

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.11 + phase: 0 /pseudo/partial
(652 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
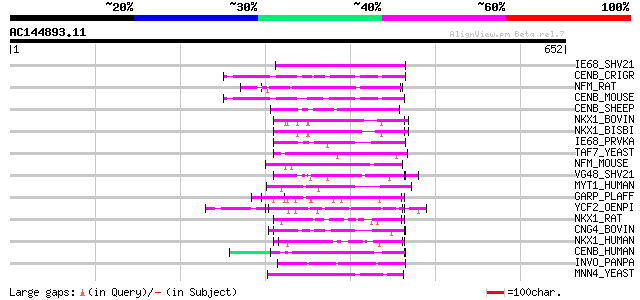
Score E
Sequences producing significant alignments: (bits) Value
IE68_SHV21 (Q01042) Immediate-early protein 65 7e-10
CENB_CRIGR (P48988) Major centromere autoantigen B (Centromere p... 64 1e-09
NFM_RAT (P12839) Neurofilament triplet M protein (160 kDa neurof... 63 2e-09
CENB_MOUSE (P27790) Major centromere autoantigen B (Centromere p... 63 2e-09
CENB_SHEEP (P49451) Major centromere autoantigen B (Centromere p... 61 1e-08
NKX1_BOVIN (Q28139) Sodium/potassium/calcium exchanger 1 precurs... 60 2e-08
NKX1_BISBI (O46383) Sodium/potassium/calcium exchanger 1 (Na(+)/... 60 2e-08
IE68_PRVKA (P24827) Immediate-early protein RSP40 59 3e-08
TAF7_YEAST (Q05021) Transcription initiation factor TFIID subuni... 59 4e-08
NFM_MOUSE (P08553) Neurofilament triplet M protein (160 kDa neur... 58 9e-08
VG48_SHV21 (Q01033) Hypothetical gene 48 protein 56 3e-07
MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI) ... 56 3e-07
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 56 3e-07
YCF2_OENPI (P31568) Protein ycf2 (Fragment) 56 3e-07
NKX1_RAT (Q9QZM6) Sodium/potassium/calcium exchanger 1 precursor... 56 3e-07
CNG4_BOVIN (Q28181) 240 kDa protein of rod photoreceptor CNG-cha... 55 6e-07
NKX1_HUMAN (O60721) Sodium/potassium/calcium exchanger 1 precurs... 54 1e-06
CENB_HUMAN (P07199) Major centromere autoantigen B (Centromere p... 54 2e-06
INVO_PANPA (P14591) Involucrin 53 2e-06
MNN4_YEAST (P36044) MNN4 protein 53 3e-06
>IE68_SHV21 (Q01042) Immediate-early protein
Length = 407
Score = 64.7 bits (156), Expect = 7e-10
Identities = 41/154 (26%), Positives = 74/154 (47%), Gaps = 1/154 (0%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
EGEG E ++E EE EA++EE EE ++E + + EE + +E E E
Sbjct: 89 EGEGREEAEEEEAEEKEAEEEEAEEAEEEAEEEEAEEAEAEEEEAEEEEAEEEEAEEAEE 148
Query: 373 SLGQDKVETLPVEKEQGEGEQMMD-FEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKG 431
++ E E+ + E E+ + E++++E E ++ E + + +
Sbjct: 149 EEAEEAEEEAEEEEAEEEAEEEAEEAEEAEEEAEEEAEEAEEAEEAEEAEEEAEEAEEEA 208
Query: 432 IDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ E+ E+ E EE E+ EEE ++ EE+E + G
Sbjct: 209 EEAEEEAEEAEEAEEAEEAEEEAEEAEEEEEEAG 242
>CENB_CRIGR (P48988) Major centromere autoantigen B (Centromere
protein B) (CENP-B)
Length = 606
Score = 64.3 bits (155), Expect = 1e-09
Identities = 62/215 (28%), Positives = 101/215 (46%), Gaps = 30/215 (13%)
Query: 252 QLGLIKEGNLEKMDWAGLMWSMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVE 311
QLGL+ E + + W +E AT E + L I + K EE
Sbjct: 358 QLGLV-----EALHFVAAAWQAVEPADIATCFREAGFGGGLNATITTSFKSEGEE----- 407
Query: 312 VEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIE 371
E E EE ++EEEEGE ++EEEEE+ + E +G ++G EE +V+E + + E
Sbjct: 408 -EEEEEEEEEEEEEEEGEGEEEEEEEE----EGEEEGGEGEEVG--EEEEVEEEGDESDE 460
Query: 372 LSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY-VGEPSLRPCHNRDRK 430
+++ E E+ EG + D+ Q E + G Y V E + P +
Sbjct: 461 ----EEEEEEEEEEESSSEGLEAEDWAQGVVEASG----GFGGYSVQEEAQCPTLHFLEG 512
Query: 431 GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
G D + + +EEE ++EE+EED+ ++D+ + G
Sbjct: 513 GED----SDSDSDEEEEDEEEDEEDEEDDDDDEDG 543
Score = 41.2 bits (95), Expect = 0.009
Identities = 32/112 (28%), Positives = 53/112 (46%), Gaps = 11/112 (9%)
Query: 386 KEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQV-KEDEGEE 444
K +GE E+ + E+ ++EE E G+ GE + +G + E+V +E+E EE
Sbjct: 402 KSEGEEEEEEEEEEEEEEEEE----GE----GEEEEEEEEEGEEEGGEGEEVGEEEEVEE 453
Query: 445 EEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEGMPSGTG--SSIQAMEAVQMP 494
E E +EEEE++ EE+E + +G+ +G E Q P
Sbjct: 454 EGDESDEEEEEEEEEEEESSSEGLEAEDWAQGVVEASGGFGGYSVQEEAQCP 505
>NFM_RAT (P12839) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 845
Score = 63.2 bits (152), Expect = 2e-09
Identities = 56/192 (29%), Positives = 96/192 (49%), Gaps = 14/192 (7%)
Query: 272 SMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAK 331
+++ +EL A+ EE A K + E+ E++ V E + EE K+EEEEG+
Sbjct: 473 TVIAEELAASAKEEKEEAEE-----KEEEPEV-EKSPVKSPEAKEEEEGEKEEEEEGQ-- 524
Query: 332 DEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEG 391
+EEEEED V D+ + G K G E+++ ++ EE E ++ E +K +G+
Sbjct: 525 EEEEEEDEGVKSDQAEEGGSEKEGSSEKDEGEQEEEGETEAEGEGEEAEAKEEKKTEGKV 584
Query: 392 EQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHE--Q 449
E+M E+ K E+ E K+ V + + + ++ KE+E EE+ E +
Sbjct: 585 EEMAIKEEIKVEKPEK----AKSPVPKSPVEEVKPKPEAKAGKDEQKEEEKVEEKKEVAK 640
Query: 450 EEEEEDDVEEDE 461
E +E+ VE+ E
Sbjct: 641 ESPKEEKVEKKE 652
Score = 45.4 bits (106), Expect = 5e-04
Identities = 47/167 (28%), Positives = 77/167 (45%), Gaps = 28/167 (16%)
Query: 296 IKSQHK---ELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDV 352
+K QHK E+ EET V + + E E+ + EE + EE+EE +E + +V
Sbjct: 445 LKVQHKFVEEIIEETKVEDEKSEMEDALTVIAEELAASAKEEKEE-----AEEKEEEPEV 499
Query: 353 KMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQ 412
+ V+ + +E EE E E+E+G+ E+ + E K ++ E G
Sbjct: 500 EKSPVKSPEAKEEEEGEKE-------------EEEEGQEEEEEEDEGVKSDQAEE---GG 543
Query: 413 KNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEE 459
G S + ++ +G E E EGEE E ++E++ E VEE
Sbjct: 544 SEKEGS-SEKDEGEQEEEG---ETEAEGEGEEAEAKEEKKTEGKVEE 586
Score = 40.4 bits (93), Expect = 0.015
Identities = 44/158 (27%), Positives = 74/158 (45%), Gaps = 11/158 (6%)
Query: 309 VVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQE--LE 366
V EV+ + E KDE++E E +E++E K+E + K V + + E ++
Sbjct: 610 VEEVKPKPEAKAGKDEQKEEEKVEEKKEVAKESPKEEKVEKKEEKPKDVPDKKKAESPVK 669
Query: 367 EHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHN 426
E +E + K + +EK+ E + + +K E E G + VG+ S
Sbjct: 670 EKAVEEMITITKSVKVSLEKDTKEEKPQQQEKVKEKAEEEG---GSEEEVGDKS-----P 721
Query: 427 RDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDV 464
++ K D E EG+EEE EQE +E+ +E+E V
Sbjct: 722 QESKKEDIAINGEVEGKEEE-EQETQEKGSGQEEEKGV 758
Score = 38.9 bits (89), Expect = 0.043
Identities = 44/182 (24%), Positives = 80/182 (43%), Gaps = 26/182 (14%)
Query: 305 EETLVVEVEGEGEEGVVKDEEE-EGEAKDEEEEEDGVVVK-------------DEVDGSG 350
EE E EGEGEE K+E++ EG+ ++ +E+ V K +EV
Sbjct: 558 EEEGETEAEGEGEEAEAKEEKKTEGKVEEMAIKEEIKVEKPEKAKSPVPKSPVEEVKPKP 617
Query: 351 DVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
+ K G E+ + +++EE + ++ + VEK++ E+ D KK E+ +
Sbjct: 618 EAKAGKDEQKEEEKVEEKK---EVAKESPKEEKVEKKE---EKPKDVPDKKKAESPVKEK 671
Query: 411 GQKNYVG-----EPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ + + SL ++ K E+VKE EE E+E ++ E + D+
Sbjct: 672 AVEEMITITKSVKVSLEK-DTKEEKPQQQEKVKEKAEEEGGSEEEVGDKSPQESKKEDIA 730
Query: 466 FH 467
+
Sbjct: 731 IN 732
Score = 35.8 bits (81), Expect = 0.36
Identities = 29/95 (30%), Positives = 45/95 (46%), Gaps = 10/95 (10%)
Query: 376 QDKVETLPVEKEQGEGEQMM-----DFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
++ +E VE E+ E E + + S KEE E +K E P + + K
Sbjct: 453 EEIIEETKVEDEKSEMEDALTVIAEELAASAKEEKEE--AEEKEEEPEVEKSPVKSPEAK 510
Query: 431 GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
E+ E E EEE E+EEEE++ V+ D+ + G
Sbjct: 511 E---EEEGEKEEEEEGQEEEEEEDEGVKSDQAEEG 542
Score = 32.7 bits (73), Expect = 3.1
Identities = 27/110 (24%), Positives = 50/110 (44%), Gaps = 8/110 (7%)
Query: 295 LIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKM 354
L K +E ++ V+ + E E G EEE G+ +E +++ + + EV+G + +
Sbjct: 687 LEKDTKEEKPQQQEKVKEKAEEEGG---SEEEVGDKSPQESKKEDIAINGEVEGKEEEEQ 743
Query: 355 GGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEE 404
E+ QE E+ + L + P E+++GE +KK E
Sbjct: 744 ETQEKGSGQEEEKGVVTNGL-----DVSPAEEKKGEDRSDDKVVVTKKVE 788
>CENB_MOUSE (P27790) Major centromere autoantigen B (Centromere
protein B) (CENP-B)
Length = 599
Score = 63.2 bits (152), Expect = 2e-09
Identities = 63/213 (29%), Positives = 93/213 (43%), Gaps = 36/213 (16%)
Query: 252 QLGLIKEGNLEKMDWAGLMWSMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVE 311
QLGL+ E + + W +E AT E + L I + K EE
Sbjct: 358 QLGLV-----EALHFVAAAWQAVEPSDIATCFREAGFGGGLNATITTSFKSEGEE----- 407
Query: 312 VEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIE 371
E E EE ++EEEEGE ++EEEEE+ G+ + G EE +E+EE
Sbjct: 408 -EEEEEEEEEEEEEEEGEGEEEEEEEE----------EGEEEGGEGEEEGEEEVEEE--- 453
Query: 372 LSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY-VGEPSLRPCHNRDRK 430
G+ E+ EG + D+ Q E + G Y V E + P +
Sbjct: 454 ---GEVDDSDEEEEESSSEGLEAEDWAQGVVEASG----GFGGYSVQEEAQFPTLHFLEG 506
Query: 431 GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
G D + + +EEE ++EE+EED+ EED+ D
Sbjct: 507 GED----SDSDSDEEEDDEEEDEEDEDEEDDED 535
Score = 34.7 bits (78), Expect = 0.81
Identities = 15/28 (53%), Positives = 21/28 (74%)
Query: 438 KEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+E+E EEEE E+EEE E + EE+E + G
Sbjct: 408 EEEEEEEEEEEEEEEGEGEEEEEEEEEG 435
Score = 34.3 bits (77), Expect = 1.1
Identities = 16/36 (44%), Positives = 25/36 (69%)
Query: 430 KGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+G + E+ +E+E EEEE E E EEE++ EE+ + G
Sbjct: 404 EGEEEEEEEEEEEEEEEEEGEGEEEEEEEEEGEEEG 439
Score = 33.5 bits (75), Expect = 1.8
Identities = 15/31 (48%), Positives = 22/31 (70%)
Query: 435 EQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
E+ +E+E EEEE + EEEE++ EE E + G
Sbjct: 410 EEEEEEEEEEEEEGEGEEEEEEEEEGEEEGG 440
Score = 32.3 bits (72), Expect = 4.0
Identities = 27/92 (29%), Positives = 42/92 (45%), Gaps = 25/92 (27%)
Query: 388 QGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEE-- 445
+ EGE+ + E+ ++EE E G++ E+ +E+EGEEE
Sbjct: 402 KSEGEEEEEEEEEEEEEEEEEGEGEE---------------------EEEEEEEGEEEGG 440
Query: 446 --EHEQEEEEEDDVEEDEHDVGFHFSTKHHLE 475
E E EEE E++ E D+ D S+ LE
Sbjct: 441 EGEEEGEEEVEEEGEVDDSDEEEEESSSEGLE 472
>CENB_SHEEP (P49451) Major centromere autoantigen B (Centromere
protein B) (CENP-B) (Fragment)
Length = 239
Score = 60.8 bits (146), Expect = 1e-08
Identities = 46/152 (30%), Positives = 77/152 (50%), Gaps = 12/152 (7%)
Query: 307 TLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELE 366
T ++ EGE EE ++EEEE +EEEEEDG +E + +G+ G E + +E+E
Sbjct: 39 TTALKSEGEEEEEEEEEEEEEEGEGEEEEEEDG----EEEEEAGE----GEELGEEEEVE 90
Query: 367 EHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEE-TEMWFLGQKNYVGEPSLRPCH 425
E ++ +++ E E+ EG + D+ Q E G + P+L
Sbjct: 91 EEGDVDTVEEEEEEE---EESSSEGLEAEDWAQGVVEAGGSFGGYGAQEEAQCPTLHFLE 147
Query: 426 NRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDV 457
+ D E+ +ED+ E+E+ E +EEE+D+V
Sbjct: 148 GEEDSESDSEEEEEDDDEDEDDEDDEEEDDEV 179
Score = 37.0 bits (84), Expect = 0.16
Identities = 17/27 (62%), Positives = 22/27 (80%)
Query: 435 EQVKEDEGEEEEHEQEEEEEDDVEEDE 461
E+ +E+E EEEE E EEEEE+D EE+E
Sbjct: 49 EEEEEEEEEEEEGEGEEEEEEDGEEEE 75
Score = 35.4 bits (80), Expect = 0.47
Identities = 18/46 (39%), Positives = 27/46 (58%)
Query: 430 KGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLE 475
+G + E+ +E+E EEE +EEEEED EE+E G + +E
Sbjct: 45 EGEEEEEEEEEEEEEEGEGEEEEEEDGEEEEEAGEGEELGEEEEVE 90
Score = 34.3 bits (77), Expect = 1.1
Identities = 14/25 (56%), Positives = 20/25 (80%)
Query: 439 EDEGEEEEHEQEEEEEDDVEEDEHD 463
+ EGEEEE E+EEEEE++ E +E +
Sbjct: 43 KSEGEEEEEEEEEEEEEEGEGEEEE 67
Score = 32.3 bits (72), Expect = 4.0
Identities = 20/56 (35%), Positives = 29/56 (51%)
Query: 406 EMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDE 461
E F G N +L+ + + + E+ +E EGEEEE E EEEE+ E +E
Sbjct: 27 EAGFGGGPNATITTALKSEGEEEEEEEEEEEEEEGEGEEEEEEDGEEEEEAGEGEE 82
>NKX1_BOVIN (Q28139) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1216
Score = 60.1 bits (144), Expect = 2e-08
Identities = 54/174 (31%), Positives = 88/174 (50%), Gaps = 30/174 (17%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEE------EDGVVVKDEVDGS------GDVK--MGG 356
EVEG+ +EG ++ E GE + +E+E E G V DE +G G+VK G
Sbjct: 859 EVEGDEDEGEIQ-AGEGGEVEGDEDEGEIQAGEAGEVEGDEDEGEIQAGEGGEVKGDEGE 917
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGE-GEQMMDFE-QSKKEETEMWFLGQKN 414
++ + E+E + E+ G+D+ E E +GE GEQ ++ E Q + ++ E G+
Sbjct: 918 IQAGEAGEVEGEDGEVEGGEDEGEIQAGEGGEGETGEQELNAEIQGEAKDDEEGVDGEGG 977
Query: 415 YVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHF 468
G G ++ +EDE EE+E E+EEEEE++ EE+E + +
Sbjct: 978 GDG-------------GDSEDEEEEDEEEEDEEEEEEEEEEEEEENEQPLSLEW 1018
Score = 53.5 bits (127), Expect = 2e-06
Identities = 56/190 (29%), Positives = 92/190 (47%), Gaps = 40/190 (21%)
Query: 311 EVEGEGEEGVVK---------DEE-------EEGEAKDEEEE------EDGVVVKDEVDG 348
EVEG+ +EG ++ DE+ E GE + +E+E E G V DE +G
Sbjct: 825 EVEGDEDEGEIQAGEGGEVEGDEDEGEIQAGEAGEVEGDEDEGEIQAGEGGEVEGDEDEG 884
Query: 349 ------SGDVKMGGVEENQVQELEEHNIELSLGQDKV-ETLPVEKEQGEGEQMMDFEQSK 401
+G+V+ G +E ++Q E ++ G+ + E VE E GE E D + +
Sbjct: 885 EIQAGEAGEVE-GDEDEGEIQAGEGGEVKGDEGEIQAGEAGEVEGEDGEVEGGEDEGEIQ 943
Query: 402 KEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCE--------QVKEDEGEEEEHEQEEEE 453
E G++ E ++ D +G+D E + +E+E EEEE E+EEEE
Sbjct: 944 AGEGGEGETGEQELNAE--IQGEAKDDEEGVDGEGGGDGGDSEDEEEEDEEEEDEEEEEE 1001
Query: 454 EDDVEEDEHD 463
E++ EE+E++
Sbjct: 1002 EEEEEEEENE 1011
Score = 39.3 bits (90), Expect = 0.033
Identities = 32/114 (28%), Positives = 56/114 (49%), Gaps = 28/114 (24%)
Query: 311 EVEGEGEEGVVK--------------DEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
EVEG +EG ++ + E +GEAKD+EE DG +G GD GG
Sbjct: 932 EVEGGEDEGEIQAGEGGEGETGEQELNAEIQGEAKDDEEGVDG-------EGGGD---GG 981
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
E++ +E EE E +++ E E+E+ E +++ ++++++ FL
Sbjct: 982 DSEDEEEEDEEEEDE----EEEEEEEEEEEEENEQPLSLEWPETRRKQAIYLFL 1031
Score = 38.1 bits (87), Expect = 0.073
Identities = 46/189 (24%), Positives = 71/189 (37%), Gaps = 17/189 (8%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
E G +GV + E GE E E E G GD +GG E + + E +I
Sbjct: 732 EDPGSQGVGAEAENTGERTGGEAEAPAEGENGERSG-GDAALGGESEGKAENESEGDIPA 790
Query: 373 SLGQDKVETLPVEKEQGEGE------QMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHN 426
D + ++ E GE + ++ E + E E Q GE +
Sbjct: 791 ERRGDDEDEGEIQAEGGEVKGDEDEGEIQAGEGGEVEGDEDEGEIQAGEGGEVE----GD 846
Query: 427 RDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEG------MPSG 480
D I + E EG+E+E E + E +VE DE + +EG + +G
Sbjct: 847 EDEGEIQAGEAGEVEGDEDEGEIQAGEGGEVEGDEDEGEIQAGEAGEVEGDEDEGEIQAG 906
Query: 481 TGSSIQAME 489
G ++ E
Sbjct: 907 EGGEVKGDE 915
Score = 35.0 bits (79), Expect = 0.62
Identities = 18/56 (32%), Positives = 32/56 (57%)
Query: 310 VEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQEL 365
V+ EG G+ G +DEEEE E +++EEEE+ ++E + + + E + Q +
Sbjct: 972 VDGEGGGDGGDSEDEEEEDEEEEDEEEEEEEEEEEEEENEQPLSLEWPETRRKQAI 1027
>NKX1_BISBI (O46383) Sodium/potassium/calcium exchanger 1
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger) (Fragment)
Length = 300
Score = 59.7 bits (143), Expect = 2e-08
Identities = 53/174 (30%), Positives = 87/174 (49%), Gaps = 31/174 (17%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEE------EDGVVVKDEVDGS------GDVK--MGG 356
EVEG+ +EG ++ E GE + +E+E E G V DE +G G+VK G
Sbjct: 93 EVEGDEDEGEIQ-AGEGGEVEGDEDEGEIQAGEGGEVEGDEDEGEIQAGEGGEVKDDEGE 151
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGE-GEQMMDFE-QSKKEETEMWFLGQKN 414
++ + E+E + E+ G+D+ E E +GE GEQ ++ E Q + ++ E G+
Sbjct: 152 IQAGEAGEVEGEDGEVEGGEDEGEIQAGEGGEGETGEQELNAEIQGEAKDDEEGVDGEGG 211
Query: 415 YVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHF 468
G D E +E++ EE+E E+EEEEE++ EE+E + +
Sbjct: 212 --------------GDGGDSEDEEEEDEEEDEEEEEEEEEEEEEENEQPLSLEW 251
Score = 54.7 bits (130), Expect = 8e-07
Identities = 52/175 (29%), Positives = 84/175 (47%), Gaps = 30/175 (17%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEE------EDGVVVKDEVDG----------SGDVKM 354
EV+G+ +EG ++ E GE + +E+E E G V DE +G GD
Sbjct: 76 EVKGDEDEGEIQ-AGEGGEVEGDEDEGEIQAGEGGEVEGDEDEGEIQAGEGGEVEGDEDE 134
Query: 355 GGVEENQVQELEEHNIELSLGQ-DKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQK 413
G ++ + E+++ E+ G+ +VE E E GE E + + + ET G++
Sbjct: 135 GEIQAGEGGEVKDDEGEIQAGEAGEVEGEDGEVEGGEDEGEIQAGEGGEGET-----GEQ 189
Query: 414 NYVGEPSLRPCHNRDRKGIDCE-----QVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
E ++ D +G+D E EDE EE+E E EEEEE++ EE+E +
Sbjct: 190 ELNAE--IQGEAKDDEEGVDGEGGGDGGDSEDEEEEDEEEDEEEEEEEEEEEEEE 242
Score = 38.5 bits (88), Expect = 0.056
Identities = 32/114 (28%), Positives = 56/114 (49%), Gaps = 29/114 (25%)
Query: 311 EVEGEGEEGVVK--------------DEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
EVEG +EG ++ + E +GEAKD+EE DG +G GD GG
Sbjct: 166 EVEGGEDEGEIQAGEGGEGETGEQELNAEIQGEAKDDEEGVDG-------EGGGD---GG 215
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
E++ +E EE + E ++ E E+E+ E +++ ++++++ FL
Sbjct: 216 DSEDEEEEDEEEDEE-----EEEEEEEEEEEENEQPLSLEWPETRRKQAIYLFL 264
Score = 37.0 bits (84), Expect = 0.16
Identities = 39/148 (26%), Positives = 59/148 (39%), Gaps = 16/148 (10%)
Query: 316 GEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLG 375
G +GV + E GE E E E G GD +GG E + + E +I
Sbjct: 3 GSQGVGAEAENTGERTGGEAEAPAEGENGERSG-GDAALGGESEGKAENESEGDIPAERR 61
Query: 376 QDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCE 435
D + ++ E GE + ++E E+ Q GE + D I
Sbjct: 62 GDDEDEGEIQAEGGE-------VKGDEDEGEI----QAGEGGEVE----GDEDEGEIQAG 106
Query: 436 QVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+ E EG+E+E E + E +VE DE +
Sbjct: 107 EGGEVEGDEDEGEIQAGEGGEVEGDEDE 134
Score = 36.6 bits (83), Expect = 0.21
Identities = 17/29 (58%), Positives = 22/29 (75%)
Query: 310 VEVEGEGEEGVVKDEEEEGEAKDEEEEED 338
V+ EG G+ G +DEEEE E +DEEEEE+
Sbjct: 206 VDGEGGGDGGDSEDEEEEDEEEDEEEEEE 234
>IE68_PRVKA (P24827) Immediate-early protein RSP40
Length = 364
Score = 59.3 bits (142), Expect = 3e-08
Identities = 48/161 (29%), Positives = 73/161 (44%), Gaps = 33/161 (20%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E E E E+G +DE+EEGE +++EEEE+G + DG DV EE+ E EE
Sbjct: 212 EDEDEEEDGEEEDEDEEGEEEEDEEEEEG-----DEDGETDV----YEEDDEAEDEEDEE 262
Query: 371 E------LSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPC 424
+ S+G D V P + GEG D + E+ +
Sbjct: 263 DGDDFDGASVGDDDVFEPPEDGSDGEGSGSDDGGDGEDEDED------------------ 304
Query: 425 HNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ D D E +++EGE+ + E+ EED+ E+ E + G
Sbjct: 305 EDEDEDEDDGEDEEDEEGEDGGEDGEDGEEDEDEDGEGEEG 345
Score = 36.6 bits (83), Expect = 0.21
Identities = 15/34 (44%), Positives = 24/34 (70%)
Query: 430 KGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+G CE EDE EEE+ E+E+E+E+ EE++ +
Sbjct: 203 RGSVCEDDGEDEDEEEDGEEEDEDEEGEEEEDEE 236
>TAF7_YEAST (Q05021) Transcription initiation factor TFIID subunit 7
(TBP-associated factor 7) (TBP-associated factor 67 kDa)
(TAFII-67)
Length = 590
Score = 58.9 bits (141), Expect = 4e-08
Identities = 48/170 (28%), Positives = 73/170 (42%), Gaps = 16/170 (9%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E EG G EG D+E++ E +E ++D D+ D GD++ G E E +E+
Sbjct: 384 EEEGSGAEG---DKEQQQEEVGDEVDQDTGGEDDDDDDDGDIEAAGGESESDDEKDENRQ 440
Query: 371 ELSLGQDKVETLP---------VEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSL 421
L D++ L + K + + KK E E ++ E S+
Sbjct: 441 HTELLADELNELETTLAHTKHKLSKATNPLLKSRFIDSIKKLEKEAELKRKQLQQTEDSV 500
Query: 422 RPCHNRDRKGIDCEQVKEDEGEEEEHEQEEE----EEDDVEEDEHDVGFH 467
+ H V+E+E EEEE E+E+E EEDD E DE + H
Sbjct: 501 QKQHQHRSDAETANNVEEEEEEEEEEEEEDEVDEDEEDDEENDEDEDNVH 550
Score = 38.1 bits (87), Expect = 0.073
Identities = 26/102 (25%), Positives = 50/102 (48%), Gaps = 7/102 (6%)
Query: 296 IKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMG 355
+K + + E+++ + + + + EEE E ++EEEEED V +E D D
Sbjct: 488 LKRKQLQQTEDSVQKQHQHRSDAETANNVEEEEEEEEEEEEEDEVDEDEEDDEENDEDED 547
Query: 356 GVEENQ-------VQELEEHNIELSLGQDKVETLPVEKEQGE 390
V E + V+EL+E E +L Q+ ++ + + +G+
Sbjct: 548 NVHEREHIQENKVVRELDEAPAEETLDQNDLDMMMLFGAEGD 589
Score = 31.2 bits (69), Expect = 8.9
Identities = 23/78 (29%), Positives = 34/78 (43%), Gaps = 5/78 (6%)
Query: 399 QSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVE 458
Q+++E W K GEP RP ++ V + EEE E EEE E
Sbjct: 317 QARQERVSSWE-NFKEEPGEPLSRPALKKEEIHTIASAVGKQGAEEEGEEGMEEE----E 371
Query: 459 EDEHDVGFHFSTKHHLEG 476
E++ D+G F ++ G
Sbjct: 372 EEDLDLGAAFESEEEGSG 389
>NFM_MOUSE (P08553) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 848
Score = 57.8 bits (138), Expect = 9e-08
Identities = 58/176 (32%), Positives = 87/176 (48%), Gaps = 19/176 (10%)
Query: 301 KELFEETLVVEVEGEGEEGVVK-----DEEEEGE------AKDEEEEEDGVVVKDEVDGS 349
KE EE E E E E+ VK +EEEEGE ++EEEEED V D+ +
Sbjct: 484 KEEKEEAEEKEEEPEAEKSPVKSPEAKEEEEEGEKEEEEEGQEEEEEEDEGVKSDQAEEG 543
Query: 350 GDVKMGGVEENQ-VQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMW 408
G K G E+++ QE EE E ++ E +K +G+ E++ E+ K E+ E
Sbjct: 544 GSEKEGSSEKDEGEQEEEEGETEAEGEGEEAEAKEEKKIEGKVEEVAVKEEIKVEKPEKA 603
Query: 409 FLGQ-KNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHE--QEEEEEDDVEEDE 461
K+ V E +P + K EQ +E++ EEE+ E +E +E+ VE+ E
Sbjct: 604 KSPMPKSPVEEVKPKP----EAKAGKGEQKEEEKVEEEKKEVTKESPKEEKVEKKE 655
Score = 43.5 bits (101), Expect = 0.002
Identities = 47/181 (25%), Positives = 80/181 (43%), Gaps = 23/181 (12%)
Query: 305 EETLVVEVEGEGEEGVVKDEEE-EGEAKDEEEEEDGVVVK-------------DEVDGSG 350
EE E EGEGEE K+E++ EG+ ++ +E+ V K +EV
Sbjct: 560 EEEGETEAEGEGEEAEAKEEKKIEGKVEEVAVKEEIKVEKPEKAKSPMPKSPVEEVKPKP 619
Query: 351 DVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
+ K G E+ + +++EE E++ K E VEK++ E+ D KK E+ +
Sbjct: 620 EAKAGKGEQKEEEKVEEEKKEVTKESPKEE--KVEKKE---EKPKDVADKKKAESPVKEK 674
Query: 411 GQKNYVG-EPSLRPCHNRD---RKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGF 466
+ + S++ +D K E+VKE EE E+E + E + D+
Sbjct: 675 AVEEVITISKSVKVSLEKDTKEEKPQPQEKVKEKAEEEGGSEEEGSDRSPQESKKEDIAI 734
Query: 467 H 467
+
Sbjct: 735 N 735
Score = 37.4 bits (85), Expect = 0.12
Identities = 43/173 (24%), Positives = 63/173 (35%), Gaps = 57/173 (32%)
Query: 296 IKSQHK---ELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDV 352
+K QHK E+ EET V + + E EE + EE + EE+EE
Sbjct: 445 LKVQHKFVEEIIEETKVEDEKSEMEETLTAIAEELAASAKEEKEE--------------- 489
Query: 353 KMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQ 412
EE E + V++ ++E+ EGE+ + E ++EE E
Sbjct: 490 ------------AEEKEEEPEAEKSPVKSPEAKEEEEEGEKEEEEEGQEEEEEED----- 532
Query: 413 KNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
E VK D+ EE E+E E D E E + G
Sbjct: 533 ----------------------EGVKSDQAEEGGSEKEGSSEKDEGEQEEEEG 563
Score = 35.8 bits (81), Expect = 0.36
Identities = 30/95 (31%), Positives = 43/95 (44%), Gaps = 18/95 (18%)
Query: 376 QDKVETLPVEKEQGEGEQMM-----DFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
++ +E VE E+ E E+ + + S KEE K E P ++
Sbjct: 453 EEIIEETKVEDEKSEMEETLTAIAEELAASAKEE--------KEEAEEKEEEP--EAEKS 502
Query: 431 GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ + KE EEEE E+EEEEE EE+E D G
Sbjct: 503 PVKSPEAKE---EEEEGEKEEEEEGQEEEEEEDEG 534
>VG48_SHV21 (Q01033) Hypothetical gene 48 protein
Length = 797
Score = 56.2 bits (134), Expect = 3e-07
Identities = 49/156 (31%), Positives = 75/156 (47%), Gaps = 7/156 (4%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E + EG+EG DE +EGE + E+E ++G DE + GD E+ + +E
Sbjct: 515 EGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGED 574
Query: 371 ELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
E G+D+ E E ++GE E + ++ + E + G + GE + +
Sbjct: 575 EGDEGEDEGED---EGDEGEDEGEDEGDEGEDEGEDE---GDEGDEGEDEGDEGEDEGDE 628
Query: 431 GIDCEQVKEDEGEEEEHEQEE-EEEDDVEEDEHDVG 465
G D EDEG+E E E +E E+E D EDE D G
Sbjct: 629 GEDEGDEGEDEGDEGEDEGDEGEDEGDEGEDEGDEG 664
Score = 54.7 bits (130), Expect = 8e-07
Identities = 51/166 (30%), Positives = 79/166 (46%), Gaps = 15/166 (9%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGD--------VKMGGVEENQV 362
E EGE EE KDE+EEGE + ++ E++G +DE + GD G E+++
Sbjct: 418 EDEGEDEED-EKDEKEEGEDEGDDGEDEG---EDEGEDEGDEGDEGDEGEDEGEDEDDEE 473
Query: 363 QELEEHNIELSLGQDKVETLPVEKEQG-EGEQMMDFEQSKKEETEMWFLGQKNYVGEPSL 421
E E+ E G+D+ + +++G EG++ D E + G + G+
Sbjct: 474 DEGEDEGDEGDEGEDEGDEGDEGEDEGDEGDEGKDEGDEGDEGKDEGDEGDEGDEGDEGE 533
Query: 422 RPCHNRDRKGIDCEQVKEDEGEEEEHEQEEE--EEDDVEEDEHDVG 465
+ +G D EDEG+E E E E+E E +D EDE D G
Sbjct: 534 DEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEG 579
Score = 53.1 bits (126), Expect = 2e-06
Identities = 50/155 (32%), Positives = 80/155 (51%), Gaps = 11/155 (7%)
Query: 311 EVEGEGEEGVVKDE-EEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHN 369
E E EG+EG DE E+EGE +D+EE+E +DE D GD G E ++ E E+
Sbjct: 448 EGEDEGDEGDEGDEGEDEGEDEDDEEDEG----EDEGD-EGD--EGEDEGDEGDEGEDEG 500
Query: 370 IELSLGQDKVETLPVEKEQG-EGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRD 428
E G+D+ + K++G EG++ + ++ + E + G+ G+ +
Sbjct: 501 DEGDEGKDEGDEGDEGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDE--GDEGEDEGDEGE 558
Query: 429 RKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+G D EDEGE+E E E+E ED+ +E E +
Sbjct: 559 DEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDE 593
Score = 50.4 bits (119), Expect = 1e-05
Identities = 48/159 (30%), Positives = 73/159 (45%), Gaps = 16/159 (10%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E E EG+EG E+EG+ ++E E++G +DE + GD E+ + +E
Sbjct: 542 EGEDEGDEG-----EDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGED 596
Query: 371 ELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
E G+D+ E E ++GE E ++ + E E G + GE + +
Sbjct: 597 EGDEGEDEGEDEGDEGDEGEDEGDEGEDEGDEGEDE----GDE---GEDEGDEGEDEGDE 649
Query: 431 GIDCEQVKEDEGEE----EEHEQEEEEEDDVEEDEHDVG 465
G D EDEG+E E+ E E+E D EDE D G
Sbjct: 650 GEDEGDEGEDEGDEGDEGEDEGDEGEDEGDEGEDEGDEG 688
Score = 50.1 bits (118), Expect = 2e-05
Identities = 52/162 (32%), Positives = 78/162 (48%), Gaps = 21/162 (12%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEE--DGVVVKDEVDGSGDVKMGGVEENQVQELEEH 368
E E EG+EG + E+EG+ DE E+E +G KDE D + K G E ++ E +E
Sbjct: 475 EGEDEGDEGD--EGEDEGDEGDEGEDEGDEGDEGKDEGDEGDEGKDEGDEGDEGDEGDEG 532
Query: 369 NIEL----SLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPC 424
E G+D+ + E ++GE E + ++ + E + G+ GE
Sbjct: 533 EDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDE--GED----- 585
Query: 425 HNRDRKGIDCEQVKEDEGEEEEHEQEEE-EEDDVEEDEHDVG 465
+G + E EDEG+E E E E+E +E D EDE D G
Sbjct: 586 -----EGDEGEDEGEDEGDEGEDEGEDEGDEGDEGEDEGDEG 622
Score = 48.9 bits (115), Expect = 4e-05
Identities = 49/162 (30%), Positives = 83/162 (50%), Gaps = 11/162 (6%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDE-EEEEDGVVVKDEVDGSGDVKMGGVEE-NQVQELEEH 368
E E EG++G ++E E E +DE +E ++G +DE + D + G +E ++ E E+
Sbjct: 433 EGEDEGDDG---EDEGEDEGEDEGDEGDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDE 489
Query: 369 NIELSLGQDKVETLPVEKEQG-EGEQMMDFEQSKKEETEMWFLGQKNY--VGEPSLRPCH 425
E G+D+ + K++G EG++ D E + +E + G+ + G+
Sbjct: 490 GDEGDEGEDEGDEGDEGKDEGDEGDEGKD-EGDEGDEGDEGDEGEDEWEDEGDEGEDEGD 548
Query: 426 NRDRKGIDCEQVKEDEGEEEEHEQEEE--EEDDVEEDEHDVG 465
+ +G + E EDEG+E E E E+E E +D EDE D G
Sbjct: 549 EGEDEGDEGEDEGEDEGDEGEDEGEDEGDEGEDEGEDEGDEG 590
Score = 47.8 bits (112), Expect = 9e-05
Identities = 55/176 (31%), Positives = 82/176 (46%), Gaps = 20/176 (11%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E E E EE +EE+EGE DEE+E+D K+E + GD E+ E +E +
Sbjct: 404 EYELENEEYNRDEEEDEGE--DEEDEKD---EKEEGEDEGDDGEDEGEDEGEDEGDEGD- 457
Query: 371 ELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
E G+D+ E E+++GE E ++ + E E G + GE +
Sbjct: 458 EGDEGEDEGEDEDDEEDEGEDEG----DEGDEGEDE----GDEGDEGEDEGDEGDEGKDE 509
Query: 431 GIDCEQVKE--DEGEE----EEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEGMPSG 480
G + ++ K+ DEG+E +E E E E+E D EDE D G + EG G
Sbjct: 510 GDEGDEGKDEGDEGDEGDEGDEGEDEWEDEGDEGEDEGDEGEDEGDEGEDEGEDEG 565
Score = 42.4 bits (98), Expect = 0.004
Identities = 41/153 (26%), Positives = 73/153 (46%), Gaps = 13/153 (8%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
E E ++ + K ++ + KDE E E+ +DE + G+ EE++ E EE E
Sbjct: 384 EKEYKKIIDKSDDRDDRDKDEYELENEEYNRDEEEDEGED-----EEDEKDEKEEGEDEG 438
Query: 373 SLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGI 432
G+D+ E E E EG++ + E ++E + + GE + +G
Sbjct: 439 DDGEDEGED-EGEDEGDEGDEGDEGEDEGEDEDD------EEDEGEDEGDEGDEGEDEGD 491
Query: 433 DCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ ++ EDEG+E + ++E +E D +DE D G
Sbjct: 492 EGDE-GEDEGDEGDEGKDEGDEGDEGKDEGDEG 523
Score = 40.8 bits (94), Expect = 0.011
Identities = 45/158 (28%), Positives = 70/158 (43%), Gaps = 16/158 (10%)
Query: 322 KDEEEEGEAKDEEEEEDG--VVVKDEVDGSGDVKMGGV--EENQVQELEEHNI-ELSLGQ 376
+DE E + DE + EDG V+ DE + V + V + Q E E I + S +
Sbjct: 338 EDEVELCISDDEVDSEDGNLCVLDDESESVNSVALRQVLTVDKQANEKEYKKIIDKSDDR 397
Query: 377 D---------KVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNR 427
D + E ++E+ EGE D + K+E + G+ GE +
Sbjct: 398 DDRDKDEYELENEEYNRDEEEDEGEDEEDEKDEKEEGEDEGDDGEDE--GEDEGEDEGDE 455
Query: 428 DRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+G + E EDE +EE+ ++E +E D EDE D G
Sbjct: 456 GDEGDEGEDEGEDEDDEEDEGEDEGDEGDEGEDEGDEG 493
Score = 40.0 bits (92), Expect = 0.019
Identities = 33/128 (25%), Positives = 61/128 (46%), Gaps = 15/128 (11%)
Query: 351 DVKMGGVEENQVQELEEHNIELSLGQDKVET----LPVEKEQGEGEQMMDFEQSKKEETE 406
D K+ V E ++ +E +EL + D+V++ L V ++ E + Q + +
Sbjct: 324 DHKLSPVSE--YEDFDEDEVELCISDDEVDSEDGNLCVLDDESESVNSVALRQVLTVDKQ 381
Query: 407 MWFLGQKNYVGEPSLRPCHNRDRKGIDCEQV----KEDEGEEEEHEQEEEEE-----DDV 457
K + + R ++D ++ E+ +EDEGE+EE E++E+EE DD
Sbjct: 382 ANEKEYKKIIDKSDDRDDRDKDEYELENEEYNRDEEEDEGEDEEDEKDEKEEGEDEGDDG 441
Query: 458 EEDEHDVG 465
E++ D G
Sbjct: 442 EDEGEDEG 449
Score = 31.2 bits (69), Expect = 8.9
Identities = 18/49 (36%), Positives = 27/49 (54%), Gaps = 3/49 (6%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEE 359
E E EG+EG DE +EG+ ++E E++G +DE D K G +
Sbjct: 680 EGEDEGDEGDEGDEGDEGDEGEDEGEDEG---EDEGDEGTKDKEGNANK 725
>MYT1_HUMAN (Q01538) Myelin transcription factor 1 (MYT1) (MYTI)
(Proteolipid protein binding protein) (PLPB1)
Length = 1121
Score = 56.2 bits (134), Expect = 3e-07
Identities = 53/177 (29%), Positives = 79/177 (43%), Gaps = 36/177 (20%)
Query: 302 ELFEETLVVEVEGEGEE---GVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVE 358
++ EETLV E G+ + G+V +E E EE E G+ ++ E + +V E
Sbjct: 173 QIAEETLVEEDLGQAAKPGPGIVHLLQEAAEGAASEEGEKGLFIQPE--DAEEVVEVTTE 230
Query: 359 ENQ---VQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY 415
+Q Q LE+ E S Q + + E E+ E E+ + E ++EE E
Sbjct: 231 RSQDLCPQSLEDAASEESSKQKGILSHEEEDEEEEEEEEEEEEDEEEEEEE--------- 281
Query: 416 VGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKH 472
E+ +E+E EEEE E+EEEEE++ EE DV F T H
Sbjct: 282 -------------------EEEEEEEEEEEEEEEEEEEEEEEEEAAPDVIFQEDTSH 319
Score = 35.0 bits (79), Expect = 0.62
Identities = 20/64 (31%), Positives = 37/64 (57%)
Query: 289 ASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDG 348
+S + ++ + ++ EE E E + EE ++EEEE E ++EEEEE+ ++E +
Sbjct: 248 SSKQKGILSHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEA 307
Query: 349 SGDV 352
+ DV
Sbjct: 308 APDV 311
Score = 34.7 bits (78), Expect = 0.81
Identities = 21/57 (36%), Positives = 31/57 (53%)
Query: 290 SHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEV 346
SH + + + +E EE E E E EE ++EEEE E ++EEEEE+ D +
Sbjct: 256 SHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEAAPDVI 312
Score = 33.1 bits (74), Expect = 2.3
Identities = 24/89 (26%), Positives = 45/89 (49%), Gaps = 6/89 (6%)
Query: 293 QHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDV 352
Q L + +E ++ ++ E E EE ++EEEE + ++EEEEE+ ++E + +
Sbjct: 238 QSLEDAASEESSKQKGILSHEEEDEEEEEEEEEEEEDEEEEEEEEEEEEEEEEEEEEEE- 296
Query: 353 KMGGVEENQVQELEEHNIELSLGQDKVET 381
EE + +E EE ++ +D T
Sbjct: 297 -----EEEEEEEEEEAAPDVIFQEDTSHT 320
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 56.2 bits (134), Expect = 3e-07
Identities = 51/168 (30%), Positives = 73/168 (43%), Gaps = 51/168 (30%)
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
K + KE+ EE+ V+ E EE V +DEEEE E ++EEEEE+ ++E
Sbjct: 552 KEESKEVQEESKEVQ---EDEEEVEEDEEEEEEEEEEEEEEE----EEE----------- 593
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYV 416
EE + +E EE E +D E E E E D E+ EE
Sbjct: 594 -EEEEEEEEEEEEDEDEEDEDDAE----EDEDDAEEDEDDAEEDDDEE------------ 636
Query: 417 GEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDV 464
D ++ +DE E+E+ E EEEEE++ EE E +
Sbjct: 637 ----------------DDDEEDDDEDEDEDEEDEEEEEEEEEESEKKI 668
Score = 53.5 bits (127), Expect = 2e-06
Identities = 48/190 (25%), Positives = 92/190 (48%), Gaps = 25/190 (13%)
Query: 297 KSQHKELFEETL---VVE--VEGEGEEGVVKDEEEEGEAKDEE----------------- 334
K +HK+ +E + VV+ +E E ++GV E+ EA +E+
Sbjct: 427 KGKHKKAKKEKVKKHVVKNVIEDEDKDGVEIINLEDKEACEEQHITVESRPLSQPQCKLI 486
Query: 335 EEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQM 394
+E + + + D+ + K ++E + + N+ K + +E+ + + ++
Sbjct: 487 DEPEQLTLMDK--SKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKH 544
Query: 395 MDFEQSKKEET-EMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEE 453
+D E+ KKEE+ E+ ++ E + + + + E+ +E+E EEEE E+EEEE
Sbjct: 545 VDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 604
Query: 454 EDDVEEDEHD 463
ED+ EEDE D
Sbjct: 605 EDEDEEDEDD 614
Score = 51.2 bits (121), Expect = 8e-06
Identities = 40/154 (25%), Positives = 75/154 (47%), Gaps = 32/154 (20%)
Query: 323 DEEEEGEAKDEEE-EEDGVVVKDEVDGS-GDV-----------KMGGVEENQVQELEEHN 369
DE E+ D+ + EE + +++++ G+ G V KM +EE ++Q+ + +
Sbjct: 487 DEPEQLTLMDKSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKHVD 546
Query: 370 IELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDR 429
E ++ E KE E E+ ++ ++ ++EE E +
Sbjct: 547 KEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEE-------------------EEEE 587
Query: 430 KGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+ + E+ +E+E EEEE +++EE+EDD EEDE D
Sbjct: 588 EEEEEEEEEEEEEEEEEEDEDEEDEDDAEEDEDD 621
Score = 48.9 bits (115), Expect = 4e-05
Identities = 59/229 (25%), Positives = 97/229 (41%), Gaps = 68/229 (29%)
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEE------EEGEAKDEEEEEDGVVVKDEVDGSG 350
+ +HKE EE EGE +EG K+EE ++ E K +E + G K + D
Sbjct: 376 EGEHKE--EE----HKEGEHKEGEHKEEEHKEEEHKKEEHKSKEHKSKGKKDKGKKDKGK 429
Query: 351 DVK------MGGVEENQVQELEEHNIELSLGQDK---------VETLPVEKEQ----GEG 391
K V +N +++ ++ +E+ +DK VE+ P+ + Q E
Sbjct: 430 HKKAKKEKVKKHVVKNVIEDEDKDGVEIINLEDKEACEEQHITVESRPLSQPQCKLIDEP 489
Query: 392 EQMMDFEQSKKEETEMWFLGQK-NYVGEPSLRPCHNRDRKGI------------------ 432
EQ+ ++SK EE + Q +G ++ P + +K +
Sbjct: 490 EQLTLMDKSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAELQKQKHVDKEE 549
Query: 433 ------------------DCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
D E+V+EDE EEEE E+EEEEE++ EE+E +
Sbjct: 550 DKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEE 598
Score = 48.1 bits (113), Expect = 7e-05
Identities = 44/184 (23%), Positives = 81/184 (43%), Gaps = 42/184 (22%)
Query: 285 ECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVV-----KDEEEEGEAKDEEEEEDG 339
+C + L ++ E+ L ++ + G G V +D ++ AK EE E
Sbjct: 482 QCKLIDEPEQLTLMDKSKVEEKNLSIQEQLIGTIGRVNVVPRRDNHKKKMAKIEEAE--- 538
Query: 340 VVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQ 399
+ + VD D K + +E++E + E+ +++VE E+E+ E E+ + E+
Sbjct: 539 LQKQKHVDKEEDKK------EESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEE 592
Query: 400 SKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEE 459
++EE E E+ +E++ +EE+ + EE+EDD EE
Sbjct: 593 EEEEEEE----------------------------EEEEEEDEDEEDEDDAEEDEDDAEE 624
Query: 460 DEHD 463
DE D
Sbjct: 625 DEDD 628
Score = 48.1 bits (113), Expect = 7e-05
Identities = 41/165 (24%), Positives = 67/165 (39%), Gaps = 51/165 (30%)
Query: 296 IKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMG 355
++ + KE+ E+ VE + E EE ++EEEE E ++EEEEE+
Sbjct: 558 VQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEE----------------- 600
Query: 356 GVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY 415
EE + E EE + +D E + E+ + E+ D E ++E E
Sbjct: 601 --EEEEEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDE--------- 649
Query: 416 VGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEED 460
++E EEEE E+EEE E ++ +
Sbjct: 650 -----------------------DEEDEEEEEEEEEESEKKIKRN 671
Score = 41.2 bits (95), Expect = 0.009
Identities = 31/119 (26%), Positives = 57/119 (47%), Gaps = 3/119 (2%)
Query: 275 EKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEE 334
+KE EE + ++ +E EE E E E EE ++EEEE E +DEE
Sbjct: 551 KKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEE 610
Query: 335 EEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQ 393
+E+D +D+ + D E++ ++ +E + + +D+ + E+E+ E E+
Sbjct: 611 DEDDAEEDEDDAEEDED---DAEEDDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEESEK 666
Score = 39.3 bits (90), Expect = 0.033
Identities = 38/181 (20%), Positives = 77/181 (41%), Gaps = 29/181 (16%)
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
K + KE+ E+ + + + + EE K +E+E + ++++E
Sbjct: 266 KQKEKEMKEQEKIEKKKKKQEEKEKKKQEKERKKQEKKER-------------------- 305
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYV 416
++ + + ++ IE + + + +K E E+ M EET N +
Sbjct: 306 -KQKEKEMKKQKKIEKERKKKEEKEKKKKKHDKENEETMQQPDQTSEET-------NNEI 357
Query: 417 GEPSLRPCHNRDRKGIDCE-QVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLE 475
P P + E + KE+E +E EH++ E +E++ +E+EH H S +H +
Sbjct: 358 MVPLPSPLTDVTTPEEHKEGEHKEEEHKEGEHKEGEHKEEEHKEEEHKKEEHKSKEHKSK 417
Query: 476 G 476
G
Sbjct: 418 G 418
Score = 33.5 bits (75), Expect = 1.8
Identities = 45/199 (22%), Positives = 81/199 (40%), Gaps = 26/199 (13%)
Query: 301 KELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEV-DGSGDVKMGGVEE 359
+E +E +V E+ + G++ + E+ G + V + S D K E
Sbjct: 189 EENLDEEMVSEINNNAQGGLLLSSPYQ------YREQGGCGIISSVHETSNDTKDNDKEN 242
Query: 360 NQVQELEEHNIELSLGQ-DKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGE 418
+ E+H E L DK E EKE E E++ + ++ K+EE E
Sbjct: 243 ISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKI-EKKKKKQEEKE------------ 289
Query: 419 PSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEGMP 478
+ ++RK + ++ K+ E E ++ ++ E+E EE E H K + E M
Sbjct: 290 ---KKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKKKH--DKENEETMQ 344
Query: 479 SGTGSSIQAMEAVQMPFGS 497
+S + + +P S
Sbjct: 345 QPDQTSEETNNEIMVPLPS 363
Score = 31.2 bits (69), Expect = 8.9
Identities = 38/163 (23%), Positives = 71/163 (43%), Gaps = 24/163 (14%)
Query: 305 EETLVVEVEGEGEEGVV---KDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQ 361
+ T ++ V + E V KD++E+ KD++E+++ K++ D + +
Sbjct: 101 DPTNIINVNDKDNENSVDKKKDKKEKKHKKDKKEKKEKKDKKEKKDKKEKKHKKEKKHKK 160
Query: 362 VQELEEHNIELSL---GQDKVETLPVEKEQGEGEQMMD----------FEQSKKEETEMW 408
++ +E++ +SL GQ K + E+ E+M+ S + E
Sbjct: 161 DKKKKENSEVMSLYKTGQHKPKNATEHGEENLDEEMVSEINNNAQGGLLLSSPYQYREQG 220
Query: 409 FLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEE 451
G + V E S D K D E + ED ++E+H+QEE
Sbjct: 221 GCGIISSVHETS------NDTKDNDKENISED--KKEDHQQEE 255
>YCF2_OENPI (P31568) Protein ycf2 (Fragment)
Length = 721
Score = 55.8 bits (133), Expect = 3e-07
Identities = 73/262 (27%), Positives = 124/262 (46%), Gaps = 45/262 (17%)
Query: 231 LASNWMLLHDDTHIMTKDVMQQLGLIKEGNLEKMDWAGLMWSMLEKELKATHLEECYYAS 290
L +++ L+ +D+ D++ L I G LE ++ A + S E+E++ T E
Sbjct: 145 LITSYGLVENDS-----DLVHGLSDIVHGLLE-LEGALVGSSPTEEEVEGTEEE----VE 194
Query: 291 HLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSG 350
+ ++ +E+ E T EVEG EE +EE EG EEE +G ++EV+G+
Sbjct: 195 GTEEEVEGTEEEV-EGTEDEEVEGTEEEVEGTEEEVEGT----EEEVEGT--EEEVEGTE 247
Query: 351 DVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
D ++ G E+ +V+ EE E+ +++VE E+ +G E++ E + E TE
Sbjct: 248 DEEVEGTEDEEVEGTEE---EVEGTEEEVEGTE-EEVEGTEEEVEGTEDEEVEGTEEEVE 303
Query: 411 GQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFST 470
G + V +G + E+V EG EEE E EEE + E++E + T
Sbjct: 304 GTEEEV-------------EGTEDEEV---EGTEEEVEGTEEEVEGTEDEEVE-----GT 342
Query: 471 KHHLEGMP---SGTGSSIQAME 489
+ +EG GT ++ E
Sbjct: 343 EEEVEGTEEEVEGTEEEVEGTE 364
Score = 50.1 bits (118), Expect = 2e-05
Identities = 54/171 (31%), Positives = 84/171 (48%), Gaps = 18/171 (10%)
Query: 301 KELFEETLVVEVEGEGEEGVVKDEEE----EGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
+E E T EVEG +E V EEE E E + EEE +G ++EV+G+ D ++ G
Sbjct: 240 EEEVEGTEDEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGT--EEEVEGTEDEEVEG 297
Query: 357 VEEN------QVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
EE +V+ E+ +E + +++VE E E E E++ E+ + E TE
Sbjct: 298 TEEEVEGTEEEVEGTEDEEVEGT--EEEVEGTEEEVEGTEDEEVEGTEE-EVEGTEEEVE 354
Query: 411 GQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDE 461
G + V E + + + E+ E EG EEE E EEE + E++E
Sbjct: 355 GTEEEV-EGTEEEVEGTEEEVEGTEE--EVEGTEEEVEGTEEEVEGTEDEE 402
Score = 47.4 bits (111), Expect = 1e-04
Identities = 52/163 (31%), Positives = 82/163 (49%), Gaps = 14/163 (8%)
Query: 305 EETLVVEVEGEGEEGVVKDEEEEGEAKDEE----EEEDGVVVKDEVDGSGDVKMGGVEEN 360
EE E E EG E V+ EEE E +EE E+E+ ++EV+G+ + ++ G E+
Sbjct: 257 EEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEDEEVEGTEEEVEGTEE-EVEGTEDE 315
Query: 361 QVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPS 420
+V+ EE E+ +++VE E+ +G E++ E+ + E TE G + V E +
Sbjct: 316 EVEGTEE---EVEGTEEEVEGTEDEEVEGTEEEVEGTEE-EVEGTEEEVEGTEEEV-EGT 370
Query: 421 LRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+ + E+ E EG EEE E E+EE VE E D
Sbjct: 371 EEEVEGTEEEVEGTEE--EVEGTEEEVEGTEDEE--VEGTEKD 409
Score = 42.0 bits (97), Expect = 0.005
Identities = 55/212 (25%), Positives = 94/212 (43%), Gaps = 32/212 (15%)
Query: 258 EGNLEKMDWAGLMWSMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGE 317
EG E+++ E+E++ T EE + ++ +E+ E T EVEG E
Sbjct: 267 EGTEEEVEGTEEEVEGTEEEVEGTEDEE---VEGTEEEVEGTEEEV-EGTEDEEVEGTEE 322
Query: 318 EGVVKDEEEEGEAKDE----EEEEDGVV-----VKDEVDGSGDVKMGGVEENQVQELEEH 368
E +EE EG +E EEE +G ++EV+G+ + G EE + E E
Sbjct: 323 EVEGTEEEVEGTEDEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVEGTEEEVE 382
Query: 369 NIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRD 428
E + + E E E+ EG + + S+ + + L LRP +
Sbjct: 383 GTEEEVEGTEEEVEGTEDEEVEGTEK---DSSQFDNDRVTLL----------LRP---KP 426
Query: 429 RKGIDCEQVKEDEGEEEEHEQEEEEEDDVEED 460
R +D +++ + +++E E EE+DD +ED
Sbjct: 427 RNPLDIQRLIY---QHQKYESELEEDDDDDED 455
Score = 41.2 bits (95), Expect = 0.009
Identities = 46/168 (27%), Positives = 70/168 (41%), Gaps = 70/168 (41%)
Query: 296 IKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMG 355
I + EL E+ L EV V EEEGEA DEE DV +
Sbjct: 495 IPEEEDELPEDALETEV-------AVWGVEEEGEADDEE----------------DVLLE 531
Query: 356 GVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNY 415
+E+++ LEE + EL +D+++ E+E E+ K+EE E+
Sbjct: 532 AQQEDEL--LEEEDEELDEEEDELD----EEE----------EEPKEEEDEL-------- 567
Query: 416 VGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+E EEEE E+EEEEED+++E++ +
Sbjct: 568 -----------------------HEEEEEEEEEEEEEEEDELQENDSE 592
Score = 40.0 bits (92), Expect = 0.019
Identities = 40/154 (25%), Positives = 66/154 (41%), Gaps = 15/154 (9%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
E E EE DE+ K E+ +V + D + E +E+ E EL
Sbjct: 443 ESELEEDDDDDEDVFAPQKMLEDLFSELVWSPRIWHPWDFLLDCEAEIPAEEIPEEEDEL 502
Query: 373 SLGQDKVETLPVE---KEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDR 429
+D +ET +E+GE + D ++E E+ + + +
Sbjct: 503 P--EDALETEVAVWGVEEEGEADDEEDVLLEAQQEDELLEEEDEEL----------DEEE 550
Query: 430 KGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+D E+ + E E+E HE+EEEEE++ EE+E D
Sbjct: 551 DELDEEEEEPKEEEDELHEEEEEEEEEEEEEEED 584
Score = 35.0 bits (79), Expect = 0.62
Identities = 30/111 (27%), Positives = 52/111 (46%), Gaps = 6/111 (5%)
Query: 237 LLHDDTHIMTKDVMQQLGLIKEGNLEKMDWAGLMWSMLEKELKATHLEECYYASHLQHLI 296
LL + I +++ ++ + E LE +W + E+ + A L+
Sbjct: 483 LLDCEAEIPAEEIPEEEDELPEDALET---EVAVWGVEEEGEADDEEDVLLEAQQEDELL 539
Query: 297 KSQHKELFEETLVVEVEGEG---EEGVVKDEEEEGEAKDEEEEEDGVVVKD 344
+ + +EL EE ++ E E EE + +EEEE E ++EEEEED + D
Sbjct: 540 EEEDEELDEEEDELDEEEEEPKEEEDELHEEEEEEEEEEEEEEEDELQEND 590
>NKX1_RAT (Q9QZM6) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1181
Score = 55.8 bits (133), Expect = 3e-07
Identities = 48/156 (30%), Positives = 73/156 (46%), Gaps = 26/156 (16%)
Query: 311 EVEGEGE------EGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQE 364
EVE E E EG + E +E + + E E E V + E + G V+ G E + E
Sbjct: 842 EVETEAERKETNHEGETEAEGKEADHEGETEAEGNVEHQGETEAEGKVEHEG--ETEAGE 899
Query: 365 LEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPC 424
+EH GQ + + E + GEGE E + +++ E G+K G
Sbjct: 900 KDEHE-----GQSETQADDTEVKDGEGEA----EANAEDQCET-AQGEKGADG------- 942
Query: 425 HNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEED 460
G D E+ +++E EEEE E+EEEEE++ E+
Sbjct: 943 -GGGSDGGDSEEEEDEEDEEEEEEEEEEEEEEESEE 977
Score = 54.7 bits (130), Expect = 8e-07
Identities = 48/164 (29%), Positives = 74/164 (44%), Gaps = 15/164 (9%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELE-EHN 369
E E EG+E V ++ E E E K+ E E + + E + G+ + G E + E E E N
Sbjct: 818 ETEAEGKE-VEQEGETEAEGKEVEHEVETEAERKETNHEGETEAEGKEADHEGETEAEGN 876
Query: 370 IELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRP---CHN 426
+E Q + E + +GE E E + ET+ K+ GE C
Sbjct: 877 VE---HQGETEAEGKVEHEGETEAGEKDEHEGQSETQADDTEVKDGEGEAEANAEDQCET 933
Query: 427 RDRK-------GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+ G D +E+E EE+E E+EEEEE++ EE+ +
Sbjct: 934 AQGEKGADGGGGSDGGDSEEEEDEEDEEEEEEEEEEEEEEESEE 977
Score = 42.7 bits (99), Expect = 0.003
Identities = 47/184 (25%), Positives = 80/184 (42%), Gaps = 15/184 (8%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKD--EVDGSGDVKMGGVEENQVQELEEHNI 370
EGE E ++E E EA+ +E + +G + EV+ G+ + G E+ Q E E
Sbjct: 765 EGETESEGKDEQEGETEAEGKEADHEGETEAEGKEVEHEGETEAEGTEDEQEGETEAEGK 824
Query: 371 ELSL-GQDKVETLPVEKE---QGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHN 426
E+ G+ + E VE E + E ++ +++ E E G+ E ++
Sbjct: 825 EVEQEGETEAEGKEVEHEVETEAERKETNHEGETEAEGKEADHEGETE--AEGNVEHQGE 882
Query: 427 RDRKGIDCEQVKEDEGEEEEHE-QEEEEEDDVE------EDEHDVGFHFSTKHHLEGMPS 479
+ +G + + + GE++EHE Q E + DD E E E + T +G
Sbjct: 883 TEAEGKVEHEGETEAGEKDEHEGQSETQADDTEVKDGEGEAEANAEDQCETAQGEKGADG 942
Query: 480 GTGS 483
G GS
Sbjct: 943 GGGS 946
Score = 42.4 bits (98), Expect = 0.004
Identities = 44/157 (28%), Positives = 70/157 (44%), Gaps = 11/157 (7%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E E +G+E ++ E E E KDE+E E K E D G+ + G E E E
Sbjct: 755 ETETKGKEK--QEGETESEGKDEQEGETEAEGK-EADHEGETEAEGKEVEHEGETEAEGT 811
Query: 371 E-LSLGQDKVETLPVEKE---QGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHN 426
E G+ + E VE+E + EG+++ +++ E E G+ G+ +
Sbjct: 812 EDEQEGETEAEGKEVEQEGETEAEGKEVEHEVETEAERKETNHEGETEAEGKEADHEGET 871
Query: 427 RDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
++ + E EG + EHE E E E+DEH+
Sbjct: 872 EAEGNVEHQGETEAEG-KVEHEGETEAG---EKDEHE 904
Score = 42.4 bits (98), Expect = 0.004
Identities = 40/152 (26%), Positives = 67/152 (43%), Gaps = 13/152 (8%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
E G + V + E G+ EE E + + + D G+ + E+ Q +E E E
Sbjct: 705 EDPGCQEDVDEAEHRGDMTGEEGERE-TEAEGKKDEEGETEAERKEDGQEEETETKGKEK 763
Query: 373 SLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGI 432
G+ + E K++ EGE + +++ E + + GE + +G
Sbjct: 764 QEGETESEG----KDEQEGETEAEGKEADHEGETEAEGKEVEHEGET--------EAEGT 811
Query: 433 DCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDV 464
+ EQ E E E +E EQE E E + +E EH+V
Sbjct: 812 EDEQEGETEAEGKEVEQEGETEAEGKEVEHEV 843
Score = 42.0 bits (97), Expect = 0.005
Identities = 39/150 (26%), Positives = 65/150 (43%), Gaps = 5/150 (3%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKD-EVDGSGDVKMGGVEENQVQELEEHN 369
E EG+ +E + E + + ++EE E G ++ E + G + G E + +E +
Sbjct: 732 EAEGKKDEEGETEAERKEDGQEEETETKGKEKQEGETESEGKDEQEGETEAEGKEADHEG 791
Query: 370 IELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNR-D 428
+ G++ E E E EQ + E KE + G+ G+ +
Sbjct: 792 ETEAEGKEVEHEGETEAEGTEDEQEGETEAEGKEVEQE---GETEAEGKEVEHEVETEAE 848
Query: 429 RKGIDCEQVKEDEGEEEEHEQEEEEEDDVE 458
RK + E E EG+E +HE E E E +VE
Sbjct: 849 RKETNHEGETEAEGKEADHEGETEAEGNVE 878
Score = 34.7 bits (78), Expect = 0.81
Identities = 29/108 (26%), Positives = 52/108 (47%), Gaps = 9/108 (8%)
Query: 299 QHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVE 358
+H+ E VE EGE E G E++E E + E + +D V DG G+ + +
Sbjct: 878 EHQGETEAEGKVEHEGETEAG----EKDEHEGQSETQADDTEV----KDGEGEAEANAED 929
Query: 359 ENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETE 406
+ + + E + G D ++ E E+ E E+ + E+ ++EE+E
Sbjct: 930 QCETAQ-GEKGADGGGGSDGGDSEEEEDEEDEEEEEEEEEEEEEEESE 976
Score = 34.3 bits (77), Expect = 1.1
Identities = 27/98 (27%), Positives = 47/98 (47%), Gaps = 9/98 (9%)
Query: 313 EGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIEL 372
E + ++ VKD E E EA E++ E K DG G G EE + +E EE
Sbjct: 908 ETQADDTEVKDGEGEAEANAEDQCETAQGEKG-ADGGGGSDGGDSEEEEDEEDEE----- 961
Query: 373 SLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
+++ E E+E+ E +++ +S++++ FL
Sbjct: 962 ---EEEEEEEEEEEEESEEPLSLEWPESRQKQAIYLFL 996
>CNG4_BOVIN (Q28181) 240 kDa protein of rod photoreceptor
CNG-channel [Contains: Glutamic acid-rich protein
(GARP); Cyclic-nucleotide-gated cation channel 4 (CNG
channel 4) (CNG-4) (Cyclic nucleotide-gated cation
channel modulatory subunit)]
Length = 1394
Score = 55.1 bits (131), Expect = 6e-07
Identities = 44/167 (26%), Positives = 75/167 (44%), Gaps = 35/167 (20%)
Query: 310 VEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHN 369
++ E E EE +D EEE E E+EEE+G ++E + + G +E + +E EE
Sbjct: 353 IQEEKEDEEEEKEDGEEEEEEGREKEEEEGEEKEEEEGREKEEEEGEKKEEEGREKEEEE 412
Query: 370 IELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDR 429
G +K + EKE+ EG + E +KEE E R +
Sbjct: 413 -----GGEKEDEEGREKEEEEGRGKEEEEGGEKEEEE-------------------GRGK 448
Query: 430 KGIDCEQVKEDEGEEEEH-----------EQEEEEEDDVEEDEHDVG 465
+ ++ + +EDE EE++H +E+ E+ +D+ +VG
Sbjct: 449 EEVEGREEEEDEEEEQDHSVLLDSYLVPQSEEDRSEESETQDQSEVG 495
Score = 52.4 bits (124), Expect = 4e-06
Identities = 51/163 (31%), Positives = 78/163 (47%), Gaps = 35/163 (21%)
Query: 305 EETLVVEVEGEGEEGVVKDEEEEGEAKD--EEEEEDGVVVKDEVDGSGDVKMGGVEENQV 362
E+ VVE + +++E+E E K+ EEEEE+G +++ + G+ K EE +
Sbjct: 338 EKGKVVEQTPRELPRIQEEKEDEEEEKEDGEEEEEEG---REKEEEEGEEK----EEEEG 390
Query: 363 QELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLR 422
+E EE E K E EKE+ EG + D E +KEE E
Sbjct: 391 REKEEEEGE------KKEEEGREKEEEEGGEKEDEEGREKEEEE---------------- 428
Query: 423 PCHNRDRKGIDCEQVKEDEGE-EEEHEQEEEEEDDVEEDEHDV 464
R ++ + + +E+EG +EE E EEEED+ EE +H V
Sbjct: 429 ---GRGKEEEEGGEKEEEEGRGKEEVEGREEEEDEEEEQDHSV 468
Score = 40.4 bits (93), Expect = 0.015
Identities = 47/153 (30%), Positives = 69/153 (44%), Gaps = 34/153 (22%)
Query: 297 KSQHKELFEETLVVEVEGEG--EEGVVKDEEEEGEAKDEE------------EEEDGVVV 342
+ + KE E E EGE EEG K+EEE GE +DEE EEE+G
Sbjct: 381 EGEEKEEEEGREKEEEEGEKKEEEGREKEEEEGGEKEDEEGREKEEEEGRGKEEEEGG-E 439
Query: 343 KDEVDGSGDVKMGGVEENQ----------------VQELEEHNIELSLGQDKVETLPVEK 386
K+E +G G ++ G EE + V + EE E S QD+ E + +
Sbjct: 440 KEEEEGRGKEEVEGREEEEDEEEEQDHSVLLDSYLVPQSEEDRSEESETQDQSE-VGGAQ 498
Query: 387 EQGE--GEQMMDFEQSKKEETEMWFLGQKNYVG 417
QGE G Q + E ++++E+ ++ VG
Sbjct: 499 AQGEVGGAQALSEESETQDQSEVGGAQDQSEVG 531
Score = 35.0 bits (79), Expect = 0.62
Identities = 22/54 (40%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Query: 412 QKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+K V E + R + D E+ KED GEEEE E E+EE++ EE E + G
Sbjct: 338 EKGKVVEQTPRELPRIQEEKEDEEEEKED-GEEEEEEGREKEEEEGEEKEEEEG 390
Score = 33.9 bits (76), Expect = 1.4
Identities = 20/72 (27%), Positives = 28/72 (38%), Gaps = 8/72 (11%)
Query: 22 PEQPPPPPPP--------PPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDED 73
PEQPP P PP P P+P + E P + P P D++ +E
Sbjct: 1284 PEQPPRPEPPAPEAPAPEPTAPEPLAPEAPAPEAPAPSSPPPASQERPEGDKDAARPEEH 1343
Query: 74 PTHPQIEDDPEP 85
P + P+P
Sbjct: 1344 PVRIHVTLGPDP 1355
>NKX1_HUMAN (O60721) Sodium/potassium/calcium exchanger 1 precursor
(Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod
Na-Ca+K exchanger)
Length = 1099
Score = 53.9 bits (128), Expect = 1e-06
Identities = 48/163 (29%), Positives = 75/163 (45%), Gaps = 32/163 (19%)
Query: 316 GEEGVVKDEE------EEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHN 369
G+E V + E EEGE E E E+ + + +G G+ + G E E E
Sbjct: 743 GQEDVAEAESTGEMPGEEGETAGEGETEEKSGGETQPEGEGETETQGKGEECEDENEAEG 802
Query: 370 IELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDR 429
+ G+D+ E + E GE + + ETE L +N+ GE D
Sbjct: 803 KGDNEGEDEGE---IHAEDGE-------MKGNEGETESQELSAENH-GEAK------NDE 845
Query: 430 KGI---------DCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
KG+ D E+ +E+E E+EE E+EEE+E++ EE+E +
Sbjct: 846 KGVEDGGGSDGGDSEEEEEEEEEQEEEEEEEEQEEEEEEEEEE 888
Score = 46.2 bits (108), Expect = 3e-04
Identities = 45/151 (29%), Positives = 67/151 (43%), Gaps = 42/151 (27%)
Query: 311 EVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNI 370
E +G+GEE ++E E E K + E ED + E G++K G E + QEL N
Sbjct: 786 ETQGKGEEC---EDENEAEGKGDNEGEDEGEIHAE---DGEMK-GNEGETESQELSAENH 838
Query: 371 ELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRK 430
G+ K + VE G G D E+ ++EE E
Sbjct: 839 ----GEAKNDEKGVE--DGGGSDGGDSEEEEEEEEE------------------------ 868
Query: 431 GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDE 461
Q +E+E EE+E E+EEEEE++ + +E
Sbjct: 869 -----QEEEEEEEEQEEEEEEEEEEEEKGNE 894
Score = 38.1 bits (87), Expect = 0.073
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 6/106 (5%)
Query: 311 EVEGE--GEEGVVKDEEEEGEAKDEEEEEDGVVVKDE---VDGSGDVKMGGVEENQVQEL 365
E EGE E+G +K E E E+++ E G DE DG G EE + +E
Sbjct: 809 EDEGEIHAEDGEMKGNEGETESQELSAENHGEAKNDEKGVEDGGGSDGGDSEEEEEEEEE 868
Query: 366 EEHNIELSLGQDKVETLPVEKEQGEGEQM-MDFEQSKKEETEMWFL 410
+E E +++ E E+E+G E + +D+ ++++++ FL
Sbjct: 869 QEEEEEEEEQEEEEEEEEEEEEKGNEEPLSLDWPETRQKQAIYLFL 914
Score = 33.9 bits (76), Expect = 1.4
Identities = 29/89 (32%), Positives = 40/89 (44%), Gaps = 15/89 (16%)
Query: 293 QHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDV 352
Q L H E + VE G + G ++EEEE E ++EEEEE+ ++E
Sbjct: 831 QELSAENHGEAKNDEKGVEDGGGSDGGDSEEEEEEEEEQEEEEEEEE---QEE------- 880
Query: 353 KMGGVEENQVQELEEHNIELSLGQDKVET 381
EE + +E EE E L D ET
Sbjct: 881 -----EEEEEEEEEEKGNEEPLSLDWPET 904
>CENB_HUMAN (P07199) Major centromere autoantigen B (Centromere
protein B) (CENP-B)
Length = 599
Score = 53.5 bits (127), Expect = 2e-06
Identities = 55/208 (26%), Positives = 84/208 (39%), Gaps = 30/208 (14%)
Query: 259 GNLEKMDWAGLMWSMLEKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEE 318
G E + + W +E A E + I + K EE E E E EE
Sbjct: 360 GLTEALHFVAAAWQAVEPSDIAACFREAGFGGGPNATITTSLKSEGEEEEEEEEEEEEEE 419
Query: 319 GVVKDEEEEGEAKDEE-EEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQD 377
G ++EEEEGE ++EE E + + ++EV+ GDV EE
Sbjct: 420 GEGEEEEEEGEEEEEEGGEGEELGEEEEVEEEGDVDSDEEEEED---------------- 463
Query: 378 KVETLPVEKEQGEGEQMMDFEQSKKEE-TEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQ 436
E+ EG + D+ Q E G + P+L G D +
Sbjct: 464 -------EESSSEGLEAEDWAQGVVEAGGSFGAYGAQEEAQCPTLHFLEG----GEDSDS 512
Query: 437 VKEDEGEEEEHEQEEEEEDDVEEDEHDV 464
E+E +EEE +++E+++DD EED +V
Sbjct: 513 DSEEEDDEEEDDEDEDDDDD-EEDGDEV 539
Score = 50.1 bits (118), Expect = 2e-05
Identities = 45/160 (28%), Positives = 76/160 (47%), Gaps = 20/160 (12%)
Query: 307 TLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELE 366
T+ ++ EGEE +EEEE +EEEEE+G ++E +G + + GG E ++ E E
Sbjct: 396 TITTSLKSEGEE----EEEEE----EEEEEEEGEGEEEEEEGEEEEEEGG-EGEELGEEE 446
Query: 367 EHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMW-FLGQKNYVGEPSLRPCH 425
E E + D+ E E+ EG + D+ Q E + G + P+L
Sbjct: 447 EVEEEGDVDSDEEEEED-EESSSEGLEAEDWAQGVVEAGGSFGAYGAQEEAQCPTLHFLE 505
Query: 426 NRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
+ D EEE ++EE++ED+ ++D+ + G
Sbjct: 506 GGE---------DSDSDSEEEDDEEEDDEDEDDDDDEEDG 536
Score = 35.0 bits (79), Expect = 0.62
Identities = 25/96 (26%), Positives = 43/96 (44%), Gaps = 2/96 (2%)
Query: 412 QKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTK 471
+ + G P+ + +G + E+ +E+E EEE +EEEEE + EE+E G +
Sbjct: 386 EAGFGGGPNATITTSLKSEGEEEEEEEEEEEEEEGEGEEEEEEGEEEEEEGGEGEELGEE 445
Query: 472 HHLEGMPSGTGSSIQAMEAVQMPFGSGIHLHDSSVG 507
+E G S + E + G+ D + G
Sbjct: 446 EEVE--EEGDVDSDEEEEEDEESSSEGLEAEDWAQG 479
Score = 34.3 bits (77), Expect = 1.1
Identities = 30/111 (27%), Positives = 50/111 (45%), Gaps = 15/111 (13%)
Query: 386 KEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEE 445
K +GE E+ + E+ ++EE E GE + +G + E++ E+E EE
Sbjct: 402 KSEGEEEEEEE-EEEEEEEGE----------GEEEEEEGEEEEEEGGEGEELGEEEEVEE 450
Query: 446 EHEQEEEEEDDVEEDEHDVGFHFSTKHHLEGMPSGTGS--SIQAMEAVQMP 494
E + + +EE+ EEDE + +G+ GS + A E Q P
Sbjct: 451 EGDVDSDEEE--EEDEESSSEGLEAEDWAQGVVEAGGSFGAYGAQEEAQCP 499
>INVO_PANPA (P14591) Involucrin
Length = 560
Score = 53.1 bits (126), Expect = 2e-06
Identities = 40/152 (26%), Positives = 81/152 (52%), Gaps = 5/152 (3%)
Query: 315 EGEEGVVKD-EEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELS 373
E +EG +K E+++G+ + E++E + + ++ +G +K +E Q++ LE +L
Sbjct: 177 EQQEGQLKHLEQQKGQLELPEQQEGQLELPEQQEGQ--LKHLEQQEGQLKHLEHQEGQLE 234
Query: 374 LGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGID 433
+ +++V L EQ EG+ +Q K+ E +GQ ++ + +P H +KG
Sbjct: 235 VPEEQVGQLKY-LEQQEGQLKHLDQQEKQPELPEQQVGQLKHLEQQEGQPKHLEQQKG-Q 292
Query: 434 CEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVG 465
E ++E EG+ + EQ+E + + +E E +G
Sbjct: 293 LEHLEEQEGQLKHLEQQEGQLEHLEHQEGQLG 324
Score = 42.4 bits (98), Expect = 0.004
Identities = 62/249 (24%), Positives = 108/249 (42%), Gaps = 46/249 (18%)
Query: 251 QQLGLIK-----EGNLEKMDW-----------AGLMWSMLEKELKATHLEECYYASHLQH 294
+Q+G +K EG L+ +D G + + ++E + HLE+ L+H
Sbjct: 238 EQVGQLKYLEQQEGQLKHLDQQEKQPELPEQQVGQLKHLEQQEGQPKHLEQ--QKGQLEH 295
Query: 295 LIKSQ----HKELFEETLVVEVEGEGEEGVVKD--------EEEEGEAKDEEEEEDGVVV 342
L + + H E E L EG+ G+ + E+EEG+ K EEEE +
Sbjct: 296 LEEQEGQLKHLEQQEGQLEHLEHQEGQLGLPEQQVQQLKQLEKEEGQPKHLEEEEG--QL 353
Query: 343 KDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQ----GEGEQMMDFE 398
K V G ++ +E Q++ L + +L + +VE L + E G+ + + + E
Sbjct: 354 KHLVQQEGQLEHLVQQEGQLEHLVQQEGQLEQQEGQVEHLEQQVEHLEQLGQLKHLEEQE 413
Query: 399 -QSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGE----EEEHEQEEEE 453
Q K E + LG VG+P N +++ E ++ EG+ E++ Q E
Sbjct: 414 GQLKHLEQQQGQLGVPEQVGQPK-----NLEQEEKQLELPEQQEGQLKHLEKQEAQLELP 468
Query: 454 EDDVEEDEH 462
E V + +H
Sbjct: 469 EQQVGQPKH 477
>MNN4_YEAST (P36044) MNN4 protein
Length = 1178
Score = 52.8 bits (125), Expect = 3e-06
Identities = 44/161 (27%), Positives = 80/161 (49%), Gaps = 9/161 (5%)
Query: 303 LFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQV 362
++E+ ++ E + K +EEE + K EEEE+ +++ + K EE +
Sbjct: 1024 VYEDYAYAKLLEERKRREKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKK 1083
Query: 363 QELEEHNIELSLGQDKVETLPVEKEQGEGEQMM-DFEQSKKEETEMWFLGQKNYVGEPSL 421
++ EE + + K + +K+Q EGE+M + E++KK E E +K E
Sbjct: 1084 KKEEEEKKKQEEEEKKKKEEEEKKKQEEGEKMKNEDEENKKNEDE-----EKKKNEEEEK 1138
Query: 422 RPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEH 462
+ +++K D E+ K+ EEEE ++ EEEE +E+ H
Sbjct: 1139 KKQEEKNKKNEDEEKKKQ---EEEEKKKNEEEEKKKQEEGH 1176
Score = 43.5 bits (101), Expect = 0.002
Identities = 44/177 (24%), Positives = 74/177 (40%), Gaps = 31/177 (17%)
Query: 300 HKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEE 359
+ +L EE E + + EE K EEEE + K+EEE++ + + + K EE
Sbjct: 1030 YAKLLEERKRREKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEE 1089
Query: 360 NQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEP 419
+ QE EE K + +K+Q EGE+M + ++ K+
Sbjct: 1090 KKKQEEEEK---------KKKEEEEKKKQEEGEKMKNEDEENKK---------------- 1124
Query: 420 SLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHDVGFHFSTKHHLEG 476
N D + E+ ++ + EE+ + E+EE+ EE+E K EG
Sbjct: 1125 ------NEDEEKKKNEEEEKKKQEEKNKKNEDEEKKKQEEEEKKKNEEEEKKKQEEG 1175
Score = 32.7 bits (73), Expect = 3.1
Identities = 32/134 (23%), Positives = 66/134 (48%), Gaps = 7/134 (5%)
Query: 275 EKELKATHLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEE 334
E+E K EE + K + +E ++ + + E EE ++EEE+ + ++EE
Sbjct: 1046 EEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKKEEEEKKKQEEEEKKKKEEEE 1105
Query: 335 --EEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGE 392
++E+G +K+E D + E+ + ++ EE + ++K +K+Q E E
Sbjct: 1106 KKKQEEGEKMKNE-----DEENKKNEDEEKKKNEEEEKKKQEEKNKKNEDEEKKKQEEEE 1160
Query: 393 QMMDFEQSKKEETE 406
+ + E+ KK++ E
Sbjct: 1161 KKKNEEEEKKKQEE 1174
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 86,081,688
Number of Sequences: 164201
Number of extensions: 4434923
Number of successful extensions: 70503
Number of sequences better than 10.0: 1465
Number of HSP's better than 10.0 without gapping: 682
Number of HSP's successfully gapped in prelim test: 832
Number of HSP's that attempted gapping in prelim test: 36827
Number of HSP's gapped (non-prelim): 11366
length of query: 652
length of database: 59,974,054
effective HSP length: 117
effective length of query: 535
effective length of database: 40,762,537
effective search space: 21807957295
effective search space used: 21807957295
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC144893.11


