

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.14 - phase: 0
(113 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
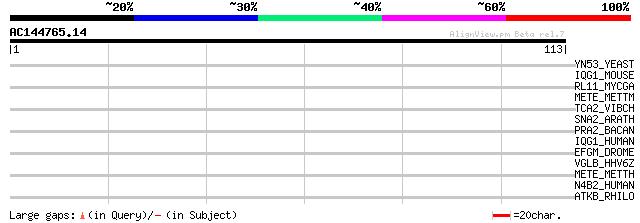
Score E
Sequences producing significant alignments: (bits) Value
YN53_YEAST (P42842) Hypothetical 102.3 kDa protein in DAL82-RFA2... 32 0.26
IQG1_MOUSE (Q9JKF1) Ras GTPase-activating-like protein IQGAP1 30 0.76
RL11_MYCGA (Q8RLD9) 50S ribosomal protein L11 28 2.9
METE_METTM (P55299) Probable methylcobalamin:homocysteine methyl... 28 2.9
TCA2_VIBCH (P23024) Toxin coregulated pilin precursor (Pilus col... 28 3.8
SNA2_ARATH (Q9SPE6) Alpha-soluble NSF attachment protein 2 (Alph... 28 4.9
PRA2_BACAN (Q81TU1) Foldase protein prsA 2 precursor (EC 5.2.1.8) 28 4.9
IQG1_HUMAN (P46940) Ras GTPase-activating-like protein IQGAP1 (p... 28 4.9
EFGM_DROME (Q9VM33) Probable elongation factor G, mitochondrial ... 28 4.9
VGLB_HHV6Z (P36320) Glycoprotein B precursor 27 6.4
METE_METTH (O26869) Probable methylcobalamin:homocysteine methyl... 27 6.4
N4B2_HUMAN (Q86UW6) Nedd4-binding protein 2 (EC 3.-.-.-) (N4BP2)... 27 8.4
ATKB_RHILO (Q98GX6) Potassium-transporting ATPase B chain (EC 3.... 27 8.4
>YN53_YEAST (P42842) Hypothetical 102.3 kDa protein in DAL82-RFA2
intergenic region
Length = 904
Score = 32.0 bits (71), Expect = 0.26
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 41 ILREAAEIFEGLGMVDSAAQCFTDLGDYERA 71
IL+ A EI+E LGM A C+ +GD ++A
Sbjct: 525 ILKSAVEIYERLGMACETALCYAAVGDEKKA 555
>IQG1_MOUSE (Q9JKF1) Ras GTPase-activating-like protein IQGAP1
Length = 1657
Score = 30.4 bits (67), Expect = 0.76
Identities = 29/93 (31%), Positives = 47/93 (50%), Gaps = 23/93 (24%)
Query: 15 GKKSKAASLRATAIRLHD----LNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYER 70
GKKSK SL+ TA RLH+ L +ED A + + +G + ++GD+E
Sbjct: 1553 GKKSKKISLKYTAARLHEKGVLLEIEDLQA--NQFKNVIFEIGPTE-------EVGDFE- 1602
Query: 71 AGINFNF-GIHVYSFMSFSIKKHY-DILEHKYQ 101
+ F G+ + +FM HY D+L+ +Y+
Sbjct: 1603 --VKAKFMGVQMETFM-----LHYQDLLQLQYE 1628
>RL11_MYCGA (Q8RLD9) 50S ribosomal protein L11
Length = 152
Score = 28.5 bits (62), Expect = 2.9
Identities = 15/40 (37%), Positives = 22/40 (54%)
Query: 15 GKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGM 54
GK SKA +L+ ++ DLN D A LR A + +G+
Sbjct: 98 GKISKAEALKIAEYKMADLNAYDTEAALRMIAGTAKQMGL 137
>METE_METTM (P55299) Probable methylcobalamin:homocysteine
methyltransferase (EC 2.1.1.-) (Methionine synthase)
Length = 309
Score = 28.5 bits (62), Expect = 2.9
Identities = 15/41 (36%), Positives = 24/41 (57%)
Query: 16 KKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVD 56
K+ +A A A+R+ +L+DA A++ + E F GMVD
Sbjct: 139 KRERAIMDMADALRVEAEHLQDAGALMIQIDEPFLSTGMVD 179
>TCA2_VIBCH (P23024) Toxin coregulated pilin precursor (Pilus
colonization factor)
Length = 224
Score = 28.1 bits (61), Expect = 3.8
Identities = 15/62 (24%), Positives = 27/62 (43%)
Query: 20 AASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFTDLGDYERAGINFNFGI 79
AA+ +A AI + L ++ ++F + + A DLGD+E + G+
Sbjct: 127 AAANKAFAISVDGLTQAQCKTLITSVGDMFPYIAIKAGGAVALADLGDFENSAAAAETGV 186
Query: 80 HV 81
V
Sbjct: 187 GV 188
>SNA2_ARATH (Q9SPE6) Alpha-soluble NSF attachment protein 2
(Alpha-SNAP2) (N-ethylmaleimide-sensitive factor
attachment protein, alpha 2)
Length = 289
Score = 27.7 bits (60), Expect = 4.9
Identities = 23/85 (27%), Positives = 34/85 (39%), Gaps = 4/85 (4%)
Query: 2 ATMCFERAGDSYW--GKKSKAASLRATAIRLH--DLNLEDANAILREAAEIFEGLGMVDS 57
A C ERA + + G+ + AA + D E A A +AAE F+ + S
Sbjct: 91 AASCLERAVNIFCEIGRLNMAARYYKEIAEYYESDQKFEQAIAYFEKAAEFFQNEEVTTS 150
Query: 58 AAQCFTDLGDYERAGINFNFGIHVY 82
A QC + Y + I +Y
Sbjct: 151 ANQCNLKVAQYAAQLEQYEKAIKIY 175
>PRA2_BACAN (Q81TU1) Foldase protein prsA 2 precursor (EC 5.2.1.8)
Length = 285
Score = 27.7 bits (60), Expect = 4.9
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 7 ERAGDSYWGKKSKAASLRATAIRLHDLNLEDANAIL 42
ER D +GKK + L+ I+++D LED I+
Sbjct: 245 ERTADPIFGKKLLQSELKKANIKINDSELEDTFTIV 280
>IQG1_HUMAN (P46940) Ras GTPase-activating-like protein IQGAP1 (p195)
Length = 1657
Score = 27.7 bits (60), Expect = 4.9
Identities = 15/27 (55%), Positives = 18/27 (66%), Gaps = 4/27 (14%)
Query: 15 GKKSKAASLRATAIRLHD----LNLED 37
GKKSK SL+ TA RLH+ L +ED
Sbjct: 1553 GKKSKKISLKYTAARLHEKGVLLEIED 1579
>EFGM_DROME (Q9VM33) Probable elongation factor G, mitochondrial
precursor (mEF-G)
Length = 745
Score = 27.7 bits (60), Expect = 4.9
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Query: 8 RAGDSYWGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAAQCFT 63
R GD+ + ++ A +RLH +ED N + A +IF G+ ++ FT
Sbjct: 373 RKGDNIFNARTNKKVRIARLVRLHSNQMEDVNEVY--AGDIFALFGVDCASGDTFT 426
>VGLB_HHV6Z (P36320) Glycoprotein B precursor
Length = 830
Score = 27.3 bits (59), Expect = 6.4
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 5/49 (10%)
Query: 48 IFEGLGMVDSAAQCFTDLGDYERAGINFNFGIHVYSFMSFSIKKHYDIL 96
+F + +++S + D DY RAG N H Y F SI K D++
Sbjct: 7 LFLAVFLMNSVLMIYCDSDDYIRAGYN-----HKYPFRICSIAKGTDLM 50
>METE_METTH (O26869) Probable methylcobalamin:homocysteine
methyltransferase (EC 2.1.1.-) (Methionine synthase)
Length = 308
Score = 27.3 bits (59), Expect = 6.4
Identities = 22/96 (22%), Positives = 47/96 (48%), Gaps = 10/96 (10%)
Query: 16 KKSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVD--SAAQCFTDLGDYERAGI 73
K+ +A A +++ +L+DA A + + E F G+VD +A + D+ AG+
Sbjct: 139 KRDRAVMDMAGVLKIEAKHLQDAGAAMIQIDEPFLSTGIVDMKTAGRAI----DHIAAGL 194
Query: 74 NFNFGIHV----YSFMSFSIKKHYDILEHKYQVYPT 105
+ +HV + +S ++ D+L+ ++ P+
Sbjct: 195 DIEVSLHVCGDIRNVLSDLLRFKVDVLDLEFAGRPS 230
>N4B2_HUMAN (Q86UW6) Nedd4-binding protein 2 (EC 3.-.-.-) (N4BP2)
(BCL-3 binding protein)
Length = 1770
Score = 26.9 bits (58), Expect = 8.4
Identities = 15/46 (32%), Positives = 25/46 (53%), Gaps = 2/46 (4%)
Query: 5 CFERAGDSYWGKKSKAASLRATAIRLHDLNLEDANAILREAAEIFE 50
C+ +A ++Y K A+ A LH+ +++AN + A EIFE
Sbjct: 1636 CYSKAKEAYRIGKKNVATFYAQQGTLHEQKMKEANHL--AAIEIFE 1679
>ATKB_RHILO (Q98GX6) Potassium-transporting ATPase B chain (EC
3.6.3.12) (Potassium-translocating ATPase B chain) (ATP
phosphohydrolase [potassium-transporting] B chain)
(Potassium binding and translocating subunit B)
Length = 697
Score = 26.9 bits (58), Expect = 8.4
Identities = 13/43 (30%), Positives = 25/43 (57%)
Query: 17 KSKAASLRATAIRLHDLNLEDANAILREAAEIFEGLGMVDSAA 59
+SK + T++++ D+ L +A I+ E+ EG+ V+ AA
Sbjct: 114 RSKFKLVPGTSLKVGDIVLVEAGDIIPSDGEVVEGVASVNEAA 156
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,261,867
Number of Sequences: 164201
Number of extensions: 436044
Number of successful extensions: 1371
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 1363
Number of HSP's gapped (non-prelim): 13
length of query: 113
length of database: 59,974,054
effective HSP length: 89
effective length of query: 24
effective length of database: 45,360,165
effective search space: 1088643960
effective search space used: 1088643960
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144765.14


