

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
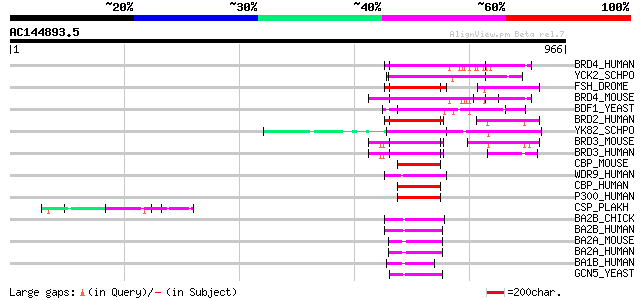
Score E
Sequences producing significant alignments: (bits) Value
BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1 prot... 115 6e-25
YCK2_SCHPO (Q9Y7N0) Hypothetical bromodomain protein C1450.02 103 2e-21
FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chor... 103 3e-21
BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic ch... 100 2e-20
BDF1_YEAST (P35817) BDF1 protein 100 2e-20
BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3 prot... 99 4e-20
YK82_SCHPO (Q9HGP4) Hypothetical bromodomain protein C631.02 96 4e-19
BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3 (Bromodomai... 92 9e-18
BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like... 91 1e-17
CBP_MOUSE (P45481) CREB-binding protein (EC 2.3.1.48) 75 6e-13
WDR9_HUMAN (Q9NSI6) WD-repeat protein 9 73 4e-12
CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48) 72 5e-12
P300_HUMAN (Q09472) E1A-associated protein p300 (EC 2.3.1.48) 71 2e-11
CSP_PLAKH (P02894) Circumsporozoite protein precursor (CS) 68 1e-10
BA2B_CHICK (Q9DE13) Bromodomain adjacent to zinc finger domain 2... 66 4e-10
BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2... 66 5e-10
BA2A_MOUSE (Q91YE5) Bromodomain adjacent to zinc finger domain 2... 64 1e-09
BA2A_HUMAN (Q9UIF9) Bromodomain adjacent to zinc finger domain 2... 64 1e-09
BA1B_HUMAN (Q9UIG0) Bromodomain adjacent to zinc finger domain p... 64 2e-09
GCN5_YEAST (Q03330) Histone acetyltransferase GCN5 (EC 2.3.1.48) 64 3e-09
>BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1
protein)
Length = 1362
Score = 115 bits (288), Expect = 6e-25
Identities = 96/337 (28%), Positives = 144/337 (42%), Gaps = 83/337 (24%)
Query: 653 SKFFKSCSSLLEKLMEHQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKN 709
S+ K CS +L+++ + YAW F PVDV+ LGLHDY II +PMD+ T+K++L
Sbjct: 351 SEQLKCCSGILKEMFAKKHAAYAWPFYKPVDVEALGLHDYCDIIKHPMDMSTIKSKLEAR 410
Query: 710 WYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMD 769
Y+ +EF DVRL F N YNP +V AMA +L +FE R+A + + +
Sbjct: 411 EYRDAQEFGADVRLMFSNCYKYNPPDHEVVAMARKLQDVFEMRFAKMPDEPEEPVVAVSS 470
Query: 770 YGAPSPLSRRVPAFTPPPPLD----------------MRRILDRQE-------------- 799
P P P + D +R+ + QE
Sbjct: 471 PAVPPPTKVVAPPSSSDSSSDSSSDSDSSTDDSEEERAQRLAELQEQLKAVHEQLAALSQ 530
Query: 800 PFARTPRSM--------------------NNTPSSRTPAPKKPK----------AKDP-- 827
P P+ N ++ P PKK K K+P
Sbjct: 531 PQQNKPKKKEKDKKEKKKEKHKRKEEVEENKKSKAKEPPPKKTKKNNSSNSNVSKKEPAP 590
Query: 828 --NKRDMTY-------------DEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*L-NQQDD 871
+K TY +EK++LS ++ LP EKL VV +++ + L N D
Sbjct: 591 MKSKPPPTYESEEEDKCKPMSYEEKRQLSLDINKLPGEKLGRVVHIIQSREPSLKNSNPD 650
Query: 872 EIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
EIE+DF+ + L EL+R+V + + K K +AE+
Sbjct: 651 EIEIDFETLKPSTLRELERYVTSCLR--KKRKPQAEK 685
Score = 87.0 bits (214), Expect = 2e-16
Identities = 63/191 (32%), Positives = 87/191 (44%), Gaps = 25/191 (13%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+L+ L +HQ+AW F PVD L L DY+ II PMD+GT+K RL N+Y + +E +D
Sbjct: 70 VLKTLWKHQFAWPFQQPVDAVKLNLPDYYKIIKTPMDMGTIKKRLENNYYWNAQECIQDF 129
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREM--------RYGMDYGAP 773
F N YN G D+ MAE L K+F + + ++ M R + G
Sbjct: 130 NTMFTNCYIYNKPGDDIVLMAEALEKLFLQKINELPTEETEIMIVQAKGRGRGRKETGTA 189
Query: 774 SPLSRRVP----AFTPP----PPLDMRRILDRQEPF-ARTPRSMNNTP--------SSRT 816
P VP A TPP P + + PF A TP + TP +T
Sbjct: 190 KPGVSTVPNTTQASTPPQTQTPQPNPPPVQATPHPFPAVTPDLIVQTPVMTVVPPQPLQT 249
Query: 817 PAPKKPKAKDP 827
P P P+ + P
Sbjct: 250 PPPVPPQPQPP 260
Score = 38.9 bits (89), Expect = 0.067
Identities = 88/453 (19%), Positives = 160/453 (34%), Gaps = 54/453 (11%)
Query: 52 SDTDNNNGTTLENIDEAPEFAVLGDGNLAKAEGSSRLEDGSSIQQQLGDQNLVEEHASSR 111
S +D+++ T + A A L + A E + L Q Q E+ +
Sbjct: 492 SSSDSDSSTDDSEEERAQRLAELQEQLKAVHEQLAALS-----QPQQNKPKKKEKDKKEK 546
Query: 112 TGDRNSPQQQLEE--LNSAQPHASFQLADGNSPQQKVENQDSAHPQASLRIGDGNSPQQQ 169
+++ ++++EE + A+ + NS V ++ A P S S ++
Sbjct: 547 KKEKHKRKEEVEENKKSKAKEPPPKKTKKNNSSNSNVSKKEPA-PMKSKPPPTYESEEED 605
Query: 170 LVEPNLAQPQASLRAGDGNSPQRQLEEE----NSAQPQASSRTGDGNLPHQQLEDQYSAQ 225
+P + + L P +L S +P + D ++E +
Sbjct: 606 KCKPMSYEEKRQLSLDINKLPGEKLGRVVHIIQSREPSLKNSNPD------EIEIDFETL 659
Query: 226 PPASLRIGDG---NSLRQKVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDD 282
P++LR + + LR+K + PQA + S + K S ++ S + D
Sbjct: 660 KPSTLRELERYVTSCLRKKRK------PQAEKVDVIAGSSKMKGFSSSESESSSESSSSD 713
Query: 283 GDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVN 342
+ ++ ++ ++ + H + + Q Q PQ +P P
Sbjct: 714 SEDSETEMAPKSKKKGHPGREQKKHHHHHHQQMQQAPAPVPQ-----QPPPPPQQPPPPP 768
Query: 343 SRLEDRSFPQPELNSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDL 402
+ + P P + + +S + P L +L GS P+ + T L
Sbjct: 769 PPQQQQQPPPPPPPPSMPQQAAPAMKSSPPPFIATQVPVLEPQLPGSVFDPIGHFTQPIL 828
Query: 403 NLAQP-----LSQSV--SDDLHINQQAEPSNLNVRQE-DDRPSSPIHRQGAISDNLHSHQ 454
+L QP L Q S H+NQ A S + +PS P +R A+
Sbjct: 829 HLPQPELPPHLPQPPEHSTPPHLNQHAVVSPPALHNALPQQPSRPSNRAAALPPKPARPP 888
Query: 455 QVEPS--------------NPNVLLEDDGPSSP 473
V P+ P VLLED+ P +P
Sbjct: 889 AVSPALTQTPLLPQPPMAQPPQVLLEDEEPPAP 921
>YCK2_SCHPO (Q9Y7N0) Hypothetical bromodomain protein C1450.02
Length = 578
Score = 103 bits (258), Expect = 2e-21
Identities = 67/241 (27%), Positives = 117/241 (47%), Gaps = 8/241 (3%)
Query: 659 CSSLLEKLMEHQY---AWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPK 715
CS++L++L + QY A+ F PVD DYF +I PMDL T++++LNKN Y + +
Sbjct: 260 CSTVLKELYKRQYESFAFPFYQPVDPVACDCPDYFDVIKEPMDLSTIQSKLNKNEYSTLE 319
Query: 716 EFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIE--SDYNREMRYGMDYGAP 773
EF D+ L F+N TYNP G VH M QL +F+++W D + + A
Sbjct: 320 EFESDILLMFNNCFTYNPPGTPVHVMGRQLENVFKEKWEARPKFDDATLVKQQEAETDAL 379
Query: 774 SPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKR--D 831
A ++ + + + ++ + + +KP+ +D K
Sbjct: 380 FDNGEEEEALMSEEEINGAKFAAVDKQISMLQDTLEAMKAKKMNRMRKPRRRDLTKEYGP 439
Query: 832 MTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRF 891
+TY + +L+ + L E+L V +++R++ L + DEIE+D + E+ + R+
Sbjct: 440 ITYAMQNELAERCNYLSAEQLSNVAEILREEMPWL-RDTDEIEIDVGNMKPEVFHRIYRY 498
Query: 892 V 892
V
Sbjct: 499 V 499
Score = 70.5 bits (171), Expect = 2e-11
Identities = 52/176 (29%), Positives = 80/176 (44%), Gaps = 31/176 (17%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K C +++ +L + + F PVD + DY TI+ NPMDLGT++ +L Y P+E
Sbjct: 91 KYCLAIVRQLKRTKNSAPFKVPVDPIKQNIPDYPTIVKNPMDLGTIEKKLTSYEYSVPQE 150
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMD----YGA 772
F +D+ L F N YN V +M + L ++FE R+++ D A
Sbjct: 151 FIDDMNLMFSNCFLYNGTESPVGSMGKALQEVFE-----------RQLKQLPDAEQPAAA 199
Query: 773 PSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMN-NTPSSRTPAPKKPKAKDP 827
P S++ A T PP RT R+ + ++ S+ A PKA P
Sbjct: 200 PVKKSKQKSASTAPP---------------RTRRNSSVSSTSASVAASTAPKAASP 240
>FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chorion
membrane protein)
Length = 2038
Score = 103 bits (256), Expect = 3e-21
Identities = 54/110 (49%), Positives = 67/110 (60%), Gaps = 3/110 (2%)
Query: 653 SKFFKSCSSLLEKLMEHQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKN 709
S KSC+ +L++L + YAW F PVD + LGLHDY II PMDLGTVK +++
Sbjct: 478 SDALKSCNEILKELFSKKHSGYAWPFYKPVDAEMLGLHDYHDIIKKPMDLGTVKRKMDNR 537
Query: 710 WYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESD 759
YKS EFA DVRL F N YNP DV AM +L +FE R+A I +
Sbjct: 538 EYKSAPEFAADVRLIFTNCYKYNPPDHDVVAMGRKLQDVFEMRYANIPDE 587
Score = 85.1 bits (209), Expect = 8e-16
Identities = 38/89 (42%), Positives = 54/89 (59%)
Query: 661 SLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAED 720
++++ + +H ++W F PVD L L DY II PMD+GT+K RL N+Y S KE +D
Sbjct: 45 TVMKVIWKHHFSWPFQQPVDAKKLNLPDYHKIIKQPMDMGTIKKRLENNYYWSAKETIQD 104
Query: 721 VRLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
F+N YN G+DV MA+ L K+F
Sbjct: 105 FNTMFNNCYVYNKPGEDVVVMAQTLEKVF 133
Score = 55.1 bits (131), Expect = 9e-07
Identities = 38/120 (31%), Positives = 65/120 (53%), Gaps = 12/120 (10%)
Query: 814 SRTPAPKKPKA-------KDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*L 866
+R + KKP ++ + M+YDEK++LS ++ LP +KL VV +++ + L
Sbjct: 927 ARGSSKKKPSQVMNFDSEEEDTAKPMSYDEKRQLSLDINKLPGDKLGRVVHIIQNREPSL 986
Query: 867 NQQD-DEIEVDFDAVDAEILWELDRFVLN--YKKSLSK--NKRKAEQARERAEALQNSVQ 921
+ DEIE+DF+ + L EL+ +V + KK+ K K K EQ E+ + L+ +Q
Sbjct: 987 RDSNPDEIEIDFETLKPSTLRELESYVASCLRKKTHKKPSGKSKDEQMAEKKQELEKRLQ 1046
Score = 32.7 bits (73), Expect = 4.8
Identities = 30/112 (26%), Positives = 46/112 (40%), Gaps = 8/112 (7%)
Query: 469 GPSSPIHRHKAVPSTEYRHSENVTEEPNLEDR-IKINLASTSKQEKQEMCLKLESELDAV 527
G SS + +H A S+ S+ E + E+R ++ + + QE KL E A
Sbjct: 623 GGSSSL-KHDASDSSSEDSSDTENESNSDEERSARLKMLESKLLGLQEEIRKLSEEASAK 681
Query: 528 RSLVKRIEVKQGYVGGYGNSNV------VLGGGITNGGGAQRAHSEAGSVGV 573
+ K+++ K+ +GG S GGG GG GSV V
Sbjct: 682 KKAKKKLKEKKKSIGGGSGSGSASHHCHATGGGANAGGAGGPGSGGHGSVSV 733
>BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic
chromosome-associated protein) (MCAP)
Length = 1400
Score = 100 bits (249), Expect = 2e-20
Identities = 76/256 (29%), Positives = 111/256 (42%), Gaps = 32/256 (12%)
Query: 625 PPAESNKKSKLNWKKQGGGDMGXGLQMGSKF---FKSCSSLLEKLMEHQ---YAWVFNTP 678
P ES++ K K G + SK K CS +L+++ + YAW F P
Sbjct: 321 PRRESSRPVKPPKKDVPDSQQHPGPEKSSKISEQLKCCSGILKEMFAKKHAAYAWPFYKP 380
Query: 679 VDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDV 738
VDV+ LGLHDY II +PMD+ T+K++L Y+ +EF DVRL F N YNP +V
Sbjct: 381 VDVEALGLHDYCDIIKHPMDMSTIKSKLESREYRDAQEFGADVRLMFSNCYKYNPPDHEV 440
Query: 739 HAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLD-------- 790
AMA +L +FE R+A + + + P P P + D
Sbjct: 441 VAMARKLQDVFEMRFAKMPDEPEEPVVTVSSPAVPPPTKVVAPPSSSDSSSDSSSDSDSS 500
Query: 791 --------MRRILDRQ-------EPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYD 835
+R+ + Q E A + N P + KK K K+ +K+ +
Sbjct: 501 TDDSEEERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKK-EKDKKEKKKEKHKKKEEVE 559
Query: 836 EKQKLSTSLHSLPVEK 851
E +K T LP +K
Sbjct: 560 ENKKSKTK--ELPPKK 573
Score = 89.0 bits (219), Expect = 6e-17
Identities = 61/197 (30%), Positives = 83/197 (41%), Gaps = 31/197 (15%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+L+ L +HQ+AW F PVD L L DY+ II PMD+GT+K RL N+Y + +E +D
Sbjct: 70 VLKTLWKHQFAWPFQQPVDAVKLNLPDYYKIIKTPMDMGTIKKRLENNYYWNAQECIQDF 129
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREM--------RYGMDYGAP 773
F N YN G D+ MAE L K+F + + ++ M R + GA
Sbjct: 130 NTMFTNCYIYNKPGDDIVLMAEALEKLFLQKINELPTEETEIMIVQAKGRGRGRKETGAA 189
Query: 774 SPLSRRVPAFT--------------PPPPLDMRR---------ILDRQEPFARTPRSMNN 810
P VP T PPPP+ ++ + P
Sbjct: 190 KPGVSTVPNTTQASTSPQTQTPQQNPPPPVQATTHPFPAVTPDLIAQPPVMTMVPPQPLQ 249
Query: 811 TPSSRTPAPKKPKAKDP 827
TPS P P P A P
Sbjct: 250 TPSPVPPQPPPPPAPVP 266
Score = 58.2 bits (139), Expect = 1e-07
Identities = 37/110 (33%), Positives = 61/110 (54%), Gaps = 10/110 (9%)
Query: 807 SMNNTPSSRTPAPKKPKA-------KDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMM 859
S N+ S + P P K K ++ + M+Y+EK++LS ++ LP EKL VV ++
Sbjct: 579 SSNSNVSKKEPVPTKTKPPPTYESEEEDKCKPMSYEEKRQLSLDINKLPGEKLGRVVHII 638
Query: 860 RKKNL*L-NQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAEQ 908
+ + L N DEIE+DF+ + L EL+R+V + + K K +AE+
Sbjct: 639 QSREPSLKNSNPDEIEIDFETLKPSTLRELERYVTSCLR--KKRKPQAEK 686
Score = 38.1 bits (87), Expect = 0.11
Identities = 52/247 (21%), Positives = 88/247 (35%), Gaps = 26/247 (10%)
Query: 249 QPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTEDVN 308
+PQA + S + K S ++ S + D + ++ ++ ++ + H D
Sbjct: 681 KPQAEKVDVIAGSSKMKGFSSSESESTSESSSSDSEDSETEMAPKSKKKGHTG---RDQK 737
Query: 309 SPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPELNSKLEDMTSQQQD 368
+ Q +P + P P + + + P P + T+
Sbjct: 738 KHHHHHHPQMQ-PAPAPVPQQPPPPPQQPPPPPPPQQQQQQPPPPPPPPSMPQQTAPAMK 796
Query: 369 NSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQP-----LSQSV--SDDLHINQQ 421
+S + P L +L GS P+ + T L+L QP L Q S H+NQ
Sbjct: 797 SSPPPFITAQVPVLEPQLPGSVFDPIGHFTQPILHLPQPELPPHLPQPPEHSTPPHLNQH 856
Query: 422 AEPSNLNVRQE-DDRPSSPIHRQGAISDNLHSHQQVEPS--------------NPNVLLE 466
A S + +PS P +R A+ V P+ P VLLE
Sbjct: 857 AVVSPPALHNALPQQPSRPSNRAAALPPKPTRPPAVSPALAQPPLLPQPPMVQPPQVLLE 916
Query: 467 DDGPSSP 473
D+ P +P
Sbjct: 917 DEEPPAP 923
>BDF1_YEAST (P35817) BDF1 protein
Length = 686
Score = 100 bits (249), Expect = 2e-20
Identities = 80/287 (27%), Positives = 131/287 (44%), Gaps = 43/287 (14%)
Query: 649 LQMGSKFFKSCSSLLEKLMEHQYA---WVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTR 705
LQ KF C S+L++LM ++A + F PVD + L YF + PMDLGT+ +
Sbjct: 314 LQQAMKF---CQSVLKELMAKKHASYNYPFLEPVDPVSMNLPTYFDYVKEPMDLGTIAKK 370
Query: 706 LNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWA----IIESDYN 761
LN Y++ ++F DVRL F N T+NP G V+ M +L ++F +WA + + D +
Sbjct: 371 LNDWQYQTMEDFERDVRLVFKNCYTFNPDGTIVNMMGHRLEEVFNSKWADRPNLDDYDSD 430
Query: 762 REMRYGMDYG------APSPLSRRVPAFTPPPPLDMRRILDRQE---------------- 799
+ R DY + S + + T P + L R +
Sbjct: 431 EDSRTQGDYDDYESEYSESDIDETI--ITNPAIQYLEEQLARMKVELQQLKKQELEKIRK 488
Query: 800 -------PFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKL 852
R RS + S + + K+ K +TYD K+ ++ ++ LP KL
Sbjct: 489 ERRLARGSKKRGKRSKGRSGSKNASSKGRRDKKNKLKTVVTYDMKRIITERINDLPTSKL 548
Query: 853 DAVVQMMRKKNL*LNQQDDEIEVDFDAVDAE-ILWELDRFVLNYKKS 898
+ + ++ KK++ +DDE+E+D D +D IL + F Y+ S
Sbjct: 549 ERAIDII-KKSMPNISEDDEVELDLDTLDNHTILTLYNTFFRQYESS 594
Score = 52.8 bits (125), Expect = 4e-06
Identities = 49/191 (25%), Positives = 72/191 (37%), Gaps = 32/191 (16%)
Query: 675 FNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPK 734
F PVD L + YF I PMDL T++ +LN Y+ P++ ED L +N++ +N
Sbjct: 173 FLQPVDPVKLDIPFYFNYIKRPMDLSTIERKLNVGAYEVPEQITEDFNLMVNNSIKFNGP 232
Query: 735 GQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRI 794
+ MA + FE +PA PP + R
Sbjct: 233 NAGISQMARNIQASFEKHML------------------------NMPAKDAPPVIAKGRR 268
Query: 795 LDRQE--PFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKL 852
QE P +N RT P PK+KD Y+ K+ S L
Sbjct: 269 SSAQEDAPIVIRRAQTHNGRPKRTIHP--PKSKD----IYPYESKKPKSKRLQQAMKFCQ 322
Query: 853 DAVVQMMRKKN 863
+ ++M KK+
Sbjct: 323 SVLKELMAKKH 333
>BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3
protein) (O27.1.1)
Length = 801
Score = 99.4 bits (246), Expect = 4e-20
Identities = 50/105 (47%), Positives = 65/105 (61%), Gaps = 3/105 (2%)
Query: 653 SKFFKSCSSLLEKLMEHQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKN 709
S+ K C+ +L++L+ + YAW F PVD LGLHDY II +PMDL TVK ++
Sbjct: 347 SEQLKHCNGILKELLSKKHAAYAWPFYKPVDASALGLHDYHDIIKHPMDLSTVKRKMENR 406
Query: 710 WYKSPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWA 754
Y+ +EFA DVRL F N YNP DV AMA +L +FE R+A
Sbjct: 407 DYRDAQEFAADVRLMFSNCYKYNPPDHDVVAMARKLQDVFEFRYA 451
Score = 81.6 bits (200), Expect = 9e-15
Identities = 39/88 (44%), Positives = 51/88 (57%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+++ L +HQ+AW F PVD LGL DY II PMD+GT+K RL N+Y + E +D
Sbjct: 86 VMKALWKHQFAWPFRQPVDAVKLGLPDYHKIIKQPMDMGTIKRRLENNYYWAASECMQDF 145
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
F N YN D+ MA+ L KIF
Sbjct: 146 NTMFTNCYIYNKPTDDIVLMAQTLEKIF 173
Score = 57.0 bits (136), Expect = 2e-07
Identities = 39/123 (31%), Positives = 63/123 (50%), Gaps = 14/123 (11%)
Query: 813 SSRTPAPKKPKAKDPNK----RDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQ 868
+++T P P D + R M+YDEK++LS ++ LP EKL VV +++ + L
Sbjct: 619 ATKTAPPALPTGYDSEEEEESRPMSYDEKRQLSLDINKLPGEKLGRVVHIIQAREPSLRD 678
Query: 869 QD-DEIEVDFDAVDAEILWELDRFVLN---------YKKSLSKNKRKAEQARERAEALQN 918
+ +EIE+DF+ + L EL+R+VL+ Y K K E A E+ L+
Sbjct: 679 SNPEEIEIDFETLKPSTLRELERYVLSCLRKKPRKPYTIKKPVGKTKEELALEKKRELEK 738
Query: 919 SVQ 921
+Q
Sbjct: 739 RLQ 741
>YK82_SCHPO (Q9HGP4) Hypothetical bromodomain protein C631.02
Length = 727
Score = 96.3 bits (238), Expect = 4e-19
Identities = 74/287 (25%), Positives = 133/287 (45%), Gaps = 24/287 (8%)
Query: 657 KSCSSLLEKLMEHQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKS 713
K C S+L++L++ Q YA+ F PV+ G DYF +I +PMDLGT++ +LN N Y S
Sbjct: 395 KFCQSVLKELLKKQHEAYAYPFYKPVNPTACGCPDYFKVIKHPMDLGTMQNKLNHNEYAS 454
Query: 714 PKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAP 773
K F D+ L F N +N G VH M ++L IF+ WA + D++ E GM
Sbjct: 455 MKAFEADMVLMFKNCYKFNSAGTPVHLMGKKLESIFQKLWA-NKPDFDSETYMGMSSVNT 513
Query: 774 SPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRD-- 831
F + D E F R ++ S+ + ++ ++R
Sbjct: 514 DYYYGDNEVFDSGDEF----LEDDGEEFEAVNRQIHKLQSTLQAMKSRARSSSVSRRSRS 569
Query: 832 ----------MTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVD 881
+TY+ + +L+ + L ++L V +++R L + DEIE+D A+
Sbjct: 570 RSLSVDIYPPITYEMQNELAEQCNYLSADQLSHVAEILRAA-LPHLRNTDEIEIDVSAMP 628
Query: 882 AEILWELDRFVLNYKKSLSKNKRKA---EQARERAEALQNSVQSSRL 925
++ +++ +V + ++ A Q +++ AL + Q+ ++
Sbjct: 629 PDVFYKVYYYVCKGDEIGAEALATASHTHQEKKKGRALSETEQAEKI 675
Score = 62.4 bits (150), Expect = 6e-09
Identities = 69/324 (21%), Positives = 118/324 (36%), Gaps = 63/324 (19%)
Query: 442 RQGAISDNLHSHQQVEPSNPNVLLEDDGPSSPIH---RHKAVPSTEYRHSENVTEEPNLE 498
++G + +S Q + P+ P +D+ PSSP H + V +HS + EE
Sbjct: 74 QKGDNQETDYSSQYLHPTPPYTNFDDESPSSPTHPSVSNITVDGDSKKHSLQLQEEEKSS 133
Query: 499 DRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRIEVKQGYVGGYGN--SNVVLGGGIT 556
+ + + + ++ L S V +K G N SN +
Sbjct: 134 ESLDSHTHPPKRVRNEDDSLTFSKT-----SPVSPSSLKDGASNTVTNDASNKIKSEASE 188
Query: 557 NGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQNSRRASEGVEKEKRMPKANQFYHNSE 616
+ + ++ + G S++ + P E V+KE+
Sbjct: 189 SASPSALQALDSTAAGSSKEHSSP--------------HDETVKKEEN------------ 222
Query: 617 FLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWVFN 676
D++PP M + K ++L +L + + F
Sbjct: 223 ---DKDQYPP------------------------MTKEQHKYIHAMLRQLRRGRDSIPFR 255
Query: 677 TPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPKGQ 736
PVD + DY TII NP+DLGT++ + + Y S + F +D+ L F N YN
Sbjct: 256 APVDPVKQNIPDYPTIIKNPIDLGTMQKKFSSGVYSSAQHFIDDMNLMFSNCFLYNGTES 315
Query: 737 DVHAMAEQLSKIFEDRWAIIESDY 760
V M + L FE + + S Y
Sbjct: 316 PVGVMGKNLQATFERQLKQLPSAY 339
>BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3
(Bromodomain-containing FSH-like protein FSRG2)
Length = 726
Score = 91.7 bits (226), Expect = 9e-18
Identities = 54/149 (36%), Positives = 80/149 (53%), Gaps = 19/149 (12%)
Query: 625 PPAESNKKSKLNWKKQGGG--------DMGXGL------QMG--SKFFKSCSSLLEKLME 668
PP K++K+ +++ GG D+ G + G S+ + C S+L +++
Sbjct: 264 PPLSEPKQAKVVARRESGGRPIKPPKKDLEDGEVPQHAGKKGKLSEHLRHCDSILREMLS 323
Query: 669 HQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTF 725
+ YAW F PVD + L LHDY II +PMDL TVK +++ Y + FA D+RL F
Sbjct: 324 KKHAAYAWPFYKPVDAEALELHDYHDIIKHPMDLSTVKRKMDSREYPDAQGFAADIRLMF 383
Query: 726 HNAMTYNPKGQDVHAMAEQLSKIFEDRWA 754
N YNP +V AMA +L +FE R+A
Sbjct: 384 SNCYKYNPPDHEVVAMARKLQDVFEMRFA 412
Score = 81.6 bits (200), Expect = 9e-15
Identities = 40/88 (45%), Positives = 51/88 (57%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+++ L +HQ+AW F PVD L L DY II NPMD+GT+K RL N+Y S E +D
Sbjct: 45 VVKTLWKHQFAWPFYQPVDAIKLNLPDYHKIIKNPMDMGTIKKRLENNYYWSASECMQDF 104
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
F N YN D+ MA+ L KIF
Sbjct: 105 NTMFTNCYIYNKPTDDIVLMAQALEKIF 132
Score = 54.7 bits (130), Expect = 1e-06
Identities = 44/143 (30%), Positives = 76/143 (52%), Gaps = 19/143 (13%)
Query: 798 QEPFARTPRSMNNTPSSRT--PAPKKPKAKDPNKRD-----MTYDEKQKLSTSLHSLPVE 850
Q+ A T ++ + T +SR K+ A ++ + M+YDEK++LS ++ LP E
Sbjct: 532 QQKKAPTKKANSTTTASRQLKKGGKQASASYDSEEEEEGLPMSYDEKRQLSLDINRLPGE 591
Query: 851 KLDAVVQMMRKKNL*LNQQD-DEIEVDFDAVDAEILWELDRFVLN-----YKKSLSKN-- 902
KL VV +++ + L + DEIE+DF+ + L EL+R+V + +K LS +
Sbjct: 592 KLGRVVHIIQSREPSLRDSNPDEIEIDFETLKPTTLRELERYVKSCLQKKQRKPLSTSGK 651
Query: 903 ----KRKAEQARERAEALQNSVQ 921
K K E A+E+ + L+ +Q
Sbjct: 652 KQAAKSKEELAQEKKKELEKRLQ 674
>BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like
protein)
Length = 726
Score = 90.9 bits (224), Expect = 1e-17
Identities = 55/149 (36%), Positives = 80/149 (52%), Gaps = 19/149 (12%)
Query: 625 PPAESNKKSKLNWKKQGGG--------DMGXGL------QMG--SKFFKSCSSLLEKLME 668
PP K++K+ +++ GG D+ G + G S+ + C S+L +++
Sbjct: 265 PPLSDPKQAKVVARRESGGRPIKPPKKDLEDGEVPQHAGKKGKLSEHLRYCDSILREMLS 324
Query: 669 HQ---YAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTF 725
+ YAW F PVD + L LHDY II +PMDL TVK +++ Y + FA DVRL F
Sbjct: 325 KKHAAYAWPFYKPVDAEALELHDYHDIIKHPMDLSTVKRKMDGREYPDAQGFAADVRLMF 384
Query: 726 HNAMTYNPKGQDVHAMAEQLSKIFEDRWA 754
N YNP +V AMA +L +FE R+A
Sbjct: 385 SNCYKYNPPDHEVVAMARKLQDVFEMRFA 413
Score = 81.6 bits (200), Expect = 9e-15
Identities = 40/88 (45%), Positives = 51/88 (57%)
Query: 662 LLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDV 721
+++ L +HQ+AW F PVD L L DY II NPMD+GT+K RL N+Y S E +D
Sbjct: 46 VVKTLWKHQFAWPFYQPVDAIKLNLPDYHKIIKNPMDMGTIKKRLENNYYWSASECMQDF 105
Query: 722 RLTFHNAMTYNPKGQDVHAMAEQLSKIF 749
F N YN D+ MA+ L KIF
Sbjct: 106 NTMFTNCYIYNKPTDDIVLMAQALEKIF 133
Score = 55.5 bits (132), Expect = 7e-07
Identities = 33/88 (37%), Positives = 53/88 (59%), Gaps = 4/88 (4%)
Query: 832 MTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQD-DEIEVDFDAVDAEILWELDR 890
M+YDEK++LS ++ LP EKL VV +++ + L + DEIE+DF+ + L EL+R
Sbjct: 572 MSYDEKRQLSLDINRLPGEKLGRVVHIIQSREPSLRDSNPDEIEIDFETLKPTTLRELER 631
Query: 891 FVLNYKKSLSKNKRKAEQARERAEALQN 918
+V K L K +RK A + +A ++
Sbjct: 632 YV---KSCLQKKQRKPFSASGKKQAAKS 656
>CBP_MOUSE (P45481) CREB-binding protein (EC 2.3.1.48)
Length = 2441
Score = 75.5 bits (184), Expect = 6e-13
Identities = 34/76 (44%), Positives = 49/76 (63%)
Query: 675 FNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPK 734
F PVD LG+ DYF I+ NPMDL T+K +L+ Y+ P ++ +DVRL F+NA YN K
Sbjct: 1112 FRQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVRLMFNNAWLYNRK 1171
Query: 735 GQDVHAMAEQLSKIFE 750
V+ +L+++FE
Sbjct: 1172 TSRVYKFCSKLAEVFE 1187
>WDR9_HUMAN (Q9NSI6) WD-repeat protein 9
Length = 2269
Score = 72.8 bits (177), Expect = 4e-12
Identities = 41/109 (37%), Positives = 61/109 (55%), Gaps = 3/109 (2%)
Query: 653 SKFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYK 712
S + K C L+ + + + + F PVD+ + DY II PMD GTV+ L+ Y
Sbjct: 1316 SNWKKQCKELVNLIFQCEDSEPFRQPVDL--VEYPDYRDIIDTPMDFGTVRETLDAGNYD 1373
Query: 713 SPKEFAEDVRLTFHNAMTYNP-KGQDVHAMAEQLSKIFEDRWAIIESDY 760
SP EF +D+RL F NA Y P K +++M +LS +FE++ I SD+
Sbjct: 1374 SPLEFCKDIRLIFSNAKAYTPNKRSKIYSMTLRLSALFEEKMKKISSDF 1422
Score = 39.3 bits (90), Expect = 0.051
Identities = 23/84 (27%), Positives = 41/84 (48%), Gaps = 2/84 (2%)
Query: 663 LEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVR 722
+++L+ A F PVD+ Y T++ P DL T++ RL +Y+ +VR
Sbjct: 1173 IDQLLNLDIAAAFAGPVDL--CTYPKYCTVVAYPTDLYTIRMRLVNRFYRRLSALVWEVR 1230
Query: 723 LTFHNAMTYNPKGQDVHAMAEQLS 746
HNA T+N + A++++
Sbjct: 1231 YIEHNARTFNEPESVIARSAKKIT 1254
>CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48)
Length = 2442
Score = 72.4 bits (176), Expect = 5e-12
Identities = 33/76 (43%), Positives = 48/76 (62%)
Query: 675 FNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPK 734
F PVD LG+ DYF I+ NPMDL T+K +L+ Y+ P ++ +DV L F+NA YN K
Sbjct: 1111 FRQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVWLMFNNAWLYNRK 1170
Query: 735 GQDVHAMAEQLSKIFE 750
V+ +L+++FE
Sbjct: 1171 TSRVYKFCSKLAEVFE 1186
Score = 35.0 bits (79), Expect = 0.97
Identities = 57/269 (21%), Positives = 95/269 (35%), Gaps = 28/269 (10%)
Query: 116 NSPQQQLEELNSAQPHASFQLADGNSPQQK-VENQDSAHPQASLRIGDGNSPQQQLVEPN 174
+SPQQQ + LN + + A K V NQ PQ L+ G PQ + +
Sbjct: 2078 SSPQQQQQVLNILKSNPQLMAAFIKQRTAKYVANQPGMQPQPGLQSQPGMQPQPGMHQQP 2137
Query: 175 LAQPQASLRAG---DGNSPQRQLEEENSAQPQASSRTGDGNLPH---------------- 215
Q +++AG G PQ+Q + Q QA + G+ P+
Sbjct: 2138 SLQNLNAMQAGVPRPGVPPQQQAMGGLNPQGQALNIMNPGHNPNMASMNPQYREMLRRQL 2197
Query: 216 -QQLEDQYSAQPPASLRIGDGNSLRQKVEGH-NLVQPQASSIIEDGNSPRQKTEDQSLAQ 273
QQ + Q Q + + + GH QPQ G P + Q + Q
Sbjct: 2198 LQQQQQQQQQQQQQQQQQQGSAGMAGGMAGHGQFQQPQG-----PGGYPPAMQQQQRMQQ 2252
Query: 274 PPVSSRTDDGDSAQQQVEDQNLAQTHV-SSRTEDVNSPRPQNSTQKDGDSPQILENSRPE 332
+ G A Q + + Q + + T ++ Q Q+ QI +P
Sbjct: 2253 HLPLQGSSMGQMAAQMGQLGQMGQPGLGADSTPNIQQALQQRILQQQQMKQQIGSPGQPN 2312
Query: 333 DASLTQPEVNSRLEDRSFPQPELNSKLED 361
S Q ++ + + P ++ + L +
Sbjct: 2313 PMSPQQHMLSGQPQASHLPGQQIATSLSN 2341
Score = 32.7 bits (73), Expect = 4.8
Identities = 42/204 (20%), Positives = 67/204 (32%), Gaps = 25/204 (12%)
Query: 135 QLADGNSPQQKVENQDSAHPQASLRIGDGNSPQQQLVEPNLAQPQASLRAGDGNSPQRQL 194
QL QQ+ + Q Q S + G + Q +P G G P
Sbjct: 2196 QLLQQQQQQQQQQQQQQQQQQGSAGMAGGMAGHGQFQQPQ----------GPGGYPPAMQ 2245
Query: 195 EEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLRQKVEGHNLVQPQASS 254
+++ Q + G + Q + QP L +++Q ++ L Q Q
Sbjct: 2246 QQQRMQQHLPLQGSSMGQMAAQMGQLGQMGQP--GLGADSTPNIQQALQQRILQQQQMKQ 2303
Query: 255 IIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLAQTHVSSRTED---VNSPR 311
I P + Q + G + Q +A T +S++ V SPR
Sbjct: 2304 QIGSPGQPNPMSPQQHMLS---------GQPQASHLPGQQIA-TSLSNQVRSPAPVQSPR 2353
Query: 312 PQNSTQKDGDSPQILENSRPEDAS 335
PQ+ SP+I P S
Sbjct: 2354 PQSQPPHSSPSPRIQPQPSPHHVS 2377
>P300_HUMAN (Q09472) E1A-associated protein p300 (EC 2.3.1.48)
Length = 2414
Score = 70.9 bits (172), Expect = 2e-11
Identities = 32/76 (42%), Positives = 48/76 (63%)
Query: 675 FNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPK 734
F PVD LG+ DYF I+ +PMDL T+K +L+ Y+ P ++ +D+ L F+NA YN K
Sbjct: 1075 FRQPVDPQLLGIPDYFDIVKSPMDLSTIKRKLDTGQYQEPWQYVDDIWLMFNNAWLYNRK 1134
Query: 735 GQDVHAMAEQLSKIFE 750
V+ +LS++FE
Sbjct: 1135 TSRVYKYCSKLSEVFE 1150
Score = 37.7 bits (86), Expect = 0.15
Identities = 95/392 (24%), Positives = 141/392 (35%), Gaps = 86/392 (21%)
Query: 84 GSSRLEDGSSIQQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADG---- 139
G S L+ G+ QQ L QNL+ S +SP QQ + L+ HA+ QL
Sbjct: 2037 GISPLKPGTVSQQAL--QNLLRTLRSP-----SSPLQQQQVLSIL--HANPQLLAAFIKQ 2087
Query: 140 ------NSPQQKVENQDSAHPQASLRIGDGNSPQQQLVEPNLAQP-----QASL-RAG-D 186
NS Q + Q PQ + P QQ V N A QA + RAG
Sbjct: 2088 RAAKYANSNPQPIPGQPGM-PQGQPGLQPPTMPGQQGVHSNPAMQNMNPMQAGVQRAGLP 2146
Query: 187 GNSPQRQLEEE-NSAQPQASSRTGDGNLPHQQLED--------QYSAQPPASLRIGDG-- 235
PQ+QL+ PQA + N Q D Q Q A IG G
Sbjct: 2147 QQQPQQQLQPPMGGMSPQAQQMNMNHNTMPSQFRDILRRQQMMQQQQQQGAGPGIGPGMA 2206
Query: 236 -NSLRQKVEG------------HNLVQPQASSIIEDGNSPRQ---------KTEDQSLAQ 273
++ Q+ +G H++ Q Q ++ + G P+ + Q L Q
Sbjct: 2207 NHNQFQQPQGVGYPPQPQQRMQHHMQQMQQGNMGQIGQLPQALGAEAGASLQAYQQRLLQ 2266
Query: 274 PPVSSRTDDGDSAQQQVEDQNLAQT-HVSSRT------------EDVNSPRPQNSTQKDG 320
+ S + QQ N AQ+ H+ + + V SPRPQ+
Sbjct: 2267 QQMGSPVQPNPMSPQQHMLPNQAQSPHLQGQQIPNSLSNQVRSPQPVPSPRPQSQPPHSS 2326
Query: 321 DSPQILENSRPEDAS---------LTQPEVNSRLEDRSFPQPELNSKLEDMTSQQQDNSI 371
SP++ P S L + N +E F P+ NS L + S + +
Sbjct: 2327 PSPRMQPQPSPHHVSPQTSSPHPGLVAAQANP-MEQGHFASPDQNSMLSQLAS---NPGM 2382
Query: 372 LEDGNSSQPQLNLRLEGSSLQPVANSTSEDLN 403
+S L L + S L + ++ D++
Sbjct: 2383 ANLHGASATDLGLSTDNSDLNSNLSQSTLDIH 2414
>CSP_PLAKH (P02894) Circumsporozoite protein precursor (CS)
Length = 363
Score = 68.2 bits (165), Expect = 1e-10
Identities = 60/223 (26%), Positives = 88/223 (38%), Gaps = 23/223 (10%)
Query: 56 NNNGTTLENID-----------EAPEFAVLGDGNLAKAEGSSRLEDGSSIQQQLGDQNLV 104
N NG + N+D A LG+ A+ + E G +++ N
Sbjct: 33 NVNGVSFNNVDTSSLGAQQVRQSASRGRGLGEKPKEGADKEKKKEKGKEKEEEPKKPN-- 90
Query: 105 EEHASSRTGDRNSPQQQLEELNSAQPHASFQLADGNSPQQKVENQDSAHPQASLRIGDGN 164
+ + PQ Q + N+ QP A A+ PQ + + ++ PQA +
Sbjct: 91 --ENKLKQPNEGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAG 148
Query: 165 SPQQQLVEPNLAQPQASLRAGDGNSPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSA 224
PQ Q N QPQA + PQ Q + N+ QPQA + P Q + +
Sbjct: 149 QPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAG 208
Query: 225 QPPASLRIGDGNSLRQ---KVEGHNLVQPQASSIIEDGNSPRQ 264
QP A GDG + Q + +G N QPQA + N PRQ
Sbjct: 209 QPQAQ---GDGANAGQPQAQGDGANAGQPQAQG--DGANVPRQ 246
Score = 56.2 bits (134), Expect = 4e-07
Identities = 42/157 (26%), Positives = 69/157 (43%), Gaps = 10/157 (6%)
Query: 167 QQQLVEPNLAQPQASLRAGDGNSPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQP 226
+ +L +PN QPQA + PQ Q + N+ QPQA + P Q + + QP
Sbjct: 91 ENKLKQPNEGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQP 150
Query: 227 PASLRIGDGNSLRQ---KVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDG 283
A GDG + Q + +G N QPQA + P+ + + + QP ++ D
Sbjct: 151 QAQ---GDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQP--QAQGDGA 205
Query: 284 DSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDG 320
++ Q Q + ++ + N+ +PQ Q DG
Sbjct: 206 NAGQPQAQGDGANAGQPQAQGDGANAGQPQ--AQGDG 240
Score = 52.4 bits (124), Expect = 6e-06
Identities = 47/163 (28%), Positives = 63/163 (37%), Gaps = 17/163 (10%)
Query: 95 QQQLGDQNLVEEHASSRTGDRNSPQQQLEELNSAQPHASFQLADGNSPQQKVENQDSAHP 154
Q Q N + A + PQ Q + N+ QP A A+ PQ + + ++ P
Sbjct: 115 QAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQP 174
Query: 155 QASLRIGDGNSPQQQLVE---PNLAQPQASLRAGDGNSPQRQLEEENSAQPQASSRTGDG 211
QA GDG + Q + N QPQA + PQ Q + N+ QPQA GDG
Sbjct: 175 QAQ---GDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQ---GDG 228
Query: 212 NLPHQQLEDQYSAQPPASLRIG--------DGNSLRQKVEGHN 246
Q A P R G +GN K +G N
Sbjct: 229 ANAGQPQAQGDGANVPRQGRNGGGAPAGGNEGNKQAGKGQGQN 271
Score = 34.7 bits (78), Expect = 1.3
Identities = 37/178 (20%), Positives = 62/178 (34%), Gaps = 11/178 (6%)
Query: 180 ASLRAGDGNSPQRQLEEENSAQPQASSRTGDGNLPHQQLEDQYSAQPPASLRIGDGNSLR 239
AS G G P+ ++E + +L+ QP A GDG +
Sbjct: 56 ASRGRGLGEKPKEGADKEKKKEKGKEKEEEPKKPNENKLKQPNEGQPQAQ---GDGANAG 112
Query: 240 Q---KVEGHNLVQPQASSIIEDGNSPRQKTEDQSLAQPPVSSRTDDGDSAQQQVEDQNLA 296
Q + +G N QPQA + P+ + + + QP + Q Q + N
Sbjct: 113 QPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAGQPQAQGDGANAG 172
Query: 297 QTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSFPQPE 354
Q + P+ Q G PQ ++ + A+ QP+ + PQ +
Sbjct: 173 QPQAQGDGANAGQPQAQGDGANAG-QPQ----AQGDGANAGQPQAQGDGANAGQPQAQ 225
>BA2B_CHICK (Q9DE13) Bromodomain adjacent to zinc finger domain 2B
(Extracellular matrix protein F22)
Length = 2130
Score = 66.2 bits (160), Expect = 4e-10
Identities = 35/104 (33%), Positives = 52/104 (49%), Gaps = 2/104 (1%)
Query: 653 SKFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYK 712
SK CS +L +L H+ AW F PV++ + Y +I PMD T++ +L Y
Sbjct: 2025 SKDLAICSMILSELETHEDAWPFLLPVNLKLVP--GYKKVIKKPMDFSTIRDKLTSGQYP 2082
Query: 713 SPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAII 756
+ + F+ DVRL F N T+N D+ + K FE +W I
Sbjct: 2083 NVEAFSLDVRLVFDNCETFNEDDSDIGRAGHNMRKYFEKKWTEI 2126
>BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2B
(hWALp4)
Length = 1972
Score = 65.9 bits (159), Expect = 5e-10
Identities = 34/101 (33%), Positives = 52/101 (50%), Gaps = 2/101 (1%)
Query: 653 SKFFKSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYK 712
SK CS +L ++ H+ AW F PV++ + Y +I PMD T++ +L+ Y
Sbjct: 1867 SKDLALCSMILTEMETHEDAWPFLLPVNLKLVP--GYKKVIKKPMDFSTIREKLSSGQYP 1924
Query: 713 SPKEFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRW 753
+ + FA DVRL F N T+N D+ + K FE +W
Sbjct: 1925 NLETFALDVRLVFDNCETFNEDDSDIGRAGHNMRKYFEKKW 1965
Score = 34.3 bits (77), Expect = 1.6
Identities = 50/251 (19%), Positives = 98/251 (38%), Gaps = 11/251 (4%)
Query: 356 NSKLEDMTSQQQDNSILEDGNSSQPQLNLRLEGSSLQPVANSTSEDLNLAQPLSQSVSDD 415
N K S+ Q D + +E SS + TS D + ++ +S S SDD
Sbjct: 13 NRKCNQEQSKNQPLDARVDKIKDKKPRKKAMESSSNSDSDSGTSSDTS-SEGISSSDSDD 71
Query: 416 LHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDNLHSHQQVEPSNPNVLLEDDGPSSPIH 475
L +++ E ++ EDD S Q ++ + H +P + IH
Sbjct: 72 LEEDEEEEDQSIE-ESEDDDSDSESEAQHKSNNQVLLHGISDPKADGQKATEKAQEKRIH 130
Query: 476 RHKAVPSTEYRHS-ENVTEEPNLEDRIKINLASTSKQEKQEMCLKLESELDAVRSLVKRI 534
+ + S HS ++ ++P + + ++ S Q K+E K S + + LV +
Sbjct: 131 QPLPLASESQTHSFQSQQKQPQVLSQ-QLPFIFQSSQAKEESVNKHTSVIQST-GLVSNV 188
Query: 535 E----VKQGYVGGYGNSNVVLGGGITNGGGAQRAHSEAGSVGVSRQPTRPLHQLSFPMFQ 590
+ V Q Y +++ G + E+ S+ + R ++ +FP
Sbjct: 189 KPLSLVNQAKKETY--MKLIVPSPDVLKAGNKNTSEESSSLTSELRSKREQYKQAFPSQL 246
Query: 591 NSRRASEGVEK 601
+ +S+ ++K
Sbjct: 247 KKQESSKSLKK 257
>BA2A_MOUSE (Q91YE5) Bromodomain adjacent to zinc finger domain 2A
(Transcription termination factor-I interacting protein
5) (TTF-I interacting protein 5) (Tip5)
Length = 1850
Score = 64.3 bits (155), Expect = 1e-09
Identities = 35/98 (35%), Positives = 48/98 (48%), Gaps = 8/98 (8%)
Query: 659 CSSLLEKLMEHQYAWVFNTPVD---VDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPK 715
C +L ++ H AW F PV+ V G Y +I NPMD T++ RL + Y S +
Sbjct: 1747 CEIILMEMESHDAAWPFLEPVNPRLVSG-----YRRVIKNPMDFSTMRERLLRGGYTSSE 1801
Query: 716 EFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRW 753
EFA D L F N T+N +V + + FE RW
Sbjct: 1802 EFAADALLVFDNCQTFNEDDSEVGKAGHVMRRFFESRW 1839
>BA2A_HUMAN (Q9UIF9) Bromodomain adjacent to zinc finger domain 2A
(Transcription termination factor-I interacting protein
5) (TTF-I interacting protein 5) (Tip5) (hWALp3)
Length = 1878
Score = 64.3 bits (155), Expect = 1e-09
Identities = 36/98 (36%), Positives = 48/98 (48%), Gaps = 8/98 (8%)
Query: 659 CSSLLEKLMEHQYAWVFNTPVD---VDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPK 715
C +L ++ H AW F PV+ V G Y II NPMD T++ RL + Y S +
Sbjct: 1775 CEIILMEMESHDAAWPFLEPVNPRLVSG-----YRRIIKNPMDFSTMRERLLRGGYTSSE 1829
Query: 716 EFAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRW 753
EFA D L F N T+N +V + + FE RW
Sbjct: 1830 EFAADALLVFDNCQTFNEDDSEVGKAGHIMRRFFESRW 1867
>BA1B_HUMAN (Q9UIG0) Bromodomain adjacent to zinc finger domain
protein 1B (Williams-Beuren syndrome chromosome region 9
protein) (WBRS9) (Williams syndrome transcription factor)
(hWALP2)
Length = 1483
Score = 63.9 bits (154), Expect = 2e-09
Identities = 29/82 (35%), Positives = 49/82 (59%), Gaps = 2/82 (2%)
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
+ C +L K+++++++W F PV D DY+ +IT+PMD TV+ + + Y+S +E
Sbjct: 1346 QKCEEILHKIVKYRFSWPFREPVTRDEA--EDYYDVITHPMDFQTVQNKCSCGSYRSVQE 1403
Query: 717 FAEDVRLTFHNAMTYNPKGQDV 738
F D++ F NA YN +G V
Sbjct: 1404 FLTDMKQVFTNAEVYNCRGSHV 1425
>GCN5_YEAST (Q03330) Histone acetyltransferase GCN5 (EC 2.3.1.48)
Length = 439
Score = 63.5 bits (153), Expect = 3e-09
Identities = 31/92 (33%), Positives = 50/92 (53%), Gaps = 2/92 (2%)
Query: 661 SLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAED 720
++L +L H AW F PV+ + + DY+ I PMDL T++ +L N Y+ ++F D
Sbjct: 338 NILTELQNHAAAWPFLQPVNKEEVP--DYYDFIKEPMDLSTMEIKLESNKYQKMEDFIYD 395
Query: 721 VRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDR 752
RL F+N YN + + A +L K F ++
Sbjct: 396 ARLVFNNCRMYNGENTSYYKYANRLEKFFNNK 427
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 118,340,385
Number of Sequences: 164201
Number of extensions: 5542996
Number of successful extensions: 17402
Number of sequences better than 10.0: 448
Number of HSP's better than 10.0 without gapping: 73
Number of HSP's successfully gapped in prelim test: 387
Number of HSP's that attempted gapping in prelim test: 15783
Number of HSP's gapped (non-prelim): 1297
length of query: 966
length of database: 59,974,054
effective HSP length: 120
effective length of query: 846
effective length of database: 40,269,934
effective search space: 34068364164
effective search space used: 34068364164
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC144893.5


