbjct: 441 TPAKPQPQPPTPAKPQPQPPT 461
>GDB0_WHEAT (P08079) Gamma-gliadin precursor (Fragment)
Length = 251
Score = 56.2 bits (134), Expect = 5e-07
Identities = 41/120 (34%), Positives = 51/120 (42%), Gaps = 10/120 (8%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
P +S P PQ T P P Q P+PQ P + QP Q QQPY PQQ
Sbjct: 40 PHQPFSQQPQQTFPQPQQTFPHQPQQQFPQPQQPQQQF---LQPQQPFPQQPQQPY-PQQ 95
Query: 485 PYQQRPYNPPQQQ--PRPQAPYNQQNQKQQFDPLP----MTYGALLPSLLAQNLVQTIPP 538
P Q P QQ P+ Q P Q +Q QQ P P ++ P + +L Q + P
Sbjct: 96 PQQPFPQTQQPQQLFPQSQQPQQQFSQPQQQFPQPQQPQQSFPQQQPPFIQPSLQQQVNP 155
Score = 41.2 bits (95), Expect = 0.016
Identities = 31/87 (35%), Positives = 39/87 (44%), Gaps = 7/87 (8%)
Query: 439 TPQMTAPQNPNYQSPRPQG-PAPYYPPLYQPLYNLQQLSQQ-PYYPQQPYQQRPYNPPQQ 496
T M + Q P+ Q P P+ P QP Q Q P+ PQQ + Q P P QQ
Sbjct: 18 TANMQVDPSSQVQWPQQQPVPQPHQPFSQQPQQTFPQPQQTFPHQPQQQFPQ-PQQPQQQ 76
Query: 497 QPRPQAPYNQQNQ----KQQFDPLPMT 519
+PQ P+ QQ Q +Q P P T
Sbjct: 77 FLQPQQPFPQQPQQPYPQQPQQPFPQT 103
Score = 32.7 bits (73), Expect = 5.8
Identities = 38/150 (25%), Positives = 52/150 (34%), Gaps = 23/150 (15%)
Query: 453 PRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQ 512
P Q P P+ QP Q QQ + QP Q P+ P QQ P Q QQ Q
Sbjct: 25 PSSQVQWPQQQPVPQPHQPFSQQPQQTF--PQPQQTFPHQPQQQFP--QPQQPQQQFLQP 80
Query: 513 FDPLPMTYGALLPSLLAQNLVQTIPP----PRIPDPLPRWYRPDLHCIYHQGAPGHDVER 568
P P P Q QT P P+ P ++ +P ++
Sbjct: 81 QQPFPQQPQQPYPQQPQQPFPQTQQPQQLFPQSQQPQQQFSQP---------------QQ 125
Query: 569 CFALKKEVQKLINSKELTFTDPDAVAQNNP 598
F ++ Q+ ++ F P Q NP
Sbjct: 126 QFPQPQQPQQSFPQQQPPFIQPSLQQQVNP 155
>SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated,
actin-dependent regulator of chromatin subfamily F
member 1 (SWI-SNF complex protein p270) (B120)
Length = 1902
Score = 55.5 bits (132), Expect = 8e-07
Identities = 51/156 (32%), Positives = 62/156 (39%), Gaps = 20/156 (12%)
Query: 393 GSSGVSSSGNKKFGNGYPKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQS 452
G SG G + N Q+ QP YS P Q + Q P Q
Sbjct: 75 GPSGYGQQGQTPYYNQQSPHPQQQ---------QPPYSQQPPSQTPHAQPSYQQQPQSQP 125
Query: 453 PRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPY-QQRPYN-PPQQQPRPQAPYNQQNQK 510
P+ Q P Y QP Q S PY QQ QQ P + PP QP+ Q+PY QQ Q
Sbjct: 126 PQLQSSQPPYSQ--QPSQPPHQQSPAPYPSQQSTTQQHPQSQPPYSQPQAQSPY-QQQQP 182
Query: 511 QQFDPLPMTYGALLPSLLAQNLVQT------IPPPR 540
QQ P ++ A P +Q QT PPP+
Sbjct: 183 QQPAPSTLSQQAAYPQPQSQQSQQTAYSQQRFPPPQ 218
Score = 45.8 bits (107), Expect = 7e-04
Identities = 47/175 (26%), Positives = 58/175 (32%), Gaps = 41/175 (23%)
Query: 406 GNGYP-----KRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAP 460
G+GYP + Q M G Q + I P QGP+
Sbjct: 31 GHGYPGQPYGSQTPQRYPMTVQGRAQSAMGGLSYTQQIPPY------------GQQGPSG 78
Query: 461 YYPPLYQPLYNLQQ---LSQQPYYPQQP----------YQQRPYNPPQQQPRPQAPYNQQ 507
Y P YN Q QQP Y QQP YQQ+P + P Q Q PY+QQ
Sbjct: 79 YGQQGQTPYYNQQSPHPQQQQPPYSQQPPSQTPHAQPSYQQQPQSQPPQLQSSQPPYSQQ 138
Query: 508 NQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRPDLHCIYHQGAP 562
+ P Y PS Q + P P + +P Y Q P
Sbjct: 139 PSQPPHQQSPAPY----PS-------QQSTTQQHPQSQPPYSQPQAQSPYQQQQP 182
>HOR1_HORVU (P06470) B1-hordein precursor
Length = 293
Score = 54.3 bits (129), Expect = 2e-06
Identities = 42/109 (38%), Positives = 49/109 (44%), Gaps = 7/109 (6%)
Query: 444 APQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAP 503
A Q P Q P PQ P PY P QP Y Q Q +PQQP Q+P PQQ PQ P
Sbjct: 19 AQQQPFPQQPIPQQPQPY-PQQPQP-YPQQPFPPQQPFPQQPVPQQPQPYPQQPFPPQQP 76
Query: 504 YNQQNQKQQFDPLPM--TYGALLPSLLAQNLVQTIPPPRIPDPLPRWYR 550
+ QQ Q P P +G P L Q Q P + P P + Y+
Sbjct: 77 FPQQPPFWQQKPFPQQPPFGLQQPILSQQ---QPCTPQQTPLPQGQLYQ 122
Score = 48.1 bits (113), Expect = 1e-04
Identities = 45/116 (38%), Positives = 57/116 (48%), Gaps = 23/116 (19%)
Query: 440 PQMTAPQNPNYQSPRPQGPAPY----YPPLYQPLYNLQQLSQQPY-YPQQPY---QQRPY 491
PQ PQ P P PQ P PY +PP Q + Q + QQP YPQQP+ Q P
Sbjct: 25 PQQPIPQQPQ---PYPQQPQPYPQQPFPP--QQPFPQQPVPQQPQPYPQQPFPPQQPFPQ 79
Query: 492 NPP--QQQPRP-QAPYNQQ----NQKQQFDP--LPMTYGALLPSLLAQNLVQTIPP 538
PP QQ+P P Q P+ Q +Q+Q P P+ G L +LL Q +Q + P
Sbjct: 80 QPPFWQQKPFPQQPPFGLQQPILSQQQPCTPQQTPLPQGQLYQTLL-QLQIQYVHP 134
Score = 36.2 bits (82), Expect = 0.53
Identities = 20/44 (45%), Positives = 26/44 (58%), Gaps = 2/44 (4%)
Query: 475 LSQQPYYPQQPYQQRPYNPPQQ-QPRPQAPYNQQNQKQQFDPLP 517
++QQ +PQQP Q+P PQQ QP PQ P+ Q Q P+P
Sbjct: 18 IAQQQPFPQQPIPQQPQPYPQQPQPYPQQPFPPQQPFPQ-QPVP 60
>FTSK_ECOLI (P46889) DNA translocase ftsK
Length = 1329
Score = 53.5 bits (127), Expect = 3e-06
Identities = 40/113 (35%), Positives = 47/113 (41%), Gaps = 18/113 (15%)
Query: 440 PQMTAPQNPNYQSPR-PQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQ--RPYNPPQQ 496
PQ P P YQ P+ P P P Y Q QQP PQQ YQQ +P P QQ
Sbjct: 755 PQQPVPPQPQYQQPQQPVAPQPQY-----------QQPQQPVAPQQQYQQPQQPVAPQQQ 803
Query: 497 QPRPQAPYNQQNQKQQFDPLPMTYG----ALLPSLLAQNLVQTIPPPRIPDPL 545
+PQ P Q Q PL M G P+ +L PPP +P+
Sbjct: 804 YQQPQQPVAPQPQDTLLHPLLMRNGDSRPLHKPTTPLPSLDLLTPPPSEVEPV 856
Score = 47.8 bits (112), Expect = 2e-04
Identities = 41/114 (35%), Positives = 50/114 (42%), Gaps = 17/114 (14%)
Query: 423 GGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYP 482
G +P+++ P V + Q P P Q +PQ P P P QP QQP P
Sbjct: 726 GPHEPLFT--PIVEPVQ-QPQQPVAPQQQYQQPQQPVPPQPQYQQP--------QQPVAP 774
Query: 483 QQPYQQ--RPYNPPQQQPRPQ---APYNQQNQKQQFDPLPMTYGALLPSLLAQN 531
Q YQQ +P P QQ +PQ AP Q Q QQ P LL LL +N
Sbjct: 775 QPQYQQPQQPVAPQQQYQQPQQPVAPQQQYQQPQQ-PVAPQPQDTLLHPLLMRN 827
Score = 38.1 bits (87), Expect = 0.14
Identities = 33/144 (22%), Positives = 52/144 (35%), Gaps = 32/144 (22%)
Query: 49 VVTAAPEGPPQPSPTTTTTIGLTQPLITDFSSGNMATMGNSGFRPLGPQGPFATPQYSMP 108
V+ APEG PQ S + +PL +P+ PQ P+ P P
Sbjct: 372 VIAPAPEGYPQQSQYAQPAVQYNEPL----------------QQPVQPQQPYYAPAAEQP 415
Query: 109 SGYPW-----GMPIATN--------GVFGPGATEMPFAQGQQTSAH---FQMSQPIPQAT 152
+ P+ P+A N F P +T QQ +A +Q QP+ Q
Sbjct: 416 AQQPYYAPAPEQPVAGNAWQAEEQQSTFAPQSTYQTEQTYQQPAAQEPLYQQPQPVEQQP 475
Query: 153 MTQAGPTVHVGPQHEEQIYHSDSI 176
+ + P V +Y+ + +
Sbjct: 476 VVEPEPVVEETKPARPPLYYFEEV 499
Score = 35.0 bits (79), Expect = 1.2
Identities = 23/73 (31%), Positives = 30/73 (40%), Gaps = 10/73 (13%)
Query: 452 SPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPY---------QQRPYNPPQQQPRPQA 502
+P P+G P QP + QQP PQQPY QQ Y P +QP
Sbjct: 374 APAPEG-YPQQSQYAQPAVQYNEPLQQPVQPQQPYYAPAAEQPAQQPYYAPAPEQPVAGN 432
Query: 503 PYNQQNQKQQFDP 515
+ + Q+ F P
Sbjct: 433 AWQAEEQQSTFAP 445
>GDBX_WHEAT (P21292) Gamma-gliadin precursor
Length = 302
Score = 51.6 bits (122), Expect = 1e-05
Identities = 33/108 (30%), Positives = 42/108 (38%), Gaps = 17/108 (15%)
Query: 410 PKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPL 469
P+R + H PQ + PQ T P P Q P+ Q P +P Q
Sbjct: 48 PQRTIPQPHQTFHHQPQQTFPQ--------PQQTYPHQPQQQFPQTQQPQQPFPQPQQTF 99
Query: 470 YNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLP 517
P PQ P+ Q+P P Q +PQ P+ Q Q QQ P P
Sbjct: 100 ---------PQQPQLPFPQQPQQPFPQPQQPQQPFPQSQQPQQPFPQP 138
Score = 49.7 bits (117), Expect = 5e-05
Identities = 36/89 (40%), Positives = 42/89 (46%), Gaps = 15/89 (16%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
PQ PF PQ T PQ P Q P PQ P +P QP QQP +PQ
Sbjct: 83 PQTQQPQQPFPQ---PQQTFPQQP--QLPFPQQPQQPFPQPQQP--------QQP-FPQS 128
Query: 485 PYQQRPYNPPQQQ-PRPQAPYNQQNQKQQ 512
Q+P+ PQQQ P+PQ P Q+QQ
Sbjct: 129 QQPQQPFPQPQQQFPQPQQPQQSFPQQQQ 157
Score = 47.0 bits (110), Expect = 3e-04
Identities = 36/109 (33%), Positives = 45/109 (41%), Gaps = 8/109 (7%)
Query: 445 PQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQP--RPQA 502
PQ P Q P+ P P+ +QP Q Q YP QP QQ P QQP +PQ
Sbjct: 40 PQQPFCQQPQRTIPQPHQTFHHQPQQTFPQPQQT--YPHQPQQQFPQTQQPQQPFPQPQQ 97
Query: 503 PYNQQNQ----KQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPR 547
+ QQ Q +Q P P P +Q Q P P+ P P+
Sbjct: 98 TFPQQPQLPFPQQPQQPFPQPQQPQQPFPQSQQPQQPFPQPQQQFPQPQ 146
Score = 46.6 bits (109), Expect = 4e-04
Identities = 29/77 (37%), Positives = 37/77 (47%), Gaps = 10/77 (12%)
Query: 440 PQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPY-YPQQPYQQRPYNPPQQQP 498
PQ+ PQ P P+PQ P +P QP QQP+ PQQ + Q P P Q P
Sbjct: 103 PQLPFPQQPQQPFPQPQQPQQPFPQSQQP--------QQPFPQPQQQFPQ-PQQPQQSFP 153
Query: 499 RPQAPYNQQNQKQQFDP 515
+ Q P Q +QQ +P
Sbjct: 154 QQQQPAIQSFLQQQMNP 170
Score = 43.1 bits (100), Expect = 0.004
Identities = 40/137 (29%), Positives = 52/137 (37%), Gaps = 12/137 (8%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
PQ + P P T P P+PQ P+ P Q + Q QQP+ Q
Sbjct: 40 PQQPFCQQPQRTIPQPHQTFHHQPQQTFPQPQQTYPHQP---QQQFPQTQQPQQPF--PQ 94
Query: 485 PYQQRPYNPPQQQP-RPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP- 542
P Q P P P +PQ P+ Q Q QQ P P + P Q Q P P+ P
Sbjct: 95 PQQTFPQQPQLPFPQQPQQPFPQPQQPQQ--PFPQSQQPQQPFPQPQ---QQFPQPQQPQ 149
Query: 543 DPLPRWYRPDLHCIYHQ 559
P+ +P + Q
Sbjct: 150 QSFPQQQQPAIQSFLQQ 166
Score = 34.7 bits (78), Expect = 1.5
Identities = 35/116 (30%), Positives = 46/116 (39%), Gaps = 5/116 (4%)
Query: 439 TPQMTAPQNPNYQSPRPQG-PAPYYPPLYQPLYNLQQLSQQPYY-PQQPYQQRPYNPPQQ 496
T M + Q P+ Q P P P QP + Q Q ++ PQQ + Q P
Sbjct: 18 TANMQVDPSGQVQWPQQQPFPQPQQPFCQQPQRTIPQPHQTFHHQPQQTFPQ-PQQTYPH 76
Query: 497 QPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP-DPLPRWYRP 551
QP+ Q P QQ Q Q F T+ Q Q P P+ P P P+ +P
Sbjct: 77 QPQQQFPQTQQPQ-QPFPQPQQTFPQQPQLPFPQQPQQPFPQPQQPQQPFPQSQQP 131
>GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor
(Prolamin)
Length = 296
Score = 51.6 bits (122), Expect = 1e-05
Identities = 42/126 (33%), Positives = 51/126 (40%), Gaps = 17/126 (13%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQS---PRPQGPAPYYPPLYQPLYNLQQLSQ-QPY 480
PQP + P P + Q P Q P+ P P P QP LQ Q QP+
Sbjct: 28 PQPQNPSQPQPQGQVPLVQQQQFPGQQQQFPPQQPYPQPQPFPSQQPYLQLQPFPQPQPF 87
Query: 481 YPQQPYQQRPYNPPQQ---QPRPQAPY----------NQQNQKQQFDPLPMTYGALLPSL 527
PQ PY Q P PQQ QP+PQ P QQ Q+QQ +L +
Sbjct: 88 PPQLPYPQPPPFSPQQPYPQPQPQYPQPQQPISQQQAQQQQQQQQQQQQQQQQQQILQQI 147
Query: 528 LAQNLV 533
L Q L+
Sbjct: 148 LQQQLI 153
Score = 43.1 bits (100), Expect = 0.004
Identities = 38/117 (32%), Positives = 48/117 (40%), Gaps = 22/117 (18%)
Query: 409 YPKRNAQEVGMVAHGGPQPVYSNHPFVA--------NITPQMTAPQNPNY--QSPRPQGP 458
+P + Q + PQP S P++ PQ+ PQ P + Q P PQ P
Sbjct: 50 FPGQQQQFPPQQPYPQPQPFPSQQPYLQLQPFPQPQPFPPQLPYPQPPPFSPQQPYPQ-P 108
Query: 459 APYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDP 515
P YP QP QQP QQ QQ+ QQQ + Q QQ +QQ P
Sbjct: 109 QPQYP---QP--------QQPISQQQAQQQQQQQQQQQQQQQQQQILQQILQQQLIP 154
Score = 40.8 bits (94), Expect = 0.021
Identities = 28/75 (37%), Positives = 36/75 (47%), Gaps = 10/75 (13%)
Query: 487 QQRPYNPPQQQPRPQAPYNQQ----NQKQQFD-----PLPMTYGALLPSLLAQNLVQTIP 537
Q +P NP Q QP+ Q P QQ Q+QQF P P + + P L Q Q P
Sbjct: 27 QPQPQNPSQPQPQGQVPLVQQQQFPGQQQQFPPQQPYPQPQPFPSQQPYLQLQPFPQPQP 86
Query: 538 -PPRIPDPLPRWYRP 551
PP++P P P + P
Sbjct: 87 FPPQLPYPQPPPFSP 101
>FTSK_SALTY (Q8ZQD5) DNA translocase ftsK
Length = 1351
Score = 51.6 bits (122), Expect = 1e-05
Identities = 47/134 (35%), Positives = 56/134 (41%), Gaps = 15/134 (11%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQ--LSQQPYY- 481
PQP Y PQ PQ P +P+PQ P P QP Y Q ++ QP Y
Sbjct: 733 PQPQYQQPQQPVAPQPQYQQPQQP--VAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 790
Query: 482 -PQQPYQQRP-YNPPQQQPRPQAPYNQ-------QNQKQQFDPLPMTYGALLPSLLAQNL 532
PQQP +P Y PQQ PQ Y Q Q Q QQ +L+ LL +N
Sbjct: 791 QPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPTAPQDSLIHPLLMRN- 849
Query: 533 VQTIPPPRIPDPLP 546
+ P R PLP
Sbjct: 850 GDSRPLQRPTTPLP 863
Score = 51.6 bits (122), Expect = 1e-05
Identities = 65/227 (28%), Positives = 91/227 (39%), Gaps = 27/227 (11%)
Query: 298 NTYLAPSRKELQSLTQK------DKESFKEYAQRFIQKAAQIRPP----LDERELSDLFY 347
N P+R+EL S K +E +E + + AQ+ + + EL+ F
Sbjct: 580 NRVRVPTRRELASYGIKLPSQRIAEEKAREAERNQYETGAQLTDEEIDAMHQDELARQFA 639
Query: 348 ETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSSGNKKFGN 407
++ Y E Q D +T E AR+ AA S SG +G + F
Sbjct: 640 QSQQHRYGETYQHDTQQAEDD--DTAAEAE-LARQFAA---SQQQRYSGEQPAGAQPFSL 693
Query: 408 GYPKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQ 467
+ +V +V G +P+ F + P+ T Q P P+PQ P P Q
Sbjct: 694 DDLDFSPMKV-LVDEGPHEPL-----FTPGVMPESTPVQQPVAPQPQPQYQQPQQPVAPQ 747
Query: 468 PLYNLQQLSQQPYYPQQPYQ--QRPYNPPQQQPRPQAPYNQQNQKQQ 512
P Y Q QQP PQ YQ Q+P P Q +PQ P Q Q QQ
Sbjct: 748 PQY---QQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQ 791
Score = 48.1 bits (113), Expect = 1e-04
Identities = 33/90 (36%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQ--LSQQPYY- 481
P+ P PQ PQ P +P+PQ P P QP Y Q ++ QP Y
Sbjct: 720 PESTPVQQPVAPQPQPQYQQPQQP--VAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 777
Query: 482 -PQQPYQQRP-YNPPQQQPRPQAPYNQQNQ 509
PQQP +P Y PQQ PQ Y Q Q
Sbjct: 778 QPQQPVAPQPQYQQPQQPVAPQPQYQQPQQ 807
>FTSK_SALTI (Q8Z814) DNA translocase ftsK
Length = 1343
Score = 51.2 bits (121), Expect = 2e-05
Identities = 45/135 (33%), Positives = 54/135 (39%), Gaps = 16/135 (11%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQ--LSQQPYY- 481
PQP Y PQ PQ P +P+PQ P P QP Y Q ++ QP Y
Sbjct: 738 PQPQYQQPQQPVASQPQYQQPQQP--VAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQ 795
Query: 482 -PQQPYQQRP-YNPPQQQPRPQAPYNQQN-----QKQQFDPLPMTYG----ALLPSLLAQ 530
PQQP +P Y PQQ PQ Y Q Q PL M G P+
Sbjct: 796 QPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPTAPQDSLIHPLLMRNGDSRPLQRPTTPLP 855
Query: 531 NLVQTIPPPRIPDPL 545
+L PPP +P+
Sbjct: 856 SLDLLTPPPSEVEPV 870
Score = 49.3 bits (116), Expect = 6e-05
Identities = 64/230 (27%), Positives = 92/230 (39%), Gaps = 25/230 (10%)
Query: 298 NTYLAPSRKELQSLTQK------DKESFKEYAQRFIQKAAQIRPP----LDERELSDLFY 347
N P+R+EL S K +E +E + + AQ+ + + EL+ F
Sbjct: 577 NRVRVPTRRELASYGIKLPSQRIAEEKAREAERNQYETGAQLTDEEIDAMHQDELARQFA 636
Query: 348 ETLSPCYSEKMIVCASQKFTDLVETGMRIEEWARKGAAVSGSSSGGSSGVSSSGNKKFGN 407
++ Y E Q D +T E AR+ AA S SG +G + F
Sbjct: 637 QSQQHRYGETYQHDTQQAEDD--DTAAEAE-LARQFAA---SQQQRYSGEQPAGAQPFSL 690
Query: 408 GYPKRNAQEVGMVAHGGPQPVYSNHPFVANITPQMTAPQNPNYQ---SPRPQGPAPYYPP 464
+ +V +V G +P+++ + Q P YQ +P+PQ P P
Sbjct: 691 DDLDFSPMKV-LVDEGPHEPLFTPGVMPESTPVQQPVAPQPQYQQPVAPQPQYQQPQQPV 749
Query: 465 LYQPLYNLQQLSQQPYYPQQPYQ--QRPYNPPQQQPRPQAPYNQQNQKQQ 512
QP Y Q QQP PQ YQ Q+P P Q +PQ P Q Q QQ
Sbjct: 750 ASQPQY---QQPQQPVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQ 796
Score = 47.0 bits (110), Expect = 3e-04
Identities = 38/96 (39%), Positives = 41/96 (42%), Gaps = 22/96 (22%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPR------PQGPAPYYPPLYQPLYNLQQLSQQ 478
PQP Y Q APQ P YQ P+ PQ P P QP Y Q QQ
Sbjct: 728 PQPQYQ----------QPVAPQ-PQYQQPQQPVASQPQYQQPQQPVAPQPQY---QQPQQ 773
Query: 479 PYYPQQPYQ--QRPYNPPQQQPRPQAPYNQQNQKQQ 512
P PQ YQ Q+P P Q +PQ P Q Q QQ
Sbjct: 774 PVAPQPQYQQPQQPVAPQPQYQQPQQPVAPQPQYQQ 809
>FTSK_ECOL6 (Q8FJC7) DNA translocase ftsK
Length = 1347
Score = 50.4 bits (119), Expect = 3e-05
Identities = 38/110 (34%), Positives = 45/110 (40%), Gaps = 14/110 (12%)
Query: 431 NHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRP 490
+ P I + PQ P +P+ Q P P QP Y Q QQP PQ YQQ
Sbjct: 741 HEPLFTPIVEPVQQPQQP--VAPQQQYQQPQQPVAPQPQY---QQPQQPVAPQPQYQQPQ 795
Query: 491 YNPPQQQPRPQAPYNQ---------QNQKQQFDPLPMTYGALLPSLLAQN 531
Y PQQ PQ Y Q Q Q+ Q +P LL LL +N
Sbjct: 796 YQQPQQPVAPQQQYQQPQQPVTQQPQYQQPQQPVVPQPQDTLLHPLLMRN 845
Score = 42.4 bits (98), Expect = 0.007
Identities = 41/126 (32%), Positives = 49/126 (38%), Gaps = 17/126 (13%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQ 484
PQ Y PQ PQ P +P+PQ P Y QP+ QQ Q PQQ
Sbjct: 761 PQQQYQQPQQPVAPQPQYQQPQQP--VAPQPQYQQPQYQQPQQPVAPQQQYQQ----PQQ 814
Query: 485 PYQQRP-YNPPQQQPRPQAPYNQQNQKQQFDPLPMTYG----ALLPSLLAQNLVQTIPPP 539
P Q+P Y PQQ P Q Q PL M G P+ +L PPP
Sbjct: 815 PVTQQPQYQQPQQ------PVVPQPQDTLLHPLLMRNGDSRPLHKPTTPLPSLDLLTPPP 868
Query: 540 RIPDPL 545
+P+
Sbjct: 869 SEVEPV 874
Score = 40.0 bits (92), Expect = 0.037
Identities = 28/83 (33%), Positives = 36/83 (42%), Gaps = 11/83 (13%)
Query: 444 APQNPNYQSPRPQGPAPYYPPLYQPLYNLQ--------QLSQQPYYP---QQPYQQRPYN 492
AP+ +QS Q Y PL QP+ Q Q QQPYY +QP QQ Y
Sbjct: 376 APEGYPHQSQYAQPAVQYNEPLQQPVQPQQPYYAPAAEQPVQQPYYAPAAEQPVQQPYYA 435
Query: 493 PPQQQPRPQAPYNQQNQKQQFDP 515
P +QP + + Q+ F P
Sbjct: 436 PAPEQPVAGNAWQAEEQQSTFAP 458
Score = 37.7 bits (86), Expect = 0.18
Identities = 22/66 (33%), Positives = 28/66 (42%), Gaps = 1/66 (1%)
Query: 452 SPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQ 511
+P P+G P+ QP + QQP PQQPY P QQP Q Q+
Sbjct: 374 APAPEG-YPHQSQYAQPAVQYNEPLQQPVQPQQPYYAPAAEQPVQQPYYAPAAEQPVQQP 432
Query: 512 QFDPLP 517
+ P P
Sbjct: 433 YYAPAP 438
>CSP_PLAYO (P06914) Circumsporozoite protein precursor (CS)
Length = 367
Score = 50.4 bits (119), Expect = 3e-05
Identities = 37/68 (54%), Positives = 37/68 (54%), Gaps = 10/68 (14%)
Query: 444 APQNPNY-QSP-RPQGP-APYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRP 500
APQ P Q P PQGP AP PP P QQ QQP PQQP QQ P PP QQPRP
Sbjct: 209 APQGPGAPQGPGAPQGPGAPQEPPQQPP----QQPPQQP--PQQPPQQPPQQPP-QQPRP 261
Query: 501 QAPYNQQN 508
Q N N
Sbjct: 262 QPDGNNNN 269
Score = 40.4 bits (93), Expect = 0.028
Identities = 33/77 (42%), Positives = 37/77 (47%), Gaps = 9/77 (11%)
Query: 444 APQNPNY-QSP-RPQGP-APYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPR- 499
APQ P Q P PQGP AP P Q Q P PQ+P QQ P PPQQ P+
Sbjct: 191 APQGPGAPQGPGAPQGPGAPQGPGAPQG----PGAPQGPGAPQEPPQQPPQQPPQQPPQQ 246
Query: 500 -PQAPYNQQNQKQQFDP 515
PQ P Q Q+ + P
Sbjct: 247 PPQQPPQQPPQQPRPQP 263
>GLTB_WHEAT (P10386) Glutenin, low molecular weight subunit 1D1
precursor
Length = 307
Score = 50.1 bits (118), Expect = 4e-05
Identities = 41/136 (30%), Positives = 54/136 (39%), Gaps = 19/136 (13%)
Query: 425 PQPVYSNHPFVANITPQMTAPQNPNYQSPRP-----QGPAPYYPPLY--QPLYNLQQLSQ 477
PQ + P + Q PQ P++ +P Q P P+ QP ++ QQ
Sbjct: 40 PQQTFPQQPLFSQQQQQQLFPQQPSFSQQQPPFWQQQPPFSQQQPILPQQPPFSQQQQLV 99
Query: 478 QPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQ------N 531
P P QQ+P PPQQ P PQ Q QQ P+ + PS+L Q
Sbjct: 100 LPQQPPFSQQQQPVLPPQQSPFPQQQQQHQQLVQQQIPV------VQPSILQQLNPCKVF 153
Query: 532 LVQTIPPPRIPDPLPR 547
L Q P +P L R
Sbjct: 154 LQQQCSPVAMPQRLAR 169
Score = 37.4 bits (85), Expect = 0.24
Identities = 30/80 (37%), Positives = 38/80 (47%), Gaps = 13/80 (16%)
Query: 441 QMTAPQNPNYQSPRPQGPAPYYPPL-YQPLYNLQQLSQQPYYPQQP---YQQRPY---NP 493
QM P + P Q P P QPL++ QQ QQ +PQQP QQ P+ P
Sbjct: 20 QMETRCIPGLERPWQQQPLPPQQTFPQQPLFSQQQ--QQQLFPQQPSFSQQQPPFWQQQP 77
Query: 494 PQQQPRP----QAPYNQQNQ 509
P Q +P Q P++QQ Q
Sbjct: 78 PFSQQQPILPQQPPFSQQQQ 97
Score = 33.5 bits (75), Expect = 3.4
Identities = 20/60 (33%), Positives = 30/60 (49%)
Query: 473 QQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNL 532
QQL Q PQQ QQ+ PQQQ Q + Q +Q Q + + +LP++ + N+
Sbjct: 228 QQLGQCVSQPQQQSQQQLGQQPQQQQLAQGTFLQPHQIAQLEVMTSIALRILPTMCSVNV 287
Score = 33.5 bits (75), Expect = 3.4
Identities = 30/87 (34%), Positives = 38/87 (43%), Gaps = 9/87 (10%)
Query: 467 QPLYNLQQLSQQPYYPQQPYQQR-PYNP--PQQQP---RPQAPYNQQNQ-KQQFDPLPMT 519
QPL Q QQP + QQ QQ P P QQQP + Q P++QQ Q P
Sbjct: 36 QPLPPQQTFPQQPLFSQQQQQQLFPQQPSFSQQQPPFWQQQPPFSQQQPILPQQPPFSQQ 95
Query: 520 YGALLPSL--LAQNLVQTIPPPRIPDP 544
+LP +Q +PP + P P
Sbjct: 96 QQLVLPQQPPFSQQQQPVLPPQQSPFP 122
Score = 32.7 bits (73), Expect = 5.8
Identities = 20/64 (31%), Positives = 29/64 (45%), Gaps = 2/64 (3%)
Query: 483 QQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIP 542
++P+QQ+P P QQ PQ P Q Q+QQ P ++ P Q + P +P
Sbjct: 30 ERPWQQQPL--PPQQTFPQQPLFSQQQQQQLFPQQPSFSQQQPPFWQQQPPFSQQQPILP 87
Query: 543 DPLP 546
P
Sbjct: 88 QQPP 91
>TAGB_DICDI (P54683) Prestalk-specific protein tagB precursor (EC
3.4.21.-)
Length = 1905
Score = 48.9 bits (115), Expect = 8e-05
Identities = 38/119 (31%), Positives = 46/119 (37%), Gaps = 16/119 (13%)
Query: 430 SNHPFVANITP--QMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQ 487
+N P ++ P Q T P + Q P P PP Q QQ QQ QQ Q
Sbjct: 1778 NNEPSSSSTPPNDQPTPPPQEQQEQKNDQPPPP--PPQEQQEQQEQQQQQQQEQQQQQQQ 1835
Query: 488 QRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLP 546
Q+ QQQ + Q QQ Q+QQ D P Y Q PPP +P P
Sbjct: 1836 QQQQQQQQQQQQQQQQQQQQQQQQQNDQPPNDYD------------QVPPPPPLPSESP 1882
Score = 34.3 bits (77), Expect = 2.0
Identities = 21/66 (31%), Positives = 25/66 (37%)
Query: 447 NPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQ 506
N N P P P P +Q + QP P QQ QQQ + Q Q
Sbjct: 1775 NNNNNEPSSSSTPPNDQPTPPPQEQQEQKNDQPPPPPPQEQQEQQEQQQQQQQEQQQQQQ 1834
Query: 507 QNQKQQ 512
Q Q+QQ
Sbjct: 1835 QQQQQQ 1840
>MYS3_HUMAN (Q92794) MYST histone acetyltransferase 3 (Runt-related
transcription factor binding protein 2) (Monocytic
leukemia zinc finger protein) (Zinc finger protein 220)
Length = 2004
Score = 48.9 bits (115), Expect = 8e-05
Identities = 45/149 (30%), Positives = 60/149 (40%), Gaps = 19/149 (12%)
Query: 383 GAAVSGSSSGGSS----GVSSSGNKKFGNGYPKRNAQEVG----MVAHGGPQP-----VY 429
G ++ G+SS SS G+SSS + + + +G M+ QP +
Sbjct: 1574 GGSICGNSSSQSSCSYGGLSSSSSLTQSSCVVTQQMASMGSSCSMMQQSSVQPAANCSIK 1633
Query: 430 SNHPFVANITPQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQLSQQPYYPQQPYQQR 489
S V P Q P +PQ P P P QP QQ QQP QP Q+
Sbjct: 1634 SPQSCVVERPPSNQQQQPPPPPPQQPQPPPPQPQPAPQPPPPQQQPQQQP----QPQPQQ 1689
Query: 490 PYNPPQQQPRPQAPYNQQNQKQQFDPLPM 518
P PP P+ Q P +Q + F P PM
Sbjct: 1690 P--PPPPPPQQQPPLSQCSMNNSFTPAPM 1716
>GDA3_WHEAT (P04723) Alpha/beta-gliadin A-III precursor (Prolamin)
Length = 282
Score = 48.9 bits (115), Expect = 8e-05
Identities = 41/114 (35%), Positives = 49/114 (42%), Gaps = 21/114 (18%)
Query: 440 PQMTAPQNPNYQSPRPQGPA---------------PYYP-PLYQPLYNLQQLSQ-QPYYP 482
PQ+ PQNP+ Q P+ Q P P P P QP + Q Q QP+ P
Sbjct: 26 PQLQ-PQNPSQQQPQEQVPLMQQQQQFPGQQEQFPPQQPYPHQQPFPSQQPYPQPQPFPP 84
Query: 483 QQPYQQRPYNPPQQ---QPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLV 533
Q PY Q PPQQ QP+PQ P QQ QQ L +L Q L+
Sbjct: 85 QLPYPQTQPFPPQQPYPQPQPQYPQPQQPISQQQAQQQQQQQQTLQQILQQQLI 138
Score = 36.2 bits (82), Expect = 0.53
Identities = 26/71 (36%), Positives = 34/71 (47%), Gaps = 7/71 (9%)
Query: 487 QQRPYNPPQQQPRPQAPYNQQ-----NQKQQFDPLPMTYGALLPSLLAQNLVQTIP-PPR 540
Q +P NP QQQP+ Q P QQ Q++QF P Y P Q Q P PP+
Sbjct: 27 QLQPQNPSQQQPQEQVPLMQQQQQFPGQQEQFPP-QQPYPHQQPFPSQQPYPQPQPFPPQ 85
Query: 541 IPDPLPRWYRP 551
+P P + + P
Sbjct: 86 LPYPQTQPFPP 96
Score = 35.4 bits (80), Expect = 0.90
Identities = 29/81 (35%), Positives = 33/81 (39%), Gaps = 9/81 (11%)
Query: 422 HGGPQPVYSNHPFVANITPQMTAPQNPNY--QSPRPQGPAPYYPPLYQPLYNLQQLSQQP 479
H P P +P PQ+ PQ + Q P PQ P P YP QP+ SQQ
Sbjct: 66 HQQPFPSQQPYPQPQPFPPQLPYPQTQPFPPQQPYPQ-PQPQYPQPQQPI------SQQQ 118
Query: 480 YYPQQPYQQRPYNPPQQQPRP 500
QQ QQ QQQ P
Sbjct: 119 AQQQQQQQQTLQQILQQQLIP 139
>GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin)
Length = 307
Score = 48.5 bits (114), Expect = 1e-04
Identities = 43/123 (34%), Positives = 50/123 (39%), Gaps = 15/123 (12%)
Query: 423 GGPQPVYSNHPFVANITPQMTAPQNPNYQ-SPRPQGPAPY-YPPLYQPLYNLQQLSQQPY 480
G QP P+ PQ Q P Q P PQ PY P L P L QP+
Sbjct: 52 GQQQPFPPQQPYPQ---PQPFPSQQPYLQLQPFPQPQLPYPQPQLPYPQPQLPYPQPQPF 108
Query: 481 YPQQPYQQRPYNPPQQQP---RPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIP 537
PQQPY PQ QP +PQ P +QQ Q+QQ +L Q L Q +
Sbjct: 109 RPQQPY-------PQSQPQYSQPQQPISQQQQQQQQQQQQKQQQQQQQQILQQILQQQLI 161
Query: 538 PPR 540
P R
Sbjct: 162 PCR 164
Score = 47.4 bits (111), Expect = 2e-04
Identities = 41/115 (35%), Positives = 49/115 (41%), Gaps = 33/115 (28%)
Query: 440 PQMTAPQNPNYQSPRPQGPAPYYPPLYQPLYNLQQL--SQQPYYPQQPYQQRPYNPPQQQ 497
PQ+ PQNP+ Q P+ Q PL QQ QQP+ PQQPY PQ Q
Sbjct: 26 PQLQ-PQNPSQQQPQEQ----------VPLVQQQQFPGQQQPFPPQQPY-------PQPQ 67
Query: 498 PRP-QAPYNQQNQKQQFDPLPMTYGALLPSLLAQNLVQTIPPPRIPDPLPRWYRP 551
P P Q PY Q P P P L P P++P P P+ +RP
Sbjct: 68 PFPSQQPY------LQLQPFPQ------PQLPYPQPQLPYPQPQLPYPQPQPFRP 110
Score = 42.4 bits (98), Expect = 0.007
Identities = 29/81 (35%), Positives = 36/81 (43%), Gaps = 5/81 (6%)
Query: 487 QQRPYNPPQQQPRPQAPYNQQ----NQKQQFDP-LPMTYGALLPSLLAQNLVQTIPPPRI 541
Q +P NP QQQP+ Q P QQ Q+Q F P P PS +Q P P++
Sbjct: 27 QLQPQNPSQQQPQEQVPLVQQQQFPGQQQPFPPQQPYPQPQPFPSQQPYLQLQPFPQPQL 86
Query: 542 PDPLPRWYRPDLHCIYHQGAP 562
P P P+ P Y Q P
Sbjct: 87 PYPQPQLPYPQPQLPYPQPQP 107
>NAB2_YEAST (P32505) Nuclear polyadenylated RNA-binding protein NAB2
Length = 525
Score = 48.1 bits (113), Expect = 1e-04
Identities = 30/73 (41%), Positives = 34/73 (46%), Gaps = 1/73 (1%)
Query: 473 QQLSQQPYYPQQPYQQRPYNPPQQQPRPQAPYNQQNQKQQFDPLPMTYGALLPSLLAQNL 532
QQ QQP QQ QQ+P PQQQP+ Q Q Q QQ L P L QN
Sbjct: 111 QQQQQQPDIAQQQPQQQPQQQPQQQPQQQPQQQPQQQPQQQPQQQPQLQPLQPQLGTQNA 170
Query: 533 VQTIPPPRIPDPL 545
+QT P P P+
Sbjct: 171 MQT-DAPATPSPI 182
>YG49_SCHPO (O60184) Protein C23E6.09 in chromosome II
Length = 1102
Score = 47.8 bits (112), Expect = 2e-04
Identities = 31/82 (37%), Positives = 41/82 (49%), Gaps = 11/82 (13%)
Query: 432 HPFVANITPQMTAPQNPNYQSPRPQGPAPYYP-PLYQPLYNLQQLSQQPYYPQQPYQQRP 490
HP + + P +++ Q P Q P P +PY P PL Q QQ PQQ QQ+
Sbjct: 133 HPQLQRMMPILSSNQ-PIQQLPLPNQASPYIPVPLQQ---------QQQSQPQQQPQQQQ 182
Query: 491 YNPPQQQPRPQAPYNQQNQKQQ 512
+ PQQ PQ P QQ Q++Q
Sbjct: 183 HQQPQQPQPPQQPLQQQQQQRQ 204
Score = 33.1 bits (74), Expect = 4.5
Identities = 39/133 (29%), Positives = 51/133 (38%), Gaps = 27/133 (20%)
Query: 425 PQPVYSNHPFVANIT--PQMTAPQNPNYQS-PRPQGPAPYYPPLYQPLYNLQQ-----LS 476
PQ Y P AN P P++ + + Q PA P L N+QQ ++
Sbjct: 36 PQTAYYASPLHANSVSLPPSHLPRSTLHPLLSQQQQPAQQSPSLGPAQQNIQQPPSVSIA 95
Query: 477 QQPYYPQ-----------QPYQQRP------YNPPQQQPRPQ--APYNQQNQKQQFDPLP 517
QP+Y + Q Y+Q P N PQQ P+ Q P NQ Q PLP
Sbjct: 96 SQPHYAEAIVPIQQVLQPQQYRQLPPNMVAATNAPQQHPQLQRMMPILSSNQPIQQLPLP 155
Query: 518 MTYGALLPSLLAQ 530
+P L Q
Sbjct: 156 NQASPYIPVPLQQ 168
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.133 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 145,159,703
Number of Sequences: 164201
Number of extensions: 6939578
Number of successful extensions: 32452
Number of sequences better than 10.0: 463
Number of HSP's better than 10.0 without gapping: 129
Number of HSP's successfully gapped in prelim test: 352
Number of HSP's that attempted gapping in prelim test: 25362
Number of HSP's gapped (non-prelim): 2947
length of query: 1155
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1034
effective length of database: 40,105,733
effective search space: 41469327922
effective search space used: 41469327922
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC145024.28


