

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
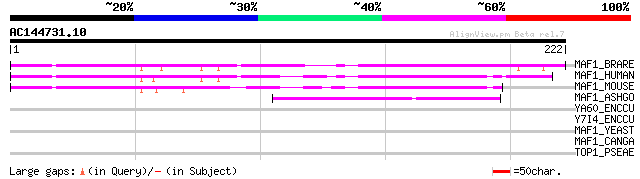
Score E
Sequences producing significant alignments: (bits) Value
MAF1_BRARE (Q6PGU2) Repressor of RNA polymerase III transcriptio... 85 1e-16
MAF1_HUMAN (Q9H063) Repressor of RNA polymerase III transcriptio... 79 6e-15
MAF1_MOUSE (Q9D0U6) Repressor of RNA polymerase III transcriptio... 79 8e-15
MAF1_ASHGO (Q757K3) Repressor of RNA polymerase III transcriptio... 43 5e-04
YA60_ENCCU (Q8STI4) Hypothetical protein ECU10_0060/ECU11_2080i 37 0.048
Y7I4_ENCCU (Q8SUU2) Hypothetical protein ECU07_1840 37 0.048
MAF1_YEAST (P41910) Repressor of RNA polymerase III transcriptio... 36 0.082
MAF1_CANGA (Q6FIZ7) Repressor of RNA polymerase III transcriptio... 34 0.24
TOP1_PSEAE (Q9HZJ5) DNA topoisomerase I (EC 5.99.1.2) (Omega-pro... 29 7.7
>MAF1_BRARE (Q6PGU2) Repressor of RNA polymerase III transcription
MAF1 homolog
Length = 247
Score = 85.1 bits (209), Expect = 1e-16
Identities = 77/257 (29%), Positives = 118/257 (44%), Gaps = 54/257 (21%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK++ E +L+ L
Sbjct: 1 MKLLENSSFEAINSLLT-IETGDCKIIGRIESYSCKMAGDDKQMFKQFCQEGQPHVLEAL 59
Query: 57 GKS-----SDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT* 92
S N +D P+ D S +T S H F+
Sbjct: 60 SPPQSSGISPNKLSHSLDDGEGPLSDKCSRKTLFYLIATLNESFRPDYDFSRTKGHDFS- 118
Query: 93 FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDD 152
PS+ + +++ L + +H L+PQ L++A+D+ +SL +
Sbjct: 119 KEPSVNWVFNAVNSSLSAAAGEAYSH------------LQPQ-----LWEALDKEISLAE 161
Query: 153 CEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIAD-----GFSLD 207
C+IY Y PD ++DP E G +WSFN+ FYN++LKRIV F S +A G LD
Sbjct: 162 CDIYSYNPDLDSDPYGEEGNLWSFNYFFYNKRLKRIVFFTCRSVSLFMAPRDSGIGTELD 221
Query: 208 EICDE--YDEEIFADMD 222
DE Y+EE +MD
Sbjct: 222 LELDEGSYEEEEEGNMD 238
>MAF1_HUMAN (Q9H063) Repressor of RNA polymerase III transcription
MAF1 homolog
Length = 256
Score = 79.3 bits (194), Expect = 6e-15
Identities = 77/246 (31%), Positives = 113/246 (45%), Gaps = 51/246 (20%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 ------GKSSDNDSPSPVDSLSSPV-DSLSART------------------SPAGRHWFT 91
G S S S P+ D S +T S A H F+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEEEGPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 92 *FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLD 151
PS++ + ++ L V F LKPQ L+ AVDE + L
Sbjct: 120 -REPSLSWVVNAVNCSL---------FSAVREDFKD---LKPQ-----LWNAVDEEICLA 161
Query: 152 DCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFSNLIADGFSLDEICD 211
+C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S + ++ E +
Sbjct: 162 ECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS-ISGSTYTPSEAGN 218
Query: 212 EYDEEI 217
E D E+
Sbjct: 219 ELDMEL 224
>MAF1_MOUSE (Q9D0U6) Repressor of RNA polymerase III transcription
MAF1 homolog
Length = 258
Score = 79.0 bits (193), Expect = 8e-15
Identities = 70/231 (30%), Positives = 102/231 (43%), Gaps = 60/231 (25%)
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE----ILDYL 56
MK LE + + ++ L+ + G+ I G IE+YSCK +G DK + E +L+ L
Sbjct: 1 MKLLENSSFEAINSQLT-VETGDAHIIGRIESYSCKMAGDDKHMFKQFCQEGQPHVLEAL 59
Query: 57 G-------------KSSDNDSPSPV-----------------DSLSSPVDSLSARTSPAG 86
KS + SP+ +S D +AR+
Sbjct: 60 SPPQTSGLSPSRLSKSQGGEDESPLSDKCSRKTLFYLIATLNESFRPDYDFSTARSHEFS 119
Query: 87 RHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDE 146
R PS+ + ++ L V F LKPQ L+ AVDE
Sbjct: 120 RE------PSLRWVVNAVNCSL---------FSAVREDF---KALKPQ-----LWNAVDE 156
Query: 147 VVSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIVTFRLSCFS 197
+ L +C+IY Y PD ++DP E G++WSFN+ FYN++LKRIV F SC S
Sbjct: 157 EICLAECDIYSYNPDLDSDPFGEDGSLWSFNYFFYNKRLKRIVFF--SCRS 205
>MAF1_ASHGO (Q757K3) Repressor of RNA polymerase III transcription
MAF1
Length = 397
Score = 43.1 bits (100), Expect = 5e-04
Identities = 26/91 (28%), Positives = 45/91 (48%), Gaps = 1/91 (1%)
Query: 106 AQLKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEAD 165
A LK T F + + ++S+F + + +++ ++ + L DC Y Y P E+
Sbjct: 239 ASLKPTDFVKSSRKQLVSKFENSIIALGKQPQEWIWETINAHMDLADCVFYQYNPQ-ESF 297
Query: 166 PLPERGAIWSFNFLFYNRKLKRIVTFRLSCF 196
E G +WS + +N+K KR+ F LS F
Sbjct: 298 LDDEPGHLWSLMWFMFNKKRKRVAYFYLSAF 328
>YA60_ENCCU (Q8STI4) Hypothetical protein ECU10_0060/ECU11_2080i
Length = 283
Score = 36.6 bits (83), Expect = 0.048
Identities = 18/42 (42%), Positives = 25/42 (58%), Gaps = 4/42 (9%)
Query: 148 VSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIV 189
VSL CEI+ Y E P +W F+FLF+N+K +R+V
Sbjct: 237 VSLGGCEIFSY----ENRMGPFEDCLWYFSFLFFNKKQRRVV 274
>Y7I4_ENCCU (Q8SUU2) Hypothetical protein ECU07_1840
Length = 161
Score = 36.6 bits (83), Expect = 0.048
Identities = 18/42 (42%), Positives = 25/42 (58%), Gaps = 4/42 (9%)
Query: 148 VSLDDCEIYGYLPDSEADPLPERGAIWSFNFLFYNRKLKRIV 189
VSL CEI+ Y E P +W F+FLF+N+K +R+V
Sbjct: 115 VSLGGCEIFSY----ENRMGPFEDCLWYFSFLFFNKKQRRVV 152
>MAF1_YEAST (P41910) Repressor of RNA polymerase III transcription
MAF1
Length = 395
Score = 35.8 bits (81), Expect = 0.082
Identities = 22/85 (25%), Positives = 41/85 (47%), Gaps = 1/85 (1%)
Query: 111 TSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPER 170
T F + + T +S+F + + +++ ++ ++L DC ++ Y P S + E
Sbjct: 257 TDFVKTSLKTFISKFENTLYSLGRQPEEWVWEVINSHMTLSDCVLFQYSP-SNSFLEDEP 315
Query: 171 GAIWSFNFLFYNRKLKRIVTFRLSC 195
G +W+ YNRK KR+ L C
Sbjct: 316 GYLWNLIGFLYNRKRKRVAYLYLIC 340
>MAF1_CANGA (Q6FIZ7) Repressor of RNA polymerase III transcription
MAF1
Length = 391
Score = 34.3 bits (77), Expect = 0.24
Identities = 19/81 (23%), Positives = 43/81 (52%), Gaps = 1/81 (1%)
Query: 108 LKLTSFSQRNHGTVLSRFLKPTCLKPQSLLDALFKAVDEVVSLDDCEIYGYLPDSEADPL 167
L+ T F + + + +++F + + +++ ++ ++L DC +Y Y P +++
Sbjct: 246 LEPTDFKRSSIKSFIAKFENSMYSLGRKPEEWIWEVINSHMTLSDCVLYQYCP-TQSFLD 304
Query: 168 PERGAIWSFNFLFYNRKLKRI 188
E G +WS F+N+K KR+
Sbjct: 305 DEPGHLWSLIGFFFNKKRKRV 325
>TOP1_PSEAE (Q9HZJ5) DNA topoisomerase I (EC 5.99.1.2)
(Omega-protein) (Relaxing enzyme) (Untwisting enzyme)
(Swivelase)
Length = 868
Score = 29.3 bits (64), Expect = 7.7
Identities = 17/40 (42%), Positives = 25/40 (62%), Gaps = 1/40 (2%)
Query: 167 LPERGAIWSFNFLFYNRKLKRIVTFRLS-CFSNLIADGFS 205
+ ERG + + N FY K+ IVT RL+ F+NL+ GF+
Sbjct: 509 IQERGYVTTHNRRFYAEKMGDIVTDRLNESFANLMDYGFT 548
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,683,744
Number of Sequences: 164201
Number of extensions: 1028064
Number of successful extensions: 2284
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 2272
Number of HSP's gapped (non-prelim): 14
length of query: 222
length of database: 59,974,054
effective HSP length: 106
effective length of query: 116
effective length of database: 42,568,748
effective search space: 4937974768
effective search space used: 4937974768
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144731.10


