

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
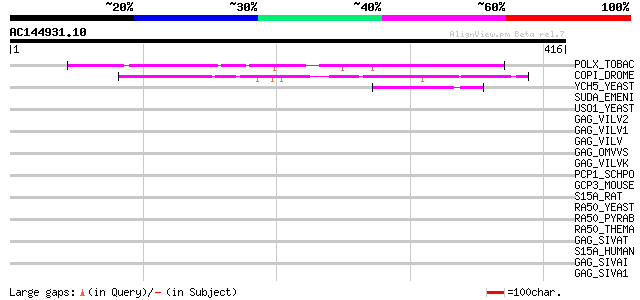
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 86 2e-16
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 62 2e-09
YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein 48 5e-05
SUDA_EMENI (Q00737) Chromosome segregation protein sudA (DA-box ... 40 0.008
USO1_YEAST (P25386) Intracellular protein transport protein USO1 40 0.014
GAG_VILV2 (P23425) Gag polyprotein [Contains: Core protein p16; ... 39 0.032
GAG_VILV1 (P23424) Gag polyprotein [Contains: Core protein p16; ... 39 0.032
GAG_VILV (P03352) Gag polyprotein [Contains: Core protein p16; C... 39 0.032
GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16; ... 39 0.032
GAG_VILVK (P35955) Gag polyprotein [Contains: Core protein p16; ... 38 0.042
PCP1_SCHPO (Q92351) Spindle pole body protein pcp1 38 0.055
GCP3_MOUSE (P58854) Gamma-tubulin complex component 3 (GCP-3) (F... 38 0.055
S15A_RAT (O54923) Exocyst complex component Sec15A (rSec15) 37 0.072
RA50_YEAST (P12753) DNA repair protein RAD50 (153 kDa protein) 37 0.072
RA50_PYRAB (Q9UZC8) DNA double-strand break repair rad50 ATPase 37 0.072
RA50_THEMA (Q9X1X1) Probable DNA double-strand break repair rad5... 37 0.094
GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17; ... 37 0.094
S15A_HUMAN (Q8TAG9) Exocyst complex component Sec15A 36 0.16
GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17; ... 36 0.21
GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17; ... 36 0.21
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 85.5 bits (210), Expect = 2e-16
Identities = 76/341 (22%), Positives = 142/341 (41%), Gaps = 29/341 (8%)
Query: 44 RALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKE 103
RA + I L DD+ I++ + A+ W +L Y T K L L+++ AL M E
Sbjct: 58 RAASAIRLHLSDDVVNNIIDEDTARGIWTRLESLYMSKTLTNK---LYLKKQLYALHMSE 114
Query: 104 TETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITV 163
+ + +ITQ+ LG + ++ +L LP ++ +++ K + I +
Sbjct: 115 GTNFLSHLNVFNGLITQLANLGVKIEEEDKAILLLNSLPSSYDNLATTILHGK--TTIEL 172
Query: 164 PELVNALQASEQRRSLRMEENVEGAFLASNKGKN--QSFKSFGEKKFPPCPHCKKDTHLD 221
++ +AL +E+ R + EN A + +G++ +S ++G + + +
Sbjct: 173 KDVTSALLLNEKMR--KKPENQGQALITEGRGRSYQRSSNNYGRSGARGKSKNRSKSRVR 230
Query: 222 KFCWYRPGVKCRACNQLGHVEKVCKNK-------TNQQEQQARVVEHHQEDEEQLF---K 271
C CNQ GH ++ C N + Q+ D LF +
Sbjct: 231 N---------CYNCNQPGHFKRDCPNPRKGKGETSGQKNDDNTAAMVQNNDNVVLFINEE 281
Query: 272 ASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVE 331
C + W++D+ ++H T F + V GN + K+ G G + ++
Sbjct: 282 EECMHLSGPESEWVVDTAASHHATPVRDLFCRYVAGDFGTVKMGNTSYSKIAGIGDICIK 341
Query: 332 TLLGIKYIL-DVLFVPEINQSLLSVGQMMEKNYSLHFKNMK 371
T +G +L DV VP++ +L+S + Y +F N K
Sbjct: 342 TNVGCTLVLKDVRHVPDLRMNLISGIALDRDGYESYFANQK 382
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 62.4 bits (150), Expect = 2e-09
Identities = 70/327 (21%), Positives = 137/327 (41%), Gaps = 42/327 (12%)
Query: 82 ERTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCL 141
ER L LR+ +LK+ ++ ++I+++ G + + + +L+ L
Sbjct: 89 ERKSLASQLALRKRLLSLKLSSEMSLLSHFHIFDELISELLAAGAKIEEMDKISHLLITL 148
Query: 142 PEMFEAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEEN-----VEGAFLASNKG- 195
P ++ I+++ E + +T+ + N L +Q ++ + N V A + +N
Sbjct: 149 PSCYDGIITAI-ETLSEENLTLAFVKNRL--LDQEIKIKNDHNDTSKKVMNAIVHNNNNT 205
Query: 196 -KNQSFKS---------FGEKKFP-PCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKV 244
KN FK+ G K+ C HC ++ H+ K C+ H +++
Sbjct: 206 YKNNLFKNRVTKPKKIFKGNSKYKVKCHHCGREGHIKKDCF--------------HYKRI 251
Query: 245 CKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKEL 304
NK + E+Q + H + K + +++DSG ++H+ N+ S + +
Sbjct: 252 LNNKNKENEKQVQTATSH--GIAFMVKEVNNTSVMDNCGFVLDSGASDHLINDESLYTDS 309
Query: 305 DESF--YSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKN 362
E V G+ + +G+V + I + DVLF E +L+SV ++ E
Sbjct: 310 VEVVPPLKIAVAKQGEFIYATKRGIVRLRNDHEIT-LEDVLFCKEAAGNLMSVKRLQEAG 368
Query: 363 YSLHFKNMKCTIFDPYGSKLMIVEMRG 389
S+ F TI + LM+V+ G
Sbjct: 369 MSIEFDKSGVTISK---NGLMVVKNSG 392
>YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein
Length = 146
Score = 47.8 bits (112), Expect = 5e-05
Identities = 25/83 (30%), Positives = 42/83 (50%), Gaps = 4/83 (4%)
Query: 273 SCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
S + + W+ D+GCT+HM ++ S F S V G G + + G G V + T
Sbjct: 68 SSTVPATKSSEWIFDTGCTSHMCHDRSIFSSFTRSSRKDFVRGVGGSIPIMGSGTVNIGT 127
Query: 333 LLGIKYILDVLFVPEINQSLLSV 355
+ + DV +VP++ +L+SV
Sbjct: 128 V----QLHDVSYVPDLPVNLISV 146
>SUDA_EMENI (Q00737) Chromosome segregation protein sudA (DA-box
protein sudA)
Length = 1211
Score = 40.4 bits (93), Expect = 0.008
Identities = 29/120 (24%), Positives = 57/120 (47%), Gaps = 2/120 (1%)
Query: 88 KVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSD-QRVVEKILVCLPEMFE 146
++ +++ E LK+ + + E + SK + Q+ L + LSD Q ++ E +
Sbjct: 270 EIAECKQQIEFLKVDKAQLEDERREA-SKALAQVELQAKSLSDNQAAAQESKARHDESLK 328
Query: 147 AKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEK 206
A S++EE + + VP ++A A + R+ E L + +G+N FK+ E+
Sbjct: 329 AVQSAIEERQTELKELVPRFISAKDAEDAARAKLTEAETARQRLYAKQGRNSRFKNKSER 388
>USO1_YEAST (P25386) Intracellular protein transport protein USO1
Length = 1790
Score = 39.7 bits (91), Expect = 0.014
Identities = 50/242 (20%), Positives = 101/242 (41%), Gaps = 34/242 (14%)
Query: 30 AQIRNHSDEVAKEGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKV 89
AQ++ + +++A + R + L+D+I E K+ ++L E + + T + +
Sbjct: 1159 AQLKKYEEQIANKERQYNEEISQLNDEITSTQQENESIKKKNDELEGEVKAMKSTSEEQS 1218
Query: 90 LNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKI 149
+ E +AL ++ E ++ + ++ I+ + E KI
Sbjct: 1219 NLKKSEIDALNLQIKELKKKNETNEASLLESIKSV------------------ESETVKI 1260
Query: 150 SSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFP 209
L++ NF E V EL + L+ASE + S +E E S K K + E K
Sbjct: 1261 KELQDECNFKEKEVSELEDKLKASEDKNSKYLELQKE-----SEKIKEELDAKTTELKI- 1314
Query: 210 PCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQL 269
+K T+L K K ++ ++L ++K + E+Q +++ + + Q
Sbjct: 1315 ---QLEKITNLSK-------AKEKSESELSRLKKTSSEERKNAEEQLEKLKNEIQIKNQA 1364
Query: 270 FK 271
F+
Sbjct: 1365 FE 1366
Score = 30.8 bits (68), Expect = 6.7
Identities = 30/129 (23%), Positives = 57/129 (43%), Gaps = 24/129 (18%)
Query: 52 ALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFS 111
+L +DI KI ++ A N+ +EE K++ NL +E E + + E F
Sbjct: 903 SLKEDIAAKITEIK----AINENLEEM-------KIQCNNLSKEKEHISKELVEYKSRFQ 951
Query: 112 DR---ISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVN 168
++K+ +++ L + D + E+ I ++EE+KN S I + L N
Sbjct: 952 SHDNLVAKLTEKLKSLANNYKDMQAEN----------ESLIKAVEESKNESSIQLSNLQN 1001
Query: 169 ALQASEQRR 177
+ + Q +
Sbjct: 1002 KIDSMSQEK 1010
>GAG_VILV2 (P23425) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 442
Score = 38.5 bits (88), Expect = 0.032
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 13/61 (21%)
Query: 191 ASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
A +KG NQ C +C K HL + C R G+ C C + GH++K C+ K
Sbjct: 378 AGHKGVNQK-----------CYNCGKPGHLARQC--RQGIICHHCGKRGHMQKDCRQKKQ 424
Query: 251 Q 251
Q
Sbjct: 425 Q 425
>GAG_VILV1 (P23424) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 442
Score = 38.5 bits (88), Expect = 0.032
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 13/61 (21%)
Query: 191 ASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
A +KG NQ C +C K HL + C R G+ C C + GH++K C+ K
Sbjct: 378 AGHKGVNQK-----------CYNCGKPGHLARQC--RQGIICHHCGKRGHMQKDCRQKKQ 424
Query: 251 Q 251
Q
Sbjct: 425 Q 425
>GAG_VILV (P03352) Gag polyprotein [Contains: Core protein p16; Core
protein p25; Core protein p14]
Length = 442
Score = 38.5 bits (88), Expect = 0.032
Identities = 21/61 (34%), Positives = 29/61 (47%), Gaps = 13/61 (21%)
Query: 191 ASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
A +KG NQ C +C K HL + C R G+ C C + GH++K C+ K
Sbjct: 378 AGHKGVNQK-----------CYNCGKPGHLARQC--RQGIICHHCGKRGHMQKDCRQKKQ 424
Query: 251 Q 251
Q
Sbjct: 425 Q 425
>GAG_OMVVS (P16900) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 446
Score = 38.5 bits (88), Expect = 0.032
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Query: 204 GEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCK----NKTNQQEQQAR 257
G++ C +C K HL + C R G+ C C + GH++K C+ N T+QQ R
Sbjct: 379 GKQGVQKCYYCGKPGHLARQC--RQGIICHHCGKRGHMQKDCRQKKGNPTSQQGNSRR 434
>GAG_VILVK (P35955) Gag polyprotein [Contains: Core protein p16;
Core protein p25; Core protein p14]
Length = 442
Score = 38.1 bits (87), Expect = 0.042
Identities = 21/61 (34%), Positives = 28/61 (45%), Gaps = 13/61 (21%)
Query: 191 ASNKGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
A KG NQ C +C K HL + C R G+ C C + GH++K C+ K
Sbjct: 378 AGQKGVNQK-----------CYNCGKPGHLARQC--RQGIICHHCGKRGHMQKDCRQKKQ 424
Query: 251 Q 251
Q
Sbjct: 425 Q 425
>PCP1_SCHPO (Q92351) Spindle pole body protein pcp1
Length = 1208
Score = 37.7 bits (86), Expect = 0.055
Identities = 33/140 (23%), Positives = 66/140 (46%), Gaps = 10/140 (7%)
Query: 59 IKILNLEIAKEAWNKLMEEYQGS-ERTKKMKVLNLRREFEALKMKETETVREFSDRISKV 117
I++L ++ KE N++M E + S T M+V L+ + K E + D+IS++
Sbjct: 361 IQVLTADLEKEKENQIMHESEASIGLTDSMQVHTLQEQLH----KANEEIEFLHDQISRM 416
Query: 118 ITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQA-SEQR 176
+ G++ D + + L ++ E+K+ +LE++ N + L N +++ Q
Sbjct: 417 NEE----GKNFEDIMLQFRSLEEERDVLESKLQTLEDDNNSLRLMTSSLGNQIESLRTQN 472
Query: 177 RSLRMEENVEGAFLASNKGK 196
R + E+N + N K
Sbjct: 473 REIDEEKNHLRLLASKNSDK 492
>GCP3_MOUSE (P58854) Gamma-tubulin complex component 3 (GCP-3)
(Fragment)
Length = 677
Score = 37.7 bits (86), Expect = 0.055
Identities = 29/109 (26%), Positives = 54/109 (48%), Gaps = 7/109 (6%)
Query: 57 IFIKILNLEIAKE-----AWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFS 111
+F +I+ L+ A++ A +L Q E+ K+ ++ A + +E + +REF
Sbjct: 548 VFDQIIELQNAQDVMYRAALEELQRRLQFEEKKKQREIEGQWGVTAAEEEEENKRIREFQ 607
Query: 112 DRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSE 160
D I K+ +Q+R+L Q VV++ LV L + + L +F+E
Sbjct: 608 DSIPKMCSQLRILTHFY--QGVVQQFLVLLTTSSDESLQFLSFRLDFNE 654
>S15A_RAT (O54923) Exocyst complex component Sec15A (rSec15)
Length = 804
Score = 37.4 bits (85), Expect = 0.072
Identities = 28/88 (31%), Positives = 47/88 (52%), Gaps = 7/88 (7%)
Query: 88 KVLNLRREFEALKMKETETVREFSDRISKVITQ----IRLLGEDLSDQRVVEKILVCLP- 142
++L +R + E LK++ T+T R F D +VI Q IR + + VVEK+ +CLP
Sbjct: 79 ELLKVRADAEKLKVQVTDTNRRFQDAGKEVIEQTEDIIRCRIQQRNITTVVEKLQLCLPV 138
Query: 143 -EMFEAKISSLEENKNFSEI-TVPELVN 168
EM+ + + +S + T+ +L N
Sbjct: 139 LEMYSKLKEQMSMQRYYSALKTMEQLEN 166
>RA50_YEAST (P12753) DNA repair protein RAD50 (153 kDa protein)
Length = 1312
Score = 37.4 bits (85), Expect = 0.072
Identities = 49/230 (21%), Positives = 96/230 (41%), Gaps = 33/230 (14%)
Query: 1 MKTYLRAQSLWD---VVEKGNNPPPLPNNPTLA---QIRNHSDEVAKEGRALAIIHAALH 54
+KTY W+ ++ K N NN + QI D + K + A ++A L
Sbjct: 477 LKTYKEKLQSWESENIIPKLNQKIEEKNNEMIILENQIEKFQDRIMKTNQQ-ADLYAKL- 534
Query: 55 DDIFIKILNLEIAKEAWNKLMEEYQGSER-------TKKMKVLNLRREFEALKMKETETV 107
+ K +N ++ + K+ E+ Q R T++ + +L +F+ L + + +
Sbjct: 535 -GLIKKSINTKL--DELQKITEKLQNDSRIRQVFPLTQEFQRADLEMDFQKLFINMQKNI 591
Query: 108 -------REFSDRISKVITQIRLLGEDLSD-QRVVEKILVCLPEMF--EAKISSLEENKN 157
E R + + + + +DL D Q+ EK++ L E + I +
Sbjct: 592 AINNKKMHELDRRYTNALYNLNTIEKDLQDNQKSKEKVIQLLSENLPEDCTIDEYNDVLE 651
Query: 158 FSEITVPELVNALQASE-----QRRSLRMEENVEGAFLASNKGKNQSFKS 202
+E++ + L+ + R++L + E +L S K +N+SFKS
Sbjct: 652 ETELSYKTALENLKMHQTTLEFNRKALEIAERDSCCYLCSRKFENESFKS 701
>RA50_PYRAB (Q9UZC8) DNA double-strand break repair rad50 ATPase
Length = 880
Score = 37.4 bits (85), Expect = 0.072
Identities = 48/224 (21%), Positives = 95/224 (41%), Gaps = 42/224 (18%)
Query: 55 DDIFIKILNLEIAKEAWNKLM---EEYQGSERTKKMKVLNLRREFEAL---------KMK 102
++I I L+ ++ + KL +EY+ R + ++ E +A+ K +
Sbjct: 283 EEIVKDIPKLQEKEKEYRKLKGFRDEYESKLRRLEKELSKWESELKAIEEVIKEGEKKKE 342
Query: 103 ETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCL--------PEMFEAKISSLEE 154
E +RE I K + +++ E+L D + V+K + L P K+ SLE+
Sbjct: 343 RAEEIREKLSEIEKRLEELKPYVEELEDAKQVQKQIERLKARLKGLSPGEVIEKLESLEK 402
Query: 155 NKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPPCPHC 214
+ E + E+ + EQ ++ RM+ E L KGK CP C
Sbjct: 403 ERTEIEEAIKEITTRIGQMEQEKNERMKAIEE---LRKAKGK--------------CPVC 445
Query: 215 KKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARV 258
++ + ++ + R ++ +E+ K +T ++E++ RV
Sbjct: 446 GRELTEE----HKKELMERYTLEIKKIEEELK-RTTEEERKLRV 484
>RA50_THEMA (Q9X1X1) Probable DNA double-strand break repair rad50
ATPase
Length = 852
Score = 37.0 bits (84), Expect = 0.094
Identities = 24/79 (30%), Positives = 42/79 (52%), Gaps = 1/79 (1%)
Query: 53 LHDDIFIKILNLEIAKEAWNKLM-EEYQGSERTKKMKVLNLRREFEALKMKETETVREFS 111
L D F ++ + I E + L+ EE + +E+ + +R E+LK E+E VR+ S
Sbjct: 584 LKSDFFDRLRKIGIGFEEFRILVKEEVKDAEKELGVVETEIRLLEESLKELESENVRDVS 643
Query: 112 DRISKVITQIRLLGEDLSD 130
+ KV Q+ L +++SD
Sbjct: 644 EDYEKVRNQLEALSQEISD 662
>GAG_SIVAT (P05892) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 519
Score = 37.0 bits (84), Expect = 0.094
Identities = 22/89 (24%), Positives = 39/89 (43%), Gaps = 2/89 (2%)
Query: 164 PELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFPP--CPHCKKDTHLD 221
P L L A + + V + + + +N + +++ PP C +C K H+
Sbjct: 350 PTLEEMLTACQGVGGPSYKAKVMAEMMQTMQNQNMVQQGGPKRQRPPLRCYNCGKFGHMQ 409
Query: 222 KFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
+ C KC C +LGH+ K C+ + N
Sbjct: 410 RQCPEPRKTKCLKCGKLGHLAKDCRGQVN 438
>S15A_HUMAN (Q8TAG9) Exocyst complex component Sec15A
Length = 804
Score = 36.2 bits (82), Expect = 0.16
Identities = 27/88 (30%), Positives = 46/88 (51%), Gaps = 7/88 (7%)
Query: 88 KVLNLRREFEALKMKETETVREFSDRISKVITQ----IRLLGEDLSDQRVVEKILVCLP- 142
++L +R + E LK++ T+T R F D +VI IR + + VVEK+ +CLP
Sbjct: 79 ELLKVRTDAEKLKVQVTDTNRRFQDAGKEVIVHTEDIIRCRIQQRNITTVVEKLQLCLPV 138
Query: 143 -EMFEAKISSLEENKNFSEI-TVPELVN 168
EM+ + + +S + T+ +L N
Sbjct: 139 LEMYSKLKEQMSAKRYYSALKTMEQLEN 166
>GAG_SIVAI (Q02843) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 513
Score = 35.8 bits (81), Expect = 0.21
Identities = 14/37 (37%), Positives = 21/37 (55%)
Query: 211 CPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKN 247
C +C K H+ + C +KC C ++GH+ K CKN
Sbjct: 392 CFNCGKFGHMQRECKAPRQIKCFKCGKIGHMAKDCKN 428
>GAG_SIVA1 (P27972) Gag polyprotein [Contains: Core protein p17;
Core protein p24; Core protein p15]
Length = 520
Score = 35.8 bits (81), Expect = 0.21
Identities = 14/42 (33%), Positives = 22/42 (52%)
Query: 209 PPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTN 250
P C +C K H+ + C +KC C + GH+ K C+ + N
Sbjct: 398 PKCYNCGKFGHMQRQCPEPRKIKCLKCGKPGHLAKDCRGQVN 439
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,186,325
Number of Sequences: 164201
Number of extensions: 2020396
Number of successful extensions: 7116
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 82
Number of HSP's that attempted gapping in prelim test: 7007
Number of HSP's gapped (non-prelim): 161
length of query: 416
length of database: 59,974,054
effective HSP length: 113
effective length of query: 303
effective length of database: 41,419,341
effective search space: 12550060323
effective search space used: 12550060323
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144931.10


