

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.2 - phase: 0
(355 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
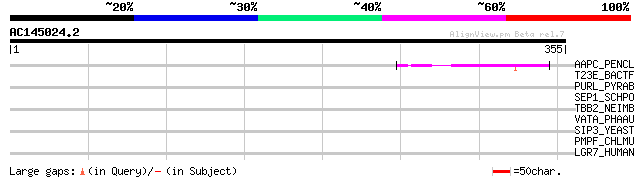
AAPC_PENCL (Q40784) Possible apospory-associated protein C 46 1e-04
T23E_BACTF (Q02403) Transposase for insertion sequence element I... 35 0.29
PURL_PYRAB (Q9UXW6) Phosphoribosylformylglycinamidine synthase I... 33 1.4
SEP1_SCHPO (O43058) Forkhead protein sep1 32 1.9
TBB2_NEIMB (Q06988) Transferrin-binding protein 2 precursor (TBP-2) 32 2.4
VATA_PHAAU (P13548) Vacuolar ATP synthase catalytic subunit A (E... 31 4.2
SIP3_YEAST (P38717) SIP3 protein 31 4.2
PMPF_CHLMU (Q9PL46) Probable outer membrane protein pmpF precurs... 30 9.3
LGR7_HUMAN (Q9HBX9) Relaxin receptor 1 (Leucine-rich repeat-cont... 30 9.3
>AAPC_PENCL (Q40784) Possible apospory-associated protein C
Length = 329
Score = 46.2 bits (108), Expect = 1e-04
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 20/105 (19%)
Query: 248 WTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIY 307
+T+QG I + E+++ +V+ A P SK IID ++ + V + G D
Sbjct: 214 FTEQGDAI-VFESEVDKVYLAAP------------SKIAIIDHEKKKTFVVTKEGLPDAV 260
Query: 308 VSSPGSMSDKYGRDY-------FVCTGPASILEPVTVNPGEEFRG 345
V +P K +D+ +C PA++ +P+T+ PGEE+RG
Sbjct: 261 VWNPWDKKAKAMQDFGDAEYKNMLCVEPAAVEKPITLKPGEEWRG 305
>T23E_BACTF (Q02403) Transposase for insertion sequence element
IS231E
Length = 478
Score = 35.0 bits (79), Expect = 0.29
Identities = 41/189 (21%), Positives = 72/189 (37%), Gaps = 44/189 (23%)
Query: 18 SSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYKP 77
S T L +PE LN +F +K + FL+ I ++N + +P + +T
Sbjct: 67 SQLHAVTGTLMSPEGLNKRFNKKAVCFLKH----IFSTLLKNKICETSLIPSSSIT---- 118
Query: 78 KVFWKDDGLEELLYTIPANETGLYKAKGGI----------------GLVLNEVLQPGAKE 121
F + L+ ++ +P + +Y GG G LN ++PG
Sbjct: 119 -YFQRIRILDATIFQVPKHLANVYPGSGGCAQTAGIKIQLEYDLHSGQFLNFQVEPGKNN 177
Query: 122 -----------LLPSTLEWTVKDVDYDAIDALQ------VELISTNRFFDMTYIVSLYPV 164
L P L ++D+ Y ++D L V IS + +M YI + +P
Sbjct: 178 DKTFGTECLATLRPGDL--CIRDLGYYSLDDLDQMDQRGVYYISRLKLNNMVYIKNEFPE 235
Query: 165 SMATAVIVK 173
++ K
Sbjct: 236 YFRNGIVKK 244
>PURL_PYRAB (Q9UXW6) Phosphoribosylformylglycinamidine synthase II
(EC 6.3.5.3) (FGAM synthase II)
Length = 705
Score = 32.7 bits (73), Expect = 1.4
Identities = 20/45 (44%), Positives = 22/45 (48%), Gaps = 2/45 (4%)
Query: 198 AAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPE 242
A KGL Y PL P +TP E M SESQ + F EPE
Sbjct: 266 AGKKGLGAVIYADRVPLREPG--MTPLEVMISESQERMLFAVEPE 308
>SEP1_SCHPO (O43058) Forkhead protein sep1
Length = 663
Score = 32.3 bits (72), Expect = 1.9
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 7/60 (11%)
Query: 203 LQTCSYCSHPPLDSPFQILTPSEAMKSES---QRLISFGAEPEMKPGSWTQQGVPITLLE 259
LQ+C+ PPL SP +PSE++++ES + S G + P +G P++ LE
Sbjct: 327 LQSCTNSPSPPLSSPASSASPSESLRNESLGIKSAKSLGLNKDDAP----VEGPPVSHLE 382
>TBB2_NEIMB (Q06988) Transferrin-binding protein 2 precursor (TBP-2)
Length = 599
Score = 32.0 bits (71), Expect = 2.4
Identities = 68/291 (23%), Positives = 108/291 (36%), Gaps = 53/291 (18%)
Query: 34 NDKFGRKGIKFLESNSIPIVELTVR---------NGSSLSLRLPDAHVTSYKPKVFWKDD 84
N + R G +L N+I I V G S LP +T YK + D
Sbjct: 138 NFTYVRSGYVYLNKNNIDIKNNIVLFGPDGYLYYKGKEPSKELPSEKIT-YKGTWDYVTD 196
Query: 85 GLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDYDAIDALQV 144
+E+ + GL A GG L+ G +L + E + D+ +V
Sbjct: 197 AMEKQRFE------GLGSAAGGDKSGALSALEEG---VLRNQAEASSGHTDFGMTSEFEV 247
Query: 145 ELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLT-NAILSHFRFKRRGGAAIKGL 203
+ F D T +LY + T +NK K T A L RFK + AA KG
Sbjct: 248 D------FSDKTIKGTLYRNNRITQNNSENKQIKTTRYTIQATLHGNRFKGKALAADKG- 300
Query: 204 QTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMS 263
+ SHP ++ S++++ +G + E G + +NK++
Sbjct: 301 --ATNGSHP-------FISDSDSLEGGF-----YGPKGEELAGKFLSN-------DNKVA 339
Query: 264 RVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSM 314
VF A K++ P+ +ID YR+ F+ + S G +
Sbjct: 340 AVFGAKQKDKKDGENAAGPATETVIDA-----YRITGEEFKKEQIDSFGDV 385
>VATA_PHAAU (P13548) Vacuolar ATP synthase catalytic subunit A (EC
3.6.3.14) (V-ATPase A subunit) (Vacuolar proton pump
alpha subunit) (V-ATPase 69 kDa subunit) (VAA3-1)
Length = 622
Score = 31.2 bits (69), Expect = 4.2
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 4/76 (5%)
Query: 256 TLLENK-MSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIY---VSSP 311
T+ EN M A PP K Y PP +Y I D E+ ++ ++ F + V +P
Sbjct: 156 TVFENTLMQHHIALPPDAMGKITYIAPPGQYSITDTVLELEFQGVKKKFTMLQTWPVRTP 215
Query: 312 GSMSDKYGRDYFVCTG 327
++ K D + TG
Sbjct: 216 RPVASKLAADTPLLTG 231
>SIP3_YEAST (P38717) SIP3 protein
Length = 1229
Score = 31.2 bits (69), Expect = 4.2
Identities = 18/57 (31%), Positives = 26/57 (45%)
Query: 139 IDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRR 195
+DAL+ + T FFD Y VS A ++ P P L+N +S+ F R
Sbjct: 41 VDALEDWIEKTVDFFDQKYKVSFEDFRRAKETLLSQLLPPPALLSNGFVSNQSFTPR 97
>PMPF_CHLMU (Q9PL46) Probable outer membrane protein pmpF precursor
(Polymorphic membrane protein F)
Length = 1025
Score = 30.0 bits (66), Expect = 9.3
Identities = 13/30 (43%), Positives = 18/30 (59%)
Query: 326 TGPASILEPVTVNPGEEFRGAQVIEHDNLS 355
T P + VT+NP +EF GA V + N+S
Sbjct: 426 TPPENSPHTVTINPSDEFSGAVVFSYKNIS 455
>LGR7_HUMAN (Q9HBX9) Relaxin receptor 1 (Leucine-rich
repeat-containing G protein-coupled receptor 7)
Length = 757
Score = 30.0 bits (66), Expect = 9.3
Identities = 39/163 (23%), Positives = 59/163 (35%), Gaps = 30/163 (18%)
Query: 66 RLPDAHVTSYKPKVFWKD------DGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGA 119
RLPD + + P++ W D L L + +N T L K I + P
Sbjct: 236 RLPDKPLCQHMPRLHWLDLEGNHIHNLRNLTFISCSNLTVLVMRKNKINHLNENTFAPLQ 295
Query: 120 K-----------ELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMAT 168
K E LP + +K++ + ++ I N+F Y+V L +S+
Sbjct: 296 KLDELDLGSNKIENLPPLIFKDLKELSQLNLSYNPIQKIQANQF---DYLVKLKSLSLEG 352
Query: 169 AVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSH 211
I + L N LSH FK+ Q C Y H
Sbjct: 353 IEISNIQQRMFRPLMN--LSHIYFKK--------FQYCGYAPH 385
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,050,476
Number of Sequences: 164201
Number of extensions: 1836214
Number of successful extensions: 4114
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4107
Number of HSP's gapped (non-prelim): 10
length of query: 355
length of database: 59,974,054
effective HSP length: 111
effective length of query: 244
effective length of database: 41,747,743
effective search space: 10186449292
effective search space used: 10186449292
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 66 (30.0 bits)
Medicago: description of AC145024.2


