

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
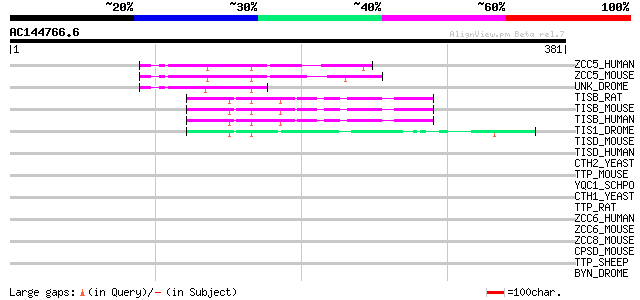
Query= AC144766.6 - phase: 0
(381 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ZCC5_HUMAN (Q9C0B0) Zinc finger CCCH type domain containing prot... 67 6e-11
ZCC5_MOUSE (Q8BL48) Zinc finger CCCH type domain containing prot... 66 1e-10
UNK_DROME (Q86B79) Unkempt protein 61 5e-09
TISB_RAT (P17431) Butyrate response factor 1 (TIS11B protein) (E... 52 3e-06
TISB_MOUSE (P23950) Butyrate response factor 1 (TIS11B protein) 52 3e-06
TISB_HUMAN (Q07352) Butyrate response factor 1 (TIS11B protein) ... 52 3e-06
TIS1_DROME (P47980) TIS11 protein (dTIS11) 44 9e-04
TISD_MOUSE (P23949) Butyrate response factor 2 (TIS11D protein) 41 0.004
TISD_HUMAN (P47974) Butyrate response factor 2 (TIS11D protein) ... 40 0.008
CTH2_YEAST (P47977) Zinc finger protein CTH2 (YTIS11 protein) 40 0.008
TTP_MOUSE (P22893) Tristetraproline (TTP) (Zinc finger protein 3... 40 0.010
YQC1_SCHPO (O74463) Hypothetical protein C1739.01 in chromosome III 39 0.017
CTH1_YEAST (P47976) Zinc finger protein CTH1 39 0.017
TTP_RAT (P47973) Tristetraproline (TTP) (Zinc finger protein 36)... 39 0.022
ZCC6_HUMAN (P61129) Zinc finger CCCH type domain containing prot... 39 0.029
ZCC6_MOUSE (Q8BYK8) Zinc finger CCCH type domain containing prot... 38 0.037
ZCC8_MOUSE (Q9JJ48) Zinc finger CCCH type domain containing prot... 37 0.064
CPSD_MOUSE (Q8BQZ5) Cleavage and polyadenylation specificity fac... 37 0.083
TTP_SHEEP (Q6S9E0) Tristetraproline (TTP) (Zinc finger protein 3... 36 0.14
BYN_DROME (P55965) T-related protein (Trp) (Brachyenteron protein) 36 0.19
>ZCC5_HUMAN (Q9C0B0) Zinc finger CCCH type domain containing protein
5
Length = 810
Score = 67.4 bits (163), Expect = 6e-11
Identities = 53/174 (30%), Positives = 72/174 (40%), Gaps = 33/174 (18%)
Query: 90 RRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGS-------CKKGDSCE 142
R C +G + CP+ H K RRR PRK+ Y + CP+ + G C+ GD+C+
Sbjct: 226 RLCRQGYA-----CPYYH-NSKDRRRSPRKHKYRSSPCPNVKHGDEWGDPGKCENGDACQ 279
Query: 143 FAHGVFECWLHPSRYRTQPCKD----GTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDS 198
+ H E HP Y++ C D G+ R P C FAH EQ P S S +
Sbjct: 280 YCHTRTEQQFHPEIYKSTKCNDMQQSGSCPRGPFCAFAH-VEQPPLSDDLQPSSAVSSPT 338
Query: 199 YDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSL---GRSVGSSS 249
G L S+ G P S P +P +L S+GS S
Sbjct: 339 QPG------------PVLYMPSAAGDSVPVSPSSPHAPDLSALLCRNSSLGSPS 380
Score = 37.0 bits (84), Expect = 0.083
Identities = 36/121 (29%), Positives = 48/121 (38%), Gaps = 21/121 (17%)
Query: 86 EFKIRRCARGRSHDWTE-----CPFSHPGEKARRRDPRK----YNYS-GTSCPDFRK--G 133
EF+ +C H T+ C H + RRR R+ +NYS C + + G
Sbjct: 39 EFRTEQCPLFVQHKCTQHRPYTCFHWHFVNQRRRRSIRRRDGTFNYSPDVYCTKYDEATG 98
Query: 134 SCKKGDSCEFAH---GVFECWLHPSRYRTQPCKDGTSCRRPV------CFFAHTTEQLRA 184
C +GD C F H G E H Y+T C T + C FAH LR+
Sbjct: 99 LCPEGDECPFLHRTTGDTERRYHLRYYKTGICIHETDSKGNCTKNGLHCAFAHGPHDLRS 158
Query: 185 P 185
P
Sbjct: 159 P 159
>ZCC5_MOUSE (Q8BL48) Zinc finger CCCH type domain containing protein
5
Length = 810
Score = 66.2 bits (160), Expect = 1e-10
Identities = 53/180 (29%), Positives = 75/180 (41%), Gaps = 33/180 (18%)
Query: 90 RRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGS-------CKKGDSCE 142
R C +G + CP+ H K RRR PRK+ Y + CP+ + G C+ GD+C+
Sbjct: 226 RLCRQGYA-----CPYYH-NSKDRRRSPRKHKYRSSPCPNVKHGDEWGDPGKCENGDACQ 279
Query: 143 FAHGVFECWLHPSRYRTQPCKD----GTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDS 198
+ H E HP Y++ C D G+ R P C FAH E P S S +
Sbjct: 280 YCHTRTEQQFHPEIYKSTKCNDMQQAGSCPRGPFCAFAH-IEPPPLSDDVQPSSAVSSPT 338
Query: 199 YDGSPLRLAFESSCVKTLQFMSSPGSVSPPV--ESPPMSPMTRSLGRSVGSSSVNEMVAS 256
G L +M S S PV SP ++ L R+ G S + + +S
Sbjct: 339 QPGPVL-------------YMPSAAGDSVPVSPSSPHAPDLSALLCRNSGLGSPSHLCSS 385
Score = 37.0 bits (84), Expect = 0.083
Identities = 36/121 (29%), Positives = 48/121 (38%), Gaps = 21/121 (17%)
Query: 86 EFKIRRCARGRSHDWTE-----CPFSHPGEKARRRDPRK----YNYS-GTSCPDFRK--G 133
EF+ +C H T+ C H + RRR R+ +NYS C + + G
Sbjct: 39 EFRTEQCPLFVQHKCTQHRPYTCFHWHFVNQRRRRSIRRRDGTFNYSPDVYCTKYDEATG 98
Query: 134 SCKKGDSCEFAH---GVFECWLHPSRYRTQPCKDGTSCRRPV------CFFAHTTEQLRA 184
C +GD C F H G E H Y+T C T + C FAH LR+
Sbjct: 99 LCPEGDECPFLHRTTGDTERRYHLRYYKTGICIHETDSKGNCTKNGLHCAFAHGPHDLRS 158
Query: 185 P 185
P
Sbjct: 159 P 159
>UNK_DROME (Q86B79) Unkempt protein
Length = 599
Score = 60.8 bits (146), Expect = 5e-09
Identities = 35/99 (35%), Positives = 47/99 (47%), Gaps = 17/99 (17%)
Query: 90 RRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKG-------SCKKGDSCE 142
R C +G + CP H K +RR PRKY Y T CP+ + G +C+ GD+C+
Sbjct: 205 RLCRQGYA-----CPQYH-NSKDKRRSPRKYKYRSTPCPNVKHGEEWGEPGNCEAGDNCQ 258
Query: 143 FAHGVFECWLHPSRYRTQPCKD----GTSCRRPVCFFAH 177
+ H E HP Y++ C D G R C FAH
Sbjct: 259 YCHTRTEQQFHPEIYKSTKCNDVQQAGYCPRSVFCAFAH 297
Score = 38.1 bits (87), Expect = 0.037
Identities = 36/128 (28%), Positives = 51/128 (39%), Gaps = 21/128 (16%)
Query: 79 NDHFRMFEFKIRRCARGRSHDWTE-----CPFSHPGEKARRRDPRK----YNYSGTS-CP 128
N + + EF++ +C H + C H + RRR RK +NYS + C
Sbjct: 19 NHYTYLKEFRVEQCQSFLQHKCNQHRPFVCFNWHFQNQRRRRPVRKRDGTFNYSADNYCT 78
Query: 129 DFRK--GSCKKGDSCEFAH---GVFECWLHPSRYRTQPCKDGTSCRRPV------CFFAH 177
+ + G C +GD C + H G E H Y+T C T R C FAH
Sbjct: 79 KYDETTGICPEGDECPYLHRTAGDTERRYHLRYYKTCMCVHDTDSRGYCVKNGLHCAFAH 138
Query: 178 TTEQLRAP 185
+ R P
Sbjct: 139 GMQDQRPP 146
>TISB_RAT (P17431) Butyrate response factor 1 (TIS11B protein)
(EGF-inducible protein CMG1)
Length = 338
Score = 51.6 bits (122), Expect = 3e-06
Identities = 54/184 (29%), Positives = 75/184 (40%), Gaps = 31/184 (16%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G+CK GD C+FAHG+ E HP +Y+T+ C+ G P C
Sbjct: 115 YKTELCRPFEENGACKYGDKCQFAHGIHELRSLTRHP-KYKTELCRTFHTIGFCPYGPRC 173
Query: 174 FFAHTTEQLRA------PTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSP 227
F H E+ RA + PR S S+ G P A ++ + SP S++P
Sbjct: 174 HFIHNAEERRALAGGRDLSADRPRLQHSF-SFAGFPSAAATAAA----TGLLDSPTSITP 228
Query: 228 PVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGP 287
PP+ LG N A + + L S + MG P GSP
Sbjct: 229 ----PPILSADDLLGSPTLPDGTNNPFAF-------SSQELASLFAPSMGLPGGGSPTTF 277
Query: 288 VIRP 291
+ RP
Sbjct: 278 LFRP 281
>TISB_MOUSE (P23950) Butyrate response factor 1 (TIS11B protein)
Length = 338
Score = 51.6 bits (122), Expect = 3e-06
Identities = 54/184 (29%), Positives = 75/184 (40%), Gaps = 31/184 (16%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G+CK GD C+FAHG+ E HP +Y+T+ C+ G P C
Sbjct: 115 YKTELCRPFEENGACKYGDKCQFAHGIHELRSLTRHP-KYKTELCRTFHTIGFCPYGPRC 173
Query: 174 FFAHTTEQLRA------PTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSP 227
F H E+ RA + PR S S+ G P A ++ + SP S++P
Sbjct: 174 HFIHNAEERRALAGGRDLSADRPRLQHSF-SFAGFPSAAATAAA----TGLLDSPTSITP 228
Query: 228 PVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGP 287
PP+ LG N A + + L S + MG P GSP
Sbjct: 229 ----PPILSADDLLGSPTLPDGTNNPFAF-------SSQELASLFAPSMGLPGGGSPTTF 277
Query: 288 VIRP 291
+ RP
Sbjct: 278 LFRP 281
>TISB_HUMAN (Q07352) Butyrate response factor 1 (TIS11B protein)
(EGF-response factor 1) (ERF-1)
Length = 338
Score = 51.6 bits (122), Expect = 3e-06
Identities = 54/184 (29%), Positives = 75/184 (40%), Gaps = 31/184 (16%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G+CK GD C+FAHG+ E HP +Y+T+ C+ G P C
Sbjct: 115 YKTELCRPFEENGACKYGDKCQFAHGIHELRSLTRHP-KYKTELCRTFHTIGFCPYGPRC 173
Query: 174 FFAHTTEQLRA------PTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSP 227
F H E+ RA + PR S S+ G P A ++ + SP S++P
Sbjct: 174 HFIHNAEERRALAGARDLSADRPRLQHSF-SFAGFPSAAATAAA----TGLLDSPTSITP 228
Query: 228 PVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGP 287
PP+ LG N A + + L S + MG P GSP
Sbjct: 229 ----PPILSADDLLGSPTLPDGTNNPFAF-------SSQELASLFAPSMGLPGGGSPTTF 277
Query: 288 VIRP 291
+ RP
Sbjct: 278 LFRP 281
>TIS1_DROME (P47980) TIS11 protein (dTIS11)
Length = 437
Score = 43.5 bits (101), Expect = 9e-04
Identities = 62/250 (24%), Positives = 95/250 (37%), Gaps = 51/250 (20%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G CK G+ C+FAHG E HP +Y+T+ C+ G P C
Sbjct: 137 YKTELCRPFEEAGECKYGEKCQFAHGSHELRNVHRHP-KYKTEYCRTFHSVGFCPYGPRC 195
Query: 174 FFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPP 233
F H ++ RA QQ+ ++ S + + ++ K+ Q + + S
Sbjct: 196 HFVHNADEARA--QQAAQAAKSSTQSQSQSQQSSSQNFSPKSNQSSNQSSNSS------- 246
Query: 234 MSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGF 293
S + S G G +S+N S L L S+ + GS R SP G
Sbjct: 247 -SSSSSSGGGGGGGNSINNNNGSQFYLPLSPPLSMST------GSDR-ESPTG------- 291
Query: 294 CSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESG--RDIRVKMFEKLSKENSFNGS 351
SL +PT S P + ++ G K S +S +G
Sbjct: 292 -SLSLSPT---------------NSLTSFPFHDALQHGYLASNGAKSNSSASSTSSASGM 335
Query: 352 GMGSGSGLGE 361
G+G G+G+
Sbjct: 336 GLGMSMGIGQ 345
>TISD_MOUSE (P23949) Butyrate response factor 2 (TIS11D protein)
Length = 367
Score = 41.2 bits (95), Expect = 0.004
Identities = 48/165 (29%), Positives = 69/165 (41%), Gaps = 17/165 (10%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G+CK G+ C+FAHG E HP +Y+T+ C+ G P C
Sbjct: 127 YKTELCRPFEESGTCKYGEKCQFAHGFHELRSLTRHP-KYKTELCRTFHTIGFCPYGPRC 185
Query: 174 FFAHTTEQLR-APTQQSPRS--VPSVDSYDGSPLRLAFE--SSCVKTLQFMSSP-GSVSP 227
F H ++ R AP+ S + + + D L A E +L F P G P
Sbjct: 186 HFIHNADERRPAPSGGGGASGDLRAFGARDALHLGFAREPRPKLHHSLSFSGFPSGHHQP 245
Query: 228 P--VESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPS 270
P +ESP + S SSS + +S + + S PS
Sbjct: 246 PGGLESPLLLDSPTSRTPPPPSSSASSCSSSASSCSSASAASTPS 290
>TISD_HUMAN (P47974) Butyrate response factor 2 (TIS11D protein)
(EGF-response factor 2) (ERF-2)
Length = 492
Score = 40.4 bits (93), Expect = 0.008
Identities = 42/137 (30%), Positives = 62/137 (44%), Gaps = 21/137 (15%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCKD----GTSCRRPVC 173
Y C F + G+CK G+ C+FAHG E HP +Y+T+ C+ G P C
Sbjct: 154 YKTELCRPFEESGTCKYGEKCQFAHGFHELRSLTRHP-KYKTELCRTFHTIGFCPYGPRC 212
Query: 174 FFAHTTEQLR-APTQQSPRSVPSVDSYDGSPLRLAF----ESSCVKTLQFMSSP-GSVSP 227
F H ++ R AP+ + + + + D L L F +L F P G P
Sbjct: 213 HFIHNADERRPAPSGGASGDLRAFGTRDA--LHLGFPREPRPKLHHSLSFSGFPSGHHQP 270
Query: 228 P--VESPPM--SPMTRS 240
P +ESP + SP +R+
Sbjct: 271 PGGLESPLLLDSPTSRT 287
>CTH2_YEAST (P47977) Zinc finger protein CTH2 (YTIS11 protein)
Length = 285
Score = 40.4 bits (93), Expect = 0.008
Identities = 22/60 (36%), Positives = 32/60 (52%), Gaps = 3/60 (5%)
Query: 106 SHPGEKARRRDPRKYNYSGTSCPDFR-KGSCKKGDSCEFAHGVFECWLHPS--RYRTQPC 162
+ P K++ ++ K Y C F KGSC G C+FAHG+ E + S +RT+PC
Sbjct: 154 AQPKVKSQVQETPKQLYKTELCESFTLKGSCPYGSKCQFAHGLGELKVKKSCKNFRTKPC 213
>TTP_MOUSE (P22893) Tristetraproline (TTP) (Zinc finger protein 36)
(Zfp-36) (TIS11A protein) (TIS11) (Growth
factor-inducible nuclear protein NUP475) (TPA induced
sequence 11)
Length = 319
Score = 40.0 bits (92), Expect = 0.010
Identities = 38/119 (31%), Positives = 49/119 (40%), Gaps = 20/119 (16%)
Query: 133 GSCKKGDSCEFAHGVFE---CWLHPSRYRTQPCK----DGTSCRRPVCFFAHT-TEQLRA 184
G C+ G C+FAHG+ E HP +Y+T+ C G C F H TE L
Sbjct: 108 GRCRYGAKCQFAHGLGELRQANRHP-KYKTELCHKFYLQGRCPYGSRCHFIHNPTEDLAL 166
Query: 185 PTQQSPRSVPSVDSYDG-------SPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSP 236
P Q P + S+ G SP F + + F SP S PP P+SP
Sbjct: 167 PGQ--PHVLRQSISFSGLPSGRRSSPPPPGFSGPSLSSCSF--SPSSSPPPPGDLPLSP 221
Score = 30.4 bits (67), Expect = 7.8
Identities = 40/167 (23%), Positives = 60/167 (34%), Gaps = 22/167 (13%)
Query: 95 GRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFR-KGSCKKGDSCEFAHGVFECWL- 152
GR +C F+H G R+ R Y C F +G C G C F H E
Sbjct: 108 GRCRYGAKCQFAH-GLGELRQANRHPKYKTELCHKFYLQGRCPYGSRCHFIHNPTEDLAL 166
Query: 153 ---------------HPSRYRTQPCKDGTS-CRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
PS R+ P G S C F+ ++ P P S +
Sbjct: 167 PGQPHVLRQSISFSGLPSGRRSSPPPPGFSGPSLSSCSFSPSSSP--PPPGDLPLSPSAF 224
Query: 197 DSYDGSPL-RLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLG 242
+ G+P+ R +C + + ++P ++ P+ SP SLG
Sbjct: 225 SAAPGTPVTRRDPNQACCPSCRRSTTPSTIWGPLGGLARSPSAHSLG 271
>YQC1_SCHPO (O74463) Hypothetical protein C1739.01 in chromosome III
Length = 547
Score = 39.3 bits (90), Expect = 0.017
Identities = 46/207 (22%), Positives = 77/207 (36%), Gaps = 24/207 (11%)
Query: 121 NYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTE 180
N C FR G+C G++C F+H + + CK G P C +H
Sbjct: 41 NLQHVPCKFFRNGTCTAGENCPFSHSLETERPICKYFLKGNCKFG-----PKCALSHALP 95
Query: 181 -QLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPP----------- 228
P S ++ S+ + G+ + + + +SS +P
Sbjct: 96 GNTNLPNGTSTNTMASMAANGGASSVASKQMGANQISPSLSSKTMKNPADKANNTTATDV 155
Query: 229 ---VESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLG-TMKSLPSSWNVQMGSPRFGSP 284
+ P P +RS GR G+S++N M+ + L G +S+ N G P S
Sbjct: 156 RGNTATSPYFPFSRSPGRHSGNSTINGMMTTPNFLSSGVNSRSVDEFNNSSSGFP--SSL 213
Query: 285 RG-PVIRPGFCSLPSTPTQVPSRGRVN 310
G P+ P + P++ + S N
Sbjct: 214 NGIPIASPPLATSPTSFSLASSASSTN 240
>CTH1_YEAST (P47976) Zinc finger protein CTH1
Length = 325
Score = 39.3 bits (90), Expect = 0.017
Identities = 25/77 (32%), Positives = 35/77 (44%), Gaps = 15/77 (19%)
Query: 101 TECPFSHPGEKARRRDP------------RKYNYSGTSCPDFR-KGSCKKGDSCEFAHGV 147
T P++ P +K + +P K Y C F KG CK G+ C+FAHG+
Sbjct: 172 TGLPYTLPIQKTTKLEPCRRAPLQLPQLVNKTLYKTELCESFTIKGYCKYGNKCQFAHGL 231
Query: 148 FECWL--HPSRYRTQPC 162
E + YRT+PC
Sbjct: 232 NELKFKKKSNNYRTKPC 248
>TTP_RAT (P47973) Tristetraproline (TTP) (Zinc finger protein 36)
(Zfp-36) (TIS11A protein) (TIS11)
Length = 320
Score = 38.9 bits (89), Expect = 0.022
Identities = 37/119 (31%), Positives = 49/119 (41%), Gaps = 20/119 (16%)
Query: 133 GSCKKGDSCEFAHG---VFECWLHPSRYRTQPCK----DGTSCRRPVCFFAHT-TEQLRA 184
G C+ G C+FAHG + + HP +Y+T+ C G C F H TE L
Sbjct: 109 GRCRYGAKCQFAHGPGELRQANRHP-KYKTELCHKFYLQGRCPYGSRCHFIHNPTEDLAL 167
Query: 185 PTQQSPRSVPSVDSYDG-------SPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSP 236
P Q P + S+ G SP F + + F SP S PP P+SP
Sbjct: 168 PGQ--PHVLRQSISFSGLPSGRRTSPPPPGFSGPSLSSCSF--SPSSSPPPPGDLPLSP 222
Score = 33.1 bits (74), Expect = 1.2
Identities = 43/168 (25%), Positives = 62/168 (36%), Gaps = 24/168 (14%)
Query: 95 GRSHDWTECPFSH-PGEKARRRDPRKYNYSGTSCPDFR-KGSCKKGDSCEFAHGVFECWL 152
GR +C F+H PGE R+ R Y C F +G C G C F H E
Sbjct: 109 GRCRYGAKCQFAHGPGEL--RQANRHPKYKTELCHKFYLQGRCPYGSRCHFIHNPTEDLA 166
Query: 153 ----------------HPSRYRTQPCKDGTS-CRRPVCFFAHTTEQLRAPTQQSPRSVPS 195
PS RT P G S C F+ ++ P P S +
Sbjct: 167 LPGQPHVLRQSISFSGLPSGRRTSPPPPGFSGPSLSSCSFSPSSSP--PPPGDLPLSPSA 224
Query: 196 VDSYDGSPL-RLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLG 242
+ G+P+ R +C + + ++P ++ P+ SP SLG
Sbjct: 225 FSAAPGTPVSRRDPTPACCPSCRRSTTPSTIWGPLGGLARSPSAHSLG 272
>ZCC6_HUMAN (P61129) Zinc finger CCCH type domain containing protein
6
Length = 1189
Score = 38.5 bits (88), Expect = 0.029
Identities = 21/79 (26%), Positives = 35/79 (43%), Gaps = 17/79 (21%)
Query: 91 RCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFEC 150
RC +G +C F H E +R++ C + +G C KG++C + H F C
Sbjct: 285 RCIKG-----DQCKFDHDAELEKRKE---------ICKFYLQGYCTKGENCIYMHNEFPC 330
Query: 151 WLHPSRYRTQPCKDGTSCR 169
+ S + C G +C+
Sbjct: 331 KFYHSGAK---CYQGDNCK 346
>ZCC6_MOUSE (Q8BYK8) Zinc finger CCCH type domain containing protein
6
Length = 1177
Score = 38.1 bits (87), Expect = 0.037
Identities = 21/79 (26%), Positives = 35/79 (43%), Gaps = 17/79 (21%)
Query: 91 RCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFEC 150
RC +G C F+H E ++++ KY + +G C KG++C + H F C
Sbjct: 283 RCIKG-----DHCKFNHDAELEKKKEVCKY---------YLQGYCTKGENCIYMHSEFPC 328
Query: 151 WLHPSRYRTQPCKDGTSCR 169
+ S + C G C+
Sbjct: 329 KFYHSGAK---CYQGDKCK 344
>ZCC8_MOUSE (Q9JJ48) Zinc finger CCCH type domain containing protein
8 (Fetal liver zinc finger protein 1)
Length = 305
Score = 37.4 bits (85), Expect = 0.064
Identities = 29/112 (25%), Positives = 47/112 (41%), Gaps = 26/112 (23%)
Query: 87 FKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHG 146
F R+C +G +C F H E ++++ KY + +G C KG++C + H
Sbjct: 214 FLERKCIKG-----DQCKFDHDAEIEKKKEMCKY---------YVQGYCTKGENCLYLHS 259
Query: 147 VFECWLHPSRYRTQPCKDGTSC-RRPVCFFAHTTEQLRAPTQQSPRSVPSVD 197
+ C + + GT C + C F+H L A TQ+ V D
Sbjct: 260 EYPCKFYHT---------GTKCYQGDHCNFSHA--PLTAETQELLAKVLDTD 300
>CPSD_MOUSE (Q8BQZ5) Cleavage and polyadenylation specificity
factor, 30 kDa subunit (CPSF 30 kDa subunit) (Clipper
homolog) (Clipper/CPSF 30K)
Length = 211
Score = 37.0 bits (84), Expect = 0.083
Identities = 34/118 (28%), Positives = 41/118 (33%), Gaps = 47/118 (39%)
Query: 103 CPFSH-PGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAH-----GVFECWLHPS- 155
CPF H GEK C + +G CKKGD CEF H + EC+ +
Sbjct: 55 CPFRHISGEKT------------VVCKHWLRGLCKKGDQCEFLHEYDMTKMPECYFYSKF 102
Query: 156 ---------------RYRTQPCKDGTSCRRPVCFFAH--------TTEQLRAPTQQSP 190
Y C +G SC+ F H TTEQ P Q P
Sbjct: 103 GPLCRHRHTRRVICVNYLVGFCPEGPSCK-----FMHPRFELPMGTTEQPPLPQQTQP 155
>TTP_SHEEP (Q6S9E0) Tristetraproline (TTP) (Zinc finger protein 36
homolog) (Zfp-36)
Length = 325
Score = 36.2 bits (82), Expect = 0.14
Identities = 54/202 (26%), Positives = 72/202 (34%), Gaps = 32/202 (15%)
Query: 122 YSGTSCPDFRK-GSCKKGDSCEFAHGVFECWLHPSR---YRTQPCK----DGTSCRRPVC 173
Y C F + G C+ G C+FAHG+ E PSR Y+T+ C G C
Sbjct: 102 YKTELCRTFSESGRCRYGAKCQFAHGLGEL-RQPSRHPKYKTELCHKFYLQGRCPYGSRC 160
Query: 174 FFAHT-TEQLRAPTQQSPRSVPSVDSYDG-------SPLRLAFESSCVKTLQFMSSPGSV 225
F H +E L AP P + S+ G SP + V + F S S
Sbjct: 161 HFIHNPSEDLAAPGH--PHVLRQSISFSGLPSGRRTSPPPASLAGPSVPSWSF-SPSSSP 217
Query: 226 SPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPR 285
PP P+SP S S + A + + T S+ +G
Sbjct: 218 PPPPGDLPLSPSAFSAAPGTPVSRRDPTPACCPSCRRATPNSV------------WGPVG 265
Query: 286 GPVIRPGFCSLPSTPTQVPSRG 307
G P SL S P + S G
Sbjct: 266 GLARSPSAHSLGSDPDEYASSG 287
>BYN_DROME (P55965) T-related protein (Trp) (Brachyenteron protein)
Length = 697
Score = 35.8 bits (81), Expect = 0.19
Identities = 40/155 (25%), Positives = 61/155 (38%), Gaps = 34/155 (21%)
Query: 184 APTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGR 243
APT+ +P S P V S GS S G+ S PP +P T +
Sbjct: 344 APTRTTPYSRPRVVSGSGSN----------------GSAGNASSTSPQPPSAPQTPTSLH 387
Query: 244 SVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTPT-- 301
S + SV+ V+S +G+ +P G + GF S+ ++PT
Sbjct: 388 STSTGSVSTSVSSSSGGGIGS-------------APSTGCFSSSYAQSGFMSVDASPTAS 434
Query: 302 --QVPSRGRVNHFDLWDQSCEEEPVMERVESGRDI 334
PS + N + W+ + P+ V SGR+I
Sbjct: 435 VFSYPSSWQSNG-NYWNATSVPGPMPMNVCSGRNI 468
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.133 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,826,956
Number of Sequences: 164201
Number of extensions: 2316709
Number of successful extensions: 7509
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 7442
Number of HSP's gapped (non-prelim): 90
length of query: 381
length of database: 59,974,054
effective HSP length: 112
effective length of query: 269
effective length of database: 41,583,542
effective search space: 11185972798
effective search space used: 11185972798
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144766.6


