

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.5 - phase: 0 /pseudo
(443 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
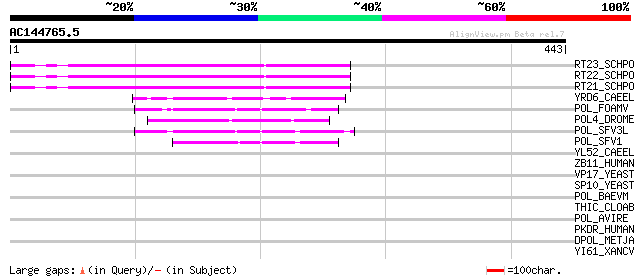
Score E
Sequences producing significant alignments: (bits) Value
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 137 6e-32
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 137 6e-32
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 137 6e-32
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 69 2e-11
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 60 1e-08
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 58 4e-08
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 56 2e-07
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 50 9e-06
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 41 0.005
ZB11_HUMAN (O95625) Zinc finger and BTB domain containing protei... 37 0.078
VP17_YEAST (P32913) Vacuolar protein sorting-associated protein ... 34 0.86
SP10_YEAST (P35208) SPT10 protein 34 0.86
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 34 0.86
THIC_CLOAB (Q97EU2) Thiamine biosynthesis protein thiC 33 1.9
POL_AVIRE (P03360) Pol polyprotein [Contains: Reverse transcript... 32 2.5
PKDR_HUMAN (Q9NTG1) Polycystic kidney disease and receptor for e... 32 2.5
DPOL_METJA (Q58295) DNA polymerase (EC 2.7.7.7) [Contains: Mja p... 31 7.3
YI61_XANCV (P25438) Insertion element IS476 hypothetical 39.2 kD... 30 9.5
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 137 bits (345), Expect = 6e-32
Identities = 77/273 (28%), Positives = 143/273 (52%), Gaps = 18/273 (6%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQ-LGMLK 59
NF +NY P A +ADALSR +V E E + + + S+ + +
Sbjct: 812 NFEINYRPGSANHIADALSR--------IVDETEPIPK--------DSEDNSINFVNQIS 855
Query: 60 INSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLI 119
I DF + D K ++L+ ++ E + ++ + ++ + +I +P++ +L + I
Sbjct: 856 ITDDFKNQVVTEYTNDTKLLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTI 915
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
+++ H+ IH G + + + F W G++K + +V C TCQ +K + +P G L
Sbjct: 916 IKKYHEEGKLIHPGIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQ 975
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYI 239
P+ E W+S+SMDF+++LP +S G+++++VVVDR +K A +P S Q A ++
Sbjct: 976 PIPPSERPWESLSMDFITALPESS-GYNALFVVVDRFSKMAILVPCTKSITAEQTARMFD 1034
Query: 240 QNIVKLHGVPSSIVSDRDPRFTSRFWRSLQMRW 272
Q ++ G P I++D D FTS+ W+ ++
Sbjct: 1035 QRVIAYFGNPKEIIADNDHIFTSQTWKDFAHKY 1067
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 137 bits (345), Expect = 6e-32
Identities = 77/273 (28%), Positives = 143/273 (52%), Gaps = 18/273 (6%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQ-LGMLK 59
NF +NY P A +ADALSR +V E E + + + S+ + +
Sbjct: 812 NFEINYRPGSANHIADALSR--------IVDETEPIPK--------DSEDNSINFVNQIS 855
Query: 60 INSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLI 119
I DF + D K ++L+ ++ E + ++ + ++ + +I +P++ +L + I
Sbjct: 856 ITDDFKNQVVTEYTNDTKLLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTI 915
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
+++ H+ IH G + + + F W G++K + +V C TCQ +K + +P G L
Sbjct: 916 IKKYHEEGKLIHPGIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQ 975
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYI 239
P+ E W+S+SMDF+++LP +S G+++++VVVDR +K A +P S Q A ++
Sbjct: 976 PIPPSERPWESLSMDFITALPESS-GYNALFVVVDRFSKMAILVPCTKSITAEQTARMFD 1034
Query: 240 QNIVKLHGVPSSIVSDRDPRFTSRFWRSLQMRW 272
Q ++ G P I++D D FTS+ W+ ++
Sbjct: 1035 QRVIAYFGNPKEIIADNDHIFTSQTWKDFAHKY 1067
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 137 bits (345), Expect = 6e-32
Identities = 77/273 (28%), Positives = 143/273 (52%), Gaps = 18/273 (6%)
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQ-LGMLK 59
NF +NY P A +ADALSR +V E E + + + S+ + +
Sbjct: 812 NFEINYRPGSANHIADALSR--------IVDETEPIPK--------DSEDNSINFVNQIS 855
Query: 60 INSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLI 119
I DF + D K ++L+ ++ E + ++ + ++ + +I +P++ +L + I
Sbjct: 856 ITDDFKNQVVTEYTNDTKLLNLLNNEDKRVEENIQLKDGLLINSKDQILLPNDTQLTRTI 915
Query: 120 LEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLT 179
+++ H+ IH G + + + F W G++K + +V C TCQ +K + +P G L
Sbjct: 916 IKKYHEEGKLIHPGIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQ 975
Query: 180 PLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYI 239
P+ E W+S+SMDF+++LP +S G+++++VVVDR +K A +P S Q A ++
Sbjct: 976 PIPPSERPWESLSMDFITALPESS-GYNALFVVVDRFSKMAILVPCTKSITAEQTARMFD 1034
Query: 240 QNIVKLHGVPSSIVSDRDPRFTSRFWRSLQMRW 272
Q ++ G P I++D D FTS+ W+ ++
Sbjct: 1035 QRVIAYFGNPKEIIADNDHIFTSQTWKDFAHKY 1067
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 68.9 bits (167), Expect = 2e-11
Identities = 50/172 (29%), Positives = 81/172 (47%), Gaps = 19/172 (11%)
Query: 99 GVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFV 158
G L R+ +P + L+K++L++ H+ H G +M Q + +W GL D+ V
Sbjct: 768 GCLLLDDRVIVP--KSLQKIVLKQLHEG----HPGIVQMKQKARSFVFWRGLDSDIENMV 821
Query: 159 YACLTCQK-SKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLT 217
C CQ+ SK+ P L P VPE W I +DF L + VVVD T
Sbjct: 822 RHCNNCQENSKMPRVVP---LNPWPVPEAPWKRIHIDFAGPLNGC-----YLLVVVDAKT 873
Query: 218 KSAHFIPINISYPVAQLAEI-YIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
K A + ++ ++ + I ++ I +HG P +I+SD + TS + +
Sbjct: 874 KYAE---VKLTRSISAVTTIDLLEEIFSIHGYPETIISDNGTQLTSHLFAQM 922
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 59.7 bits (143), Expect = 1e-08
Identities = 44/163 (26%), Positives = 75/163 (45%), Gaps = 9/163 (5%)
Query: 100 VLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVY 159
V R G IP + +K++L+ NL+ H G + L+WW ++KDV + +
Sbjct: 593 VSRPEGVKIIPPQSDRQKIVLQA---HNLA-HTGREATLLKIANLYWWPNMRKDVVKQLG 648
Query: 160 ACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKS 219
C C + ++ +L P D P+ +D +D++ LP S+G+ + VVVD +T
Sbjct: 649 RCQQCLITNASNKASGPILRP-DRPQKPFDKFFIDYIGPLP-PSQGYLYVLVVVDGMTGF 706
Query: 220 AHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTS 262
P A + + N++ +P I SD+ FTS
Sbjct: 707 TWLYPTKAPSTSATVKSL---NVLTSIAIPKVIHSDQGAAFTS 746
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-); Reverse
transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 58.2 bits (139), Expect = 4e-08
Identities = 37/147 (25%), Positives = 72/147 (48%), Gaps = 5/147 (3%)
Query: 111 DNEELKKLILEEGHKSNLSI-HLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSK- 168
+NE+ K+ IL H + H G TK +K+ ++W + K + +V C CQK+K
Sbjct: 888 NNEKEKEAILSTLHDDPIQGGHTGITKTLAKVKRHYYWKNMSKYIKEYVRKCQKCQKAKT 947
Query: 169 VEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINIS 228
+H + +T + PE +D + +D + LP + G++ ++ LTK IPI +
Sbjct: 948 TKHTKTP--MTITETPEHAFDRVVVDTIGPLPKSENGNEYAVTLICDLTKYLVAIPI-AN 1004
Query: 229 YPVAQLAEIYIQNIVKLHGVPSSIVSD 255
+A+ ++ + +G + ++D
Sbjct: 1005 KSAKTVAKAIFESFILKYGPMKTFITD 1031
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 55.8 bits (133), Expect = 2e-07
Identities = 44/176 (25%), Positives = 74/176 (42%), Gaps = 13/176 (7%)
Query: 100 VLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVY 159
V R G+ IP + ++IL+ + + H G + + +WW L+KDV + +
Sbjct: 803 VTRPNGKRIIPPKSDRPQIILQAHNIA----HTGRDSTFLKVSSKYWWPNLRKDVVKVIR 858
Query: 160 ACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKS 219
C C + +L P + P +D +D++ LP S G+ + VVVD +T
Sbjct: 859 QCKQCLVTNAATLAAPPILRP-ERPVKPFDKFFIDYIGPLP-PSNGYLHVLVVVDSMTGF 916
Query: 220 AHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQMRWVRN 275
P + A + N++ VP I SD+ FTS + W +N
Sbjct: 917 VWLYPTKAP---STSATVKALNMLTSIAVPKVIHSDQGAAFTSATFAD----WAKN 965
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 50.4 bits (119), Expect = 9e-06
Identities = 33/132 (25%), Positives = 57/132 (43%), Gaps = 5/132 (3%)
Query: 131 HLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDS 190
H G + + +WW L+KDV + + C C + + +L P+ P +D
Sbjct: 828 HTGRDATFLKVSSKYWWPNLRKDVVKSIRQCKQCLVTNATNLTSPPILRPVK-PLKPFDK 886
Query: 191 ISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPS 250
+D++ LP S G+ + VVVD +T P + A + N++ +P
Sbjct: 887 FYIDYIGPLP-PSNGYLHVLVVVDSMTGFVWLYPTKAP---STSATVKALNMLTSIAIPK 942
Query: 251 SIVSDRDPRFTS 262
+ SD+ FTS
Sbjct: 943 VLHSDQGAAFTS 954
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 41.2 bits (95), Expect = 0.005
Identities = 27/113 (23%), Positives = 57/113 (49%), Gaps = 3/113 (2%)
Query: 113 EELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQ 172
E+++ +L+E H+ L+ H G KM++ + + F+W ++ V V C C + +H
Sbjct: 1462 EKIRTPLLKELHEGMLAGHFGIKKMWRMVHRKFYWPQMRVCVENCVRTCAKCLCAN-DHS 1520
Query: 173 RPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPI 225
+ LTP + + + ++ D + + + +G+ I ++D TK +PI
Sbjct: 1521 KLTSSLTPYRM-TFPLEIVACDLM-DVGLSVQGNRYILTIIDLFTKYGTAVPI 1571
>ZB11_HUMAN (O95625) Zinc finger and BTB domain containing protein
11
Length = 1053
Score = 37.4 bits (85), Expect = 0.078
Identities = 17/61 (27%), Positives = 30/61 (48%), Gaps = 1/61 (1%)
Query: 114 ELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQR 173
E ++ ++E H H + + L K +WW G+ K V ++ C CQ+ K++ R
Sbjct: 69 ERRRDLIEAAHLGPGGTHHTRHQTWHYLSKTYWWRGILKQVKDYIKQCSKCQE-KLDRSR 127
Query: 174 P 174
P
Sbjct: 128 P 128
>VP17_YEAST (P32913) Vacuolar protein sorting-associated protein
VPS17
Length = 551
Score = 33.9 bits (76), Expect = 0.86
Identities = 23/73 (31%), Positives = 37/73 (50%)
Query: 9 DKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKINSDFLGSI 68
+K+K +A L RKTL A E L +FR L + SQS+Q +L+++ + +
Sbjct: 231 NKSKSLASGLKRKTLKQLAPPYDEITELAEFRPLVKSIYVVSQSLQEKLLRVSRNRKMMV 290
Query: 69 REAQQVDVKFVDL 81
+E FV+L
Sbjct: 291 QEENAFGQDFVNL 303
>SP10_YEAST (P35208) SPT10 protein
Length = 640
Score = 33.9 bits (76), Expect = 0.86
Identities = 21/80 (26%), Positives = 40/80 (49%), Gaps = 7/80 (8%)
Query: 99 GVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFV 158
G L +G+ I D ++ ++ LE +L HLG K+ + + + W G+K V+ +
Sbjct: 327 GKLMTKGKEIIYDTDQQIQIALE----IHLMEHLGINKVTSKIGEKYHWRGIKSTVSEVI 382
Query: 159 YACLTCQKSKVEHQRPAGLL 178
CQK K+ ++ G++
Sbjct: 383 ---SRCQKCKMRYKDGTGVI 399
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 33.9 bits (76), Expect = 0.86
Identities = 41/166 (24%), Positives = 67/166 (39%), Gaps = 12/166 (7%)
Query: 105 GRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKL-FWWSGLKKDVARFVYACLT 163
G+I +P E L + + + HLG K+ ++K F + + AC
Sbjct: 828 GKIVLPQKEALAMI-----QQMHAWTHLGNRKLKLLIEKTDFLIPRASTLIEQVTSACKV 882
Query: 164 CQKSKVEHQR-PAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHF 222
CQ+ R PAG T + P W+ +DF P+ + G+ + V VD +
Sbjct: 883 CQQVNAGATRVPAGKRTRGNRPGVYWE---IDFTEVKPHYA-GYKYLLVFVDTFSGWVEA 938
Query: 223 IPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
P +A+ ++ I G+P I SD P F S+ + L
Sbjct: 939 FPTR-QETAHIVAKKILEEIFPRFGLPKVIGSDNGPAFVSQVSQGL 983
>THIC_CLOAB (Q97EU2) Thiamine biosynthesis protein thiC
Length = 436
Score = 32.7 bits (73), Expect = 1.9
Identities = 18/57 (31%), Positives = 33/57 (57%), Gaps = 3/57 (5%)
Query: 88 AEESDFKVDEQGVLRFRGRICIPDNEELKKLILE---EGHKSNLSIHLGATKMYQDL 141
AE+ ++ E +L G+I IP N+ K L E +G K+ ++++LG +K Q++
Sbjct: 23 AEKEQMEISELKMLMAEGKIVIPANKNHKSLSAEGVGQGLKTKINVNLGISKDCQNV 79
>POL_AVIRE (P03360) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 473
Score = 32.3 bits (72), Expect = 2.5
Identities = 36/170 (21%), Positives = 72/170 (42%), Gaps = 19/170 (11%)
Query: 105 GRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTC 164
GR+ +P + + +LE+ H++ HLG +K+ + ++K + G+ + C+ C
Sbjct: 111 GRLLLP--RAVGRKVLEQTHRAT---HLGESKLTELVRKHYPICGIYRAARDITTRCVAC 165
Query: 165 QKSKVEHQRPAGLLTPLD------VPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTK 218
+ + R A + L+ P W+ +DF + G+ + V+VD +
Sbjct: 166 AQV---NPRAAPVEKGLNSRIRGAAPGEHWE---VDFTEMI-TAKGGYKYLLVLVDTFSG 218
Query: 219 SAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
P + + I +I+ G+P I SD P F ++ + L
Sbjct: 219 WVEAYPAKRETSQVVIKHL-ILDIIPRFGLPVQIGSDNGPAFVAKVTQQL 267
>PKDR_HUMAN (Q9NTG1) Polycystic kidney disease and receptor for egg
jelly related protein precursor (PKD and REJ homolog)
Length = 2253
Score = 32.3 bits (72), Expect = 2.5
Identities = 20/58 (34%), Positives = 26/58 (44%), Gaps = 2/58 (3%)
Query: 214 DRLTKSAHFIPINISYPVAQLAEI--YIQNIVKLHGVPSSIVSDRDPRFTSRFWRSLQ 269
DR+ H I + PV+ L EI + I KL PS D R T R W++ Q
Sbjct: 729 DRVNLRKHLIDQSFLLPVSTLVEIGQVVMTITKLTQKPSEFTWDAQKRATMRVWQANQ 786
>DPOL_METJA (Q58295) DNA polymerase (EC 2.7.7.7) [Contains: Mja pol-1
intein; Mja pol-2 intein]
Length = 1634
Score = 30.8 bits (68), Expect = 7.3
Identities = 21/60 (35%), Positives = 33/60 (55%), Gaps = 5/60 (8%)
Query: 66 GSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHK 125
G I E + DV+F DL+VV + VD++ V+ R+ D EE+K L++ + HK
Sbjct: 986 GKIVEVKGDDVRFGDLIVVPKKLT----CVDKEVVINIPKRLINADEEEIKDLVITK-HK 1040
>YI61_XANCV (P25438) Insertion element IS476 hypothetical 39.2 kDa
protein
Length = 346
Score = 30.4 bits (67), Expect = 9.5
Identities = 27/93 (29%), Positives = 40/93 (42%), Gaps = 4/93 (4%)
Query: 172 QRPAGLLTPLDVPEWKWDSISMDFV-SSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYP 230
+RP G L D+ SMDFV +L N R VVD T+ + I ++
Sbjct: 158 KRPVGERQKLLASSMPNDTWSMDFVFDALANARR--IKCLTVVDDFTRESVDIAVDHGIS 215
Query: 231 VAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSR 263
A + + + G P ++ +D P FTSR
Sbjct: 216 GAYVVRL-LDQAACFRGYPRAVRTDNGPEFTSR 247
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.341 0.148 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,408,752
Number of Sequences: 164201
Number of extensions: 1630988
Number of successful extensions: 5900
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 5876
Number of HSP's gapped (non-prelim): 21
length of query: 443
length of database: 59,974,054
effective HSP length: 113
effective length of query: 330
effective length of database: 41,419,341
effective search space: 13668382530
effective search space used: 13668382530
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144765.5


