

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144759.9 - phase: 0
(749 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
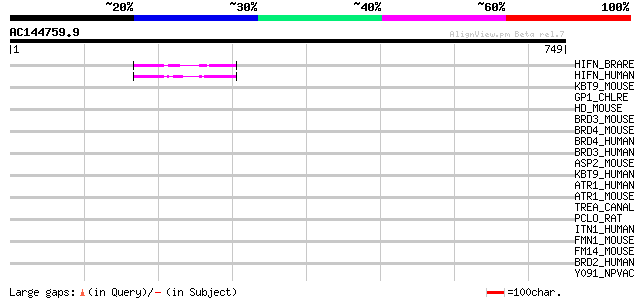
Score E
Sequences producing significant alignments: (bits) Value
HIFN_BRARE (P59723) Hypoxia-inducible factor 1 alpha inhibitor (... 58 8e-08
HIFN_HUMAN (Q9NWT6) Hypoxia-inducible factor 1 alpha inhibitor (... 57 2e-07
KBT9_MOUSE (Q80T74) Kelch repeat and BTB domain containing prote... 41 0.013
GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor (H... 40 0.029
HD_MOUSE (P42859) Huntingtin (Huntington's disease protein homol... 39 0.038
BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3 (Bromodomai... 39 0.050
BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic ch... 38 0.086
BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1 prot... 38 0.086
BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like... 38 0.086
ASP2_MOUSE (Q8CG79) Apoptosis stimulating of p53 protein 2 (Tumo... 38 0.086
KBT9_HUMAN (Q96CT2) Kelch repeat and BTB domain containing prote... 38 0.11
ATR1_HUMAN (Q9H6X2) Anthrax toxin receptor 1 precursor (Tumor en... 38 0.11
ATR1_MOUSE (Q9CZ52) Anthrax toxin receptor 1 precursor (Tumor en... 37 0.15
TREA_CANAL (P52494) Neutral trehalase (EC 3.2.1.28) (Alpha,alpha... 37 0.19
PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic cytom... 37 0.25
ITN1_HUMAN (Q15811) Intersectin 1 (SH3 domain-containing protein... 37 0.25
FMN1_MOUSE (Q05860) Formin 1 isoforms I/II/III (Limb deformity p... 37 0.25
FM14_MOUSE (Q05859) Formin 1 isoform IV (Limb deformity protein) 37 0.25
BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3 prot... 37 0.25
Y091_NPVAC (P41479) Hypothetical 24.1 kDa protein in LEF4-P33 in... 36 0.43
>HIFN_BRARE (P59723) Hypoxia-inducible factor 1 alpha inhibitor (EC
1.14.11.16) (Hypoxia-inducible factor asparagine
hydroxylase)
Length = 344
Score = 58.2 bits (139), Expect = 8e-08
Identities = 41/139 (29%), Positives = 59/139 (41%), Gaps = 28/139 (20%)
Query: 168 LWIGNKHSSTWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEF 227
L IG + + T H D +N +A + G K +LFPP Y P + +
Sbjct: 178 LLIGMEGNVTPAHYDEQQNFFAQIKGHKRCILFPPEQFDCLY----PYPVHHPCDRQSQV 233
Query: 228 DLELDKPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFH 287
D E NP + KFP + N +E V G++LY+P W+H
Sbjct: 234 DFE-------------NP---------DYDKFPNFKNAVG-YEAVVGPGDVLYIPMYWWH 270
Query: 288 HVRQSGDDGELTIAVNYWY 306
H+ + GE TI VN+WY
Sbjct: 271 HIESLLNGGE-TITVNFWY 288
>HIFN_HUMAN (Q9NWT6) Hypoxia-inducible factor 1 alpha inhibitor (EC
1.14.11.16) (Hypoxia-inducible factor asparagine
hydroxylase) (Factor inhibiting HIF-1) (FIH-1)
Length = 349
Score = 57.0 bits (136), Expect = 2e-07
Identities = 39/139 (28%), Positives = 62/139 (44%), Gaps = 28/139 (20%)
Query: 168 LWIGNKHSSTWFHKDHYENLYAVVTGQKHFLLFPPTDVHRFYIRNYPAATYKYYMETGEF 227
L IG + + T H D +N +A + G K +LFPP Y YP ++ +
Sbjct: 187 LLIGMEGNVTPAHYDEQQNFFAQIKGYKRCILFPPDQFECLY--PYPV----HHPCDRQS 240
Query: 228 DLELDKPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFECTVKAGEILYLPSMWFH 287
++ D P + +FP F +E V G++LY+P W+H
Sbjct: 241 QVDFDNP--------------------DYERFP-NFQNVVGYETVVGPGDVLYIPMYWWH 279
Query: 288 HVRQSGDDGELTIAVNYWY 306
H+ +S +G +TI VN+WY
Sbjct: 280 HI-ESLLNGGITITVNFWY 297
>KBT9_MOUSE (Q80T74) Kelch repeat and BTB domain containing protein
9
Length = 655
Score = 40.8 bits (94), Expect = 0.013
Identities = 39/183 (21%), Positives = 79/183 (42%), Gaps = 31/183 (16%)
Query: 334 PIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPP 393
P++ PPPPA P+ + +T PP+ H L++ L I T P
Sbjct: 40 PLLPPPPPAQPSAALPSSVPATNGPPTTDSAHGLQM----------LRTIGVGKYEFTDP 89
Query: 394 -------KKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDP--ETAIELH 444
K+ ++R+ + + +++ F +++ +++ C+L + +++
Sbjct: 90 GHPKEMLKELNQQRRAKAFTDLKIVVEGREF----EVHQNVLASCSLYFKDLIQRSVQDS 145
Query: 445 TQIITRGIELPLTLLN----RILIMFVSCGLL----ENARRVFDVMSVRDFHSWATLFVS 496
+Q +EL L+ L +L+ FV G L NA+ + + S FH++ + VS
Sbjct: 146 SQSSREKLELVLSNLQADVLELLLEFVYTGSLVIDSANAKTLLEAASKFQFHTFCKVCVS 205
Query: 497 YYE 499
+ E
Sbjct: 206 FLE 208
>GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor
(Hydroxyproline-rich glycoprotein 1)
Length = 555
Score = 39.7 bits (91), Expect = 0.029
Identities = 27/68 (39%), Positives = 32/68 (46%), Gaps = 5/68 (7%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRN-PKPKNLSLIHP 385
+P SP+ P APP PAPP+ + A PPS P R P N P P + P
Sbjct: 240 APPSPVPPSPAPPSPAPPSPKPPA----PPPPPSPPPPPPPRPPFPANTPMPPSPPSPPP 295
Query: 386 SSQPITPP 393
S P TPP
Sbjct: 296 SPAPPTPP 303
Score = 39.7 bits (91), Expect = 0.029
Identities = 24/68 (35%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLS-LIHP 385
+P SP P APP PAPP+ A + + P + P PL +P P + S + P
Sbjct: 102 APPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPP 161
Query: 386 SSQPITPP 393
S P PP
Sbjct: 162 SPSPPVPP 169
Score = 37.7 bits (86), Expect = 0.11
Identities = 25/71 (35%), Positives = 31/71 (43%), Gaps = 5/71 (7%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKN-LSLIHP 385
SP P P APP PAPP+ + + +PPS P P +P P + S P
Sbjct: 72 SPAPPSPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPS----PAPPSPAPPSPPSPAPP 127
Query: 386 SSQPITPPKKS 396
S P PP S
Sbjct: 128 SPSPPAPPSPS 138
Score = 36.6 bits (83), Expect = 0.25
Identities = 24/72 (33%), Positives = 29/72 (39%), Gaps = 3/72 (4%)
Query: 325 YRSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIH 384
+ P SP P APP PAPP+ + + PPS P P P P S
Sbjct: 37 FNCPPSPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPP---SPPSPAPPSPAPPSPAP 93
Query: 385 PSSQPITPPKKS 396
PS P +P S
Sbjct: 94 PSPAPPSPAPPS 105
Score = 35.8 bits (81), Expect = 0.43
Identities = 23/67 (34%), Positives = 28/67 (41%), Gaps = 6/67 (8%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
SP+ P P PP PAPP+ + PP + P P R P P N + P
Sbjct: 235 SPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPP----PPPRPPFPANTPM--PP 288
Query: 387 SQPITPP 393
S P PP
Sbjct: 289 SPPSPPP 295
Score = 35.0 bits (79), Expect = 0.73
Identities = 21/66 (31%), Positives = 28/66 (41%), Gaps = 4/66 (6%)
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSS 387
P P P P PP+PP+ + + +PPS P P+ +P P S PS
Sbjct: 275 PPRPPFPANTPMPPSPPSPPPSPAPPTPPTPPSPSPPS----PVPPSPAPVPPSPAPPSP 330
Query: 388 QPITPP 393
P PP
Sbjct: 331 APSPPP 336
Score = 34.7 bits (78), Expect = 0.95
Identities = 21/67 (31%), Positives = 28/67 (41%), Gaps = 7/67 (10%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
SP+ P P +PP PAPP A + + P + P + P +P P PS
Sbjct: 128 SPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTP-------PS 180
Query: 387 SQPITPP 393
P PP
Sbjct: 181 PSPPVPP 187
Score = 34.3 bits (77), Expect = 1.2
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 4/67 (5%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
+P SP P+ PP PAPP+ A + + P + P P+ +P P S PS
Sbjct: 203 APPSPAPPV--PPSPAPPSPPSPAPPSPPSPAPPSPSPPAPPSPVPPSPAPP--SPAPPS 258
Query: 387 SQPITPP 393
+P PP
Sbjct: 259 PKPPAPP 265
Score = 33.5 bits (75), Expect = 2.1
Identities = 20/67 (29%), Positives = 27/67 (39%), Gaps = 7/67 (10%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPS 386
SP P+ P APP P+PP + + P + P P+ +P P PS
Sbjct: 141 SPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSPPVPPSPAP-------PS 193
Query: 387 SQPITPP 393
P PP
Sbjct: 194 PAPPVPP 200
Score = 33.1 bits (74), Expect = 2.8
Identities = 22/68 (32%), Positives = 29/68 (42%), Gaps = 2/68 (2%)
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSS 387
P SP P +PP PAPP+ A + + P + P P P PK + P S
Sbjct: 212 PPSPAPP--SPPSPAPPSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPS 269
Query: 388 QPITPPKK 395
P PP +
Sbjct: 270 PPPPPPPR 277
Score = 33.1 bits (74), Expect = 2.8
Identities = 23/68 (33%), Positives = 29/68 (41%), Gaps = 6/68 (8%)
Query: 327 SPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKN-LSLIHP 385
SP+ P+ P APP PAPP A + P + P P +P P + S P
Sbjct: 180 SPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAP-----PSPPSPAPPSPPSPAPP 234
Query: 386 SSQPITPP 393
S P PP
Sbjct: 235 SPSPPAPP 242
Score = 32.7 bits (73), Expect = 3.6
Identities = 22/68 (32%), Positives = 26/68 (37%), Gaps = 5/68 (7%)
Query: 326 RSPTSPMMPIVAPP-PPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIH 384
+ P P P PP PP PP S SPP + P P P P S +
Sbjct: 260 KPPAPPPPPSPPPPPPPRPPFPANTPMPPSPPSPPPSPAPPTPPTP----PSPSPPSPVP 315
Query: 385 PSSQPITP 392
PS P+ P
Sbjct: 316 PSPAPVPP 323
Score = 32.3 bits (72), Expect = 4.7
Identities = 20/66 (30%), Positives = 27/66 (40%), Gaps = 4/66 (6%)
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSS 387
P SP P PP P+PP A + P + P P+ +P P + PS
Sbjct: 168 PPSPAPPSPTPPSPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPS----PPSP 223
Query: 388 QPITPP 393
P +PP
Sbjct: 224 APPSPP 229
Score = 32.0 bits (71), Expect = 6.1
Identities = 22/70 (31%), Positives = 28/70 (39%), Gaps = 3/70 (4%)
Query: 327 SPTSPMMPIVAPP---PPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLI 383
SP P P APP PPAPP+ + + +PPS P P P P
Sbjct: 222 SPAPPSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFP 281
Query: 384 HPSSQPITPP 393
+ P +PP
Sbjct: 282 ANTPMPPSPP 291
>HD_MOUSE (P42859) Huntingtin (Huntington's disease protein homolog)
(HD protein)
Length = 3119
Score = 39.3 bits (90), Expect = 0.038
Identities = 29/97 (29%), Positives = 38/97 (38%), Gaps = 20/97 (20%)
Query: 317 FNFLQSIDYRSPTSPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPK 376
F L+S + P P APPPP PP PP QP P + P
Sbjct: 11 FESLKSFQQQQQQQP--PPQAPPPPPPP-------------PPQPPQP-----PPQGQPP 50
Query: 377 PKNLSLIHPSSQPITPPKKSKRRRKCDTTSHILPLMD 413
P L P+ +P+ PKK K D +H L + +
Sbjct: 51 PPPPPLPGPAEEPLHRPKKELSATKKDRVNHCLTICE 87
>BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3
(Bromodomain-containing FSH-like protein FSRG2)
Length = 726
Score = 38.9 bits (89), Expect = 0.050
Identities = 22/70 (31%), Positives = 34/70 (48%), Gaps = 2/70 (2%)
Query: 328 PTSPMMPIVAPPPPAPPTR--RRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHP 385
P +P++P+V P PP + +R ADTT+ T+ PL + K ++
Sbjct: 221 PATPIVPVVPPTPPVVKKKGVKRKADTTTPTTSAITASRSESPPPLSEPKQAKVVARRES 280
Query: 386 SSQPITPPKK 395
+PI PPKK
Sbjct: 281 GGRPIKPPKK 290
>BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic
chromosome-associated protein) (MCAP)
Length = 1400
Score = 38.1 bits (87), Expect = 0.086
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 7/71 (9%)
Query: 328 PTSPMMPIVAPPPPAPPTRR---RNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIH 384
P PI+A P T++ R ADTT+ T+ H+P L PK L
Sbjct: 268 PVQSHPPIIATTPQPVKTKKGVKRKADTTTPTTIDPIHEPP----SLAPEPKTAKLGPRR 323
Query: 385 PSSQPITPPKK 395
SS+P+ PPKK
Sbjct: 324 ESSRPVKPPKK 334
Score = 33.1 bits (74), Expect = 2.8
Identities = 26/106 (24%), Positives = 40/106 (37%), Gaps = 27/106 (25%)
Query: 319 FLQSIDYRSPTSPMMPIVA---------PPPPAPPTRRRNADTTSTTSPPSNHQP----- 364
+LQ + P +P++P V PPPP P +++ PP QP
Sbjct: 933 YLQQLQKVQPPTPLLPSVKVQSQPPPPLPPPPHPSVQQQQLQPQPPPPPPPQPQPPPQQQ 992
Query: 365 --------HLLRLP----LRRNPKPKNLSLIH-PSSQPITPPKKSK 397
HL +P +++ P P H P Q PP+ +K
Sbjct: 993 HQPPPRPVHLPSMPFSAHIQQPPPPPGQQPTHPPPGQQPPPPQPAK 1038
>BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1
protein)
Length = 1362
Score = 38.1 bits (87), Expect = 0.086
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 7/71 (9%)
Query: 328 PTSPMMPIVAPPPPAPPTRR---RNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIH 384
P PI+A P T++ R ADTT+ T+ H+P L PK L
Sbjct: 267 PVQSHPPIIAATPQPVKTKKGVKRKADTTTPTTIDPIHEPP----SLPPEPKTTKLGQRR 322
Query: 385 PSSQPITPPKK 395
SS+P+ PPKK
Sbjct: 323 ESSRPVKPPKK 333
Score = 33.5 bits (75), Expect = 2.1
Identities = 22/82 (26%), Positives = 35/82 (41%), Gaps = 7/82 (8%)
Query: 319 FLQSIDYRSPTSPMMPIVA----PPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRN 374
+LQ + P +P++P V PPPP PP + PP P P +++
Sbjct: 931 YLQQLQKVQPPTPLLPSVKVQSQPPPPLPPPPHPSVQQQLQQQPPPPPPPQPQPPPQQQH 990
Query: 375 ---PKPKNLSLIHPSSQPITPP 393
P+P +L + S+ PP
Sbjct: 991 QPPPRPVHLQPMQFSTHIQQPP 1012
>BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like
protein)
Length = 726
Score = 38.1 bits (87), Expect = 0.086
Identities = 22/70 (31%), Positives = 34/70 (48%), Gaps = 2/70 (2%)
Query: 328 PTSPMMPIVAPPPPAPPTR--RRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHP 385
P +P++P+V P PP + +R ADTT+ T+ PL + K ++
Sbjct: 222 PATPIVPVVPPTPPVVKKKGVKRKADTTTPTTSAITASRSESPPPLSDPKQAKVVARRES 281
Query: 386 SSQPITPPKK 395
+PI PPKK
Sbjct: 282 GGRPIKPPKK 291
>ASP2_MOUSE (Q8CG79) Apoptosis stimulating of p53 protein 2 (Tumor
suppressor p53-binding protein 2) (Fragment)
Length = 1088
Score = 38.1 bits (87), Expect = 0.086
Identities = 36/111 (32%), Positives = 52/111 (46%), Gaps = 24/111 (21%)
Query: 24 ERLESAPTPLQFHRNFIT----PNKPCIIS-----NSISHWPSLSLWSHPSYLTQSLSST 74
++L +AP PL+ R+ IT PN P I +I+ ++S+ SHPS +S S
Sbjct: 682 KKLSNAPRPLK-KRSSITEPEGPNGPNIQKLLYQRTTIAAMETISVPSHPS---KSPGSV 737
Query: 75 TVS-----------LHLTPTGSADSLTPLPSSPSSLCFASAHVQNLPFPEA 114
TV+ LH+ P SL P P SP + AS ++P P A
Sbjct: 738 TVNPESSVEIPNPYLHVEPEKEVGSLVPEPLSPEDMGSASTENSDVPAPSA 788
>KBT9_HUMAN (Q96CT2) Kelch repeat and BTB domain containing protein
9
Length = 655
Score = 37.7 bits (86), Expect = 0.11
Identities = 39/183 (21%), Positives = 78/183 (42%), Gaps = 31/183 (16%)
Query: 334 PIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPITPP 393
P++ PPPPA P+ + +T PP+ H L++ L I T P
Sbjct: 40 PLLPPPPPAQPSATLPSGAPATNGPPTTDSAHGLQM----------LRTIGVGKYEFTDP 89
Query: 394 -------KKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDP--ETAIELH 444
K+ ++R+ + + +++ F +++ +++ C+L + +++
Sbjct: 90 GHPREMLKELNQQRRAKAFTDLKIVVEGREF----EVHQNVLASCSLYFKDLIQRSVQDS 145
Query: 445 TQIITRGIELPLTLLN----RILIMFVSCGLL----ENARRVFDVMSVRDFHSWATLFVS 496
Q +EL L+ L +L+ FV G L NA+ + + S FH++ + VS
Sbjct: 146 GQGGREKLELVLSNLQADVLELLLEFVYTGSLVIDSANAKTLLEAASKFQFHTFCKVCVS 205
Query: 497 YYE 499
+ E
Sbjct: 206 FLE 208
>ATR1_HUMAN (Q9H6X2) Anthrax toxin receptor 1 precursor (Tumor
endothelial marker 8)
Length = 564
Score = 37.7 bits (86), Expect = 0.11
Identities = 22/55 (40%), Positives = 25/55 (45%), Gaps = 2/55 (3%)
Query: 325 YRSPTSPMMPIVAPPPPAP--PTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKP 377
Y + + P PI PPPPAP P +A T SPPS P P R P P
Sbjct: 501 YHTSSPPPAPIYTPPPPAPHCPPPPPSAPTPPIPSPPSTLPPPPQAPPPNRAPPP 555
>ATR1_MOUSE (Q9CZ52) Anthrax toxin receptor 1 precursor (Tumor
endothelial marker 8)
Length = 562
Score = 37.4 bits (85), Expect = 0.15
Identities = 22/55 (40%), Positives = 24/55 (43%), Gaps = 2/55 (3%)
Query: 325 YRSPTSPMMPIVAPPPPAP--PTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKP 377
Y + P PI PPPPAP P +A T SPPS P P R P P
Sbjct: 499 YHPSSPPPAPIYTPPPPAPHCPPPAPSAPTPPIPSPPSTLPPPPQAPPPNRAPPP 553
>TREA_CANAL (P52494) Neutral trehalase (EC 3.2.1.28)
(Alpha,alpha-trehalase) (Alpha,alpha-trehalose
glucohydrolase)
Length = 907
Score = 37.0 bits (84), Expect = 0.19
Identities = 37/127 (29%), Positives = 57/127 (44%), Gaps = 14/127 (11%)
Query: 292 SGDDGELTIAVNYWYDMQFDI--KYAYFNFLQSIDYRS-PTSPMMPIVAPPPPAPPTRRR 348
S DD +A Y+ + + + F+ +S Y + P SP+ P PPP T
Sbjct: 13 SSDDDPFDVAEKYYGEERKKKLNRVRTFSAFESTKYGAGPISPLRPTYEPPPVIKETSEP 72
Query: 349 NADTTSTTSPPS-NHQPHLLRLPLRRNPKPKNLSLIHPSSQ-------PITPPKKSKRRR 400
+ ++ST+SPP+ Q ++ L K KN S IH S+ P++ KK +
Sbjct: 73 LSSSSSTSSPPTLTPQTSQVQFNLGVG-KTKNGSAIHSDSEEEEEDEDPVSNTKKGEADE 131
Query: 401 K--CDTT 405
K DTT
Sbjct: 132 KDPFDTT 138
>PCLO_RAT (Q9JKS6) Piccolo protein (Multidomain presynaptic
cytomatrix protein)
Length = 5085
Score = 36.6 bits (83), Expect = 0.25
Identities = 28/118 (23%), Positives = 50/118 (41%), Gaps = 14/118 (11%)
Query: 331 PMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQPI 390
P P A P P PPT + T + P + QP L + P + P K++S + +P+
Sbjct: 447 PQPPTPAKPQPQPPTATKPQPQPPTATKPHHQQPGLAK-PSAQQP-TKSISQT-VTGRPL 503
Query: 391 TPPKKSKRRRK-----------CDTTSHILPLMDALHFPITIDIYTSLVKECTLSTDP 437
PP S + C+TT +L + + +F + +++ C + +P
Sbjct: 504 QPPPTSAAQTPAQGLSKTICPLCNTTELLLHIPEKANFNTCTECQSTVCSLCGFNPNP 561
Score = 34.3 bits (77), Expect = 1.2
Identities = 22/65 (33%), Positives = 28/65 (42%), Gaps = 6/65 (9%)
Query: 330 SPMMPIVAPPPPAPPTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHPSSQP 389
SP PI A P P P + T S P+ QP + P P+P+ + P QP
Sbjct: 396 SPQQPIPAKPQPQQPVATK---TQPQQSAPAKPQP---QQPAPAKPQPQQPTPAKPQPQP 449
Query: 390 ITPPK 394
TP K
Sbjct: 450 PTPAK 454
>ITN1_HUMAN (Q15811) Intersectin 1 (SH3 domain-containing protein
1A) (SH3P17)
Length = 1721
Score = 36.6 bits (83), Expect = 0.25
Identities = 41/185 (22%), Positives = 72/185 (38%), Gaps = 27/185 (14%)
Query: 233 KPTRYVPWCSVNPFPSPENLEDEISKFPLYFNGPPPFEC------TVKAGEILYLPSMWF 286
KP PW + P + ++ + +Y+ PFE T++ G+I+ + W
Sbjct: 717 KPAVQAPWSTAEKGPLTISAQENVKV--VYYRALYPFESRSHDEITIQPGDIVMVKGEWV 774
Query: 287 HHVRQSGDDGELTIAVNYWYDMQFDIKYAYF--NFLQSIDYRSPTSPMMPIV---APPPP 341
Q+G+ G W + K +F N+ + I +P+ P+ + P P
Sbjct: 775 DE-SQTGEPG--------WLGGELKGKTGWFPANYAEKIPENEVPAPVKPVTDSTSAPAP 825
Query: 342 APPTRRRNADTTSTTSPPS----NHQPHLLRLPLRRNPKPKNLSLIHPSSQP-ITPPKKS 396
R A T+S PS N P N KP+ + ++QP +T P
Sbjct: 826 KLALRETPAPLAVTSSEPSTTPNNWADFSSTWPTSTNEKPETDNWDAWAAQPSLTVPSAG 885
Query: 397 KRRRK 401
+ R++
Sbjct: 886 QLRQR 890
>FMN1_MOUSE (Q05860) Formin 1 isoforms I/II/III (Limb deformity
protein)
Length = 1468
Score = 36.6 bits (83), Expect = 0.25
Identities = 23/68 (33%), Positives = 28/68 (40%), Gaps = 9/68 (13%)
Query: 328 PTSPMMPIVAPPPPAP--PTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHP 385
P P+ P+ PPPP P PT +D PP P++L LP P P P
Sbjct: 902 PIPPVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPPPLPNVLALPNSGGPPP-------P 954
Query: 386 SSQPITPP 393
P PP
Sbjct: 955 PPPPPPPP 962
>FM14_MOUSE (Q05859) Formin 1 isoform IV (Limb deformity protein)
Length = 1206
Score = 36.6 bits (83), Expect = 0.25
Identities = 23/68 (33%), Positives = 28/68 (40%), Gaps = 9/68 (13%)
Query: 328 PTSPMMPIVAPPPPAP--PTRRRNADTTSTTSPPSNHQPHLLRLPLRRNPKPKNLSLIHP 385
P P+ P+ PPPP P PT +D PP P++L LP P P P
Sbjct: 676 PIPPVCPVSPPPPPPPPPPTPVPPSDGPPPPPPPPPPLPNVLALPNSGGPPP-------P 728
Query: 386 SSQPITPP 393
P PP
Sbjct: 729 PPPPPPPP 736
>BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3
protein) (O27.1.1)
Length = 801
Score = 36.6 bits (83), Expect = 0.25
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 27/91 (29%)
Query: 320 LQSIDYRSPTSPMMPIVAPPPPAPPTR----RRNADTTSTT-----------SPPSNHQP 364
L+S+ P P++ + A PP P + +R ADTT+ T SPP + +P
Sbjct: 251 LKSLHSAGP--PLLAVTAAPPAQPLAKKKGVKRKADTTTPTPTAILAPGSPASPPGSLEP 308
Query: 365 HLLRLPLRRNPKPKNLSLIHPSSQPITPPKK 395
RLP R S +PI PP+K
Sbjct: 309 KAARLPPMR----------RESGRPIKPPRK 329
>Y091_NPVAC (P41479) Hypothetical 24.1 kDa protein in LEF4-P33
intergenic region
Length = 224
Score = 35.8 bits (81), Expect = 0.43
Identities = 29/105 (27%), Positives = 38/105 (35%), Gaps = 4/105 (3%)
Query: 328 PTSPMMPIVAPPPPAPPTRRRNADTTSTTSP---PSNHQPHLLRLPLRRNPKPK-NLSLI 383
PT P P +PP P PP T T P P P P+ P P S I
Sbjct: 52 PTPPPTPTPSPPSPTPPPTPIPPTPTPTPPPTPIPPTPTPTPPPSPIPPTPTPSPPPSPI 111
Query: 384 HPSSQPITPPKKSKRRRKCDTTSHILPLMDALHFPITIDIYTSLV 428
P+ P PP PL + +++P ID +L+
Sbjct: 112 PPTPTPSPPPSPIPPTPTPSPPPSPSPLGEPMYYPSNIDTNEALI 156
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 93,987,728
Number of Sequences: 164201
Number of extensions: 4271957
Number of successful extensions: 16038
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 84
Number of HSP's that attempted gapping in prelim test: 15187
Number of HSP's gapped (non-prelim): 505
length of query: 749
length of database: 59,974,054
effective HSP length: 118
effective length of query: 631
effective length of database: 40,598,336
effective search space: 25617550016
effective search space used: 25617550016
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144759.9


